Abstract
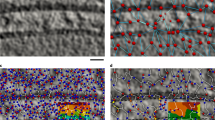
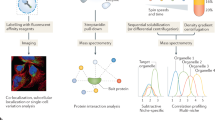
We developed a computational imaging approach that describes the three-dimensional spatial organization of endomembranes from micromanipulation-normalized mammalian cells with probabilistic density maps. Applied to several well-known marker proteins, this approach revealed the average steady-state organization of early endosomes, multivesicular bodies or lysosomes, endoplasmic reticulum exit sites, the Golgi apparatus and Golgi-derived transport carriers in crossbow-shaped cells. The steady-state organization of each tested endomembranous population was well-defined, unique and in some cases depended on the cellular adhesion geometry. Density maps of all endomembrane populations became stable when pooling several tens of cells only and were reproducible in independent experiments, allowing construction of a standardized cell model. We detected subtle changes in steady-state organization induced by disruption of the cellular cytoskeleton, with statistical significance observed for just 20 cells. Thus, combining micropatterning with construction of endomembrane density maps allows the systematic study of intracellular trafficking determinants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bornens, M. Organelle positioning and cell polarity. Nat. Rev. Mol. Cell Biol. 9, 874–886 (2008).
Caviston, J.P. & Holzbaur, E.L. Microtubule motors at the intersection of trafficking and transport. Trends Cell Biol. 16, 530–537 (2006).
Lanzetti, L. Actin in membrane trafficking. Curr. Opin. Cell Biol. 19, 453–458 (2007).
Ross, J.L., Ali, M.Y. & Warshaw, D.M. Cargo transport: molecular motors navigate a complex cytoskeleton. Curr. Opin. Cell Biol. 20, 41–47 (2008).
Insall, R.H. & Machesky, L.M. Actin dynamics at the leading edge: from simple machinery to complex networks. Dev. Cell 17, 310–322 (2009).
Schmoranzer, J. et al. Par3 and dynein associate to regulate local microtubule dynamics and centrosome orientation during migration. Curr. Biol. 19, 1065–1074 (2009).
Egea, G., Lazaro-Dieguez, F. & Vilella, M. Actin dynamics at the Golgi complex in mammalian cells. Curr. Opin. Cell Biol. 18, 168–178 (2006).
Rivero, S., Cardenas, J., Bornens, M. & Rios, R.M. Microtubule nucleation at the cis-side of the Golgi apparatus requires AKAP450 and GM130. EMBO J. 28, 1016–1028 (2009).
Semenova, I. et al. Actin dynamics is essential for myosin-based transport of membrane organelles. Curr. Biol. 18, 1581–1586 (2008).
Taunton, J. Actin filament nucleation by endosomes, lysosomes and secretory vesicles. Curr. Opin. Cell Biol. 13, 85–91 (2001).
Sachs, K., Perez, O., Pe'er, D., Lauffenburger, D.A. & Nolan, G.P. Causal protein-signaling networks derived from multiparameter single-cell data. Science 308, 523–529 (2005).
Sigal, A. et al. Variability and memory of protein levels in human cells. Nature 444, 643–646 (2006).
Snijder, B. et al. Population context determines cell-to-cell variability in endocytosis and virus infection. Nature 461, 520–523 (2009).
Liu, W.F. & Chen, C.S. Cellular and multicellular form and function. Adv. Drug Deliv. Rev. 59, 1319–1328 (2007).
Thery, M., Pepin, A., Dressaire, E., Chen, Y. & Bornens, M. Cell distribution of stress fibres in response to the geometry of the adhesive environment. Cell Motil. Cytoskeleton 63, 341–355 (2006).
Thery, M. et al. Anisotropy of cell adhesive microenvironment governs cell internal organization and orientation of polarity. Proc. Natl. Acad. Sci. USA 103, 19771–19776 (2006).
Racine, V. et al. Visualization and quantification of vesicle trafficking on a three-dimensional cytoskeleton network in living cells. J. Microsc. 225, 214–228 (2007).
Bowman, A.W. & Foster, P. Density based exploration of bivariate data. Stat. Comput. 3, 171–177 (1993).
Hyndman, R. Computing and graphing highest density regions. Am. Stat. 50, 120–126 (1996).
Pols, M.S. & Klumperman, J. Trafficking and function of the tetraspanin CD63. Exp. Cell Res. 315, 1584–1592 (2009).
Chavrier, P., Parton, R.G., Hauri, H.P., Simons, K. & Zerial, M. Localization of low molecular weight GTP binding proteins to exocytic and endocytic compartments. Cell 62, 317–329 (1990).
Tang, B.L. et al. The mammalian homolog of yeast Sec13p is enriched in the intermediate compartment and is essential for protein transport from the endoplasmic reticulum to the Golgi apparatus. Mol. Cell. Biol. 17, 256–266 (1997).
Antony, C. et al. The small GTP-binding protein rab6p is distributed from medial Golgi to the trans-Golgi network as determined by a confocal microscopic approach. J. Cell Sci. 103, 785–796 (1992).
Grigoriev, I. et al. Rab6 regulates transport and targeting of exocytotic carriers. Dev. Cell 13, 305–314 (2007).
White, J. et al. Rab6 coordinates a novel Golgi to ER retrograde transport pathway in live cells. J. Cell Biol. 147, 743–760 (1999).
Gretton, A., Borgwardt, K.M., Rasch, M.J., Schoelkopf, B. & Smola, A. A kernel method for the two-sample problem. Advances in Neural Information Processing Systems 19: Proceedings of the 2006 Conference 513–520 (MIT Press, Cambridge, Masssachusetts, USA, 2007).
Wodarz, A. & Näthke, I. Cell polarity in development and cancer. Nat. Cell Biol. 9, 1016–1024 (2007).
Rodriguez Boulan, E. & Sabatini, D.D. Asymmetric budding of viruses in epithelial monlayers: a model system for study of epithelial polarity. Proc. Natl. Acad. Sci. USA 75, 5071–5075 (1978).
Snider, J. et al. Intracellular actin-based transport: how far you go depends on how often you switch. Proc. Natl. Acad. Sci. USA 101, 13204–13209 (2004).
Azioune, A., Storch, M., Bornens, M., Théry, M. & Piel, M. Simple and rapid process for single cell micro-patterning. Lab Chip 9, 1640–1642 (2009).
Sibarita, J.B. Deconvolution microscopy. Adv. Biochem. Eng. Biotechnol. 95, 201–243 (2005).
Simonoff, J.S. Smoothing Methods for Statistics. (Springer, New York, 1996).
Duong, T. & Hazelton, M.L. Plug-in bandwidth matrices for bivariate kernel density estimations. J. Nonparametr. Stat. 17, 17–30 (2003).
Acknowledgements
We acknowledge L. Sengmanivong of the Nikon Imaging Centre at Institut Curie–Centre National de la Recherche Scientifique and V. Fraisier of the Plate-forme Imagerie Cellulaire et Tissulaire–Infrastructures en Biologie Santé et Agronomie Imaging Facility for their extensive help with microscopes and in particular their help using the deconvolution service of the facility. We thank J.-B. Sibarita for advice on image analysis including use of the multidimensional image analysis program and fruitful discussion during early phases of the project; I. Brito for statistical advice; W. Hong (Institute of Molecular and Cell Biology, Singapore) for providing the Sec13 antibody; M. Piel, A. Azioune and J. Fink for help with microprinting; and G. Egea, S. Miserey, A. Echard and J. Enninga for critical reading of the manuscript. K.S. received funding from the Fondation pour la Recherche Médicale en France and Association pour la Recherche sur le Cancer. This project was supported by grants from the Centre National de la Recherche Scientifique and Institut Curie.
Author information
Authors and Affiliations
Contributions
K.S. and B.G. designed the research, K.S. performed the experiments and analysis and wrote the manuscript, T.D. developed the density calculation, K.B. developed the statistical analysis and edited the manuscript, S.B. adjusted patterning techniques and M.B. contributed to the conception of the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Table 1 and Supplementary Notes 1–2 (PDF 2412 kb)
Supplementary Video 1
Maximum intensity projection of the deconvolved fluorescence of GFP-Rab6-positive cells (n = 82) under control conditions. (AVI 2507 kb)
Supplementary Video 2
Maximum intensity projection of the deconvolved fluorescence of GFP-Rab6-positive cells (n = 47) after nocodazole treatment. (AVI 1383 kb)
Supplementary Video 3
Maximum intensity projection of the deconvolved fluorescence of GFP-Rab6-positive cells (n = 50) after cytochalasin D treatment. (AVI 1667 kb)
Rights and permissions
About this article
Cite this article
Schauer, K., Duong, T., Bleakley, K. et al. Probabilistic density maps to study global endomembrane organization. Nat Methods 7, 560–566 (2010). https://doi.org/10.1038/nmeth.1462
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1462
This article is cited by
-
Transcription factor EB regulates phosphatidylinositol-3-phosphate levels that control lysosome positioning in the bladder cancer model
Communications Biology (2023)
-
Cellular harmonics for the morphology-invariant analysis of molecular organization at the cell surface
Nature Computational Science (2023)
-
Integrated intracellular organization and its variations in human iPS cells
Nature (2023)
-
Cell shape regulates subcellular organelle location to control early Ca2+ signal dynamics in vascular smooth muscle cells
Scientific Reports (2020)
-
Actomyosin-driven force patterning controls endocytosis at the immune synapse
Nature Communications (2019)