Abstract
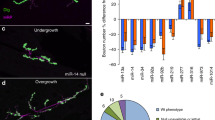
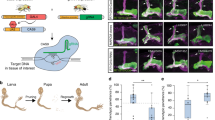
MicroRNAs are important regulators of gene expression, yet the functional outputs of most microRNA-target interactions remain elusive. Here we introduce the Drosophila melanogaster microRNA sponge (miR-SP) as a powerful transgenic technology to dissect the function of every microRNA with precise spatiotemporal resolution. miR-SPs can be used to characterize tissue-specific microRNA loss-of-function phenotypes, define the spatial regulation of their effectors and uncover interactions between microRNAs and other genes. Using themiR-SP system, we identified an essential role of the conserved microRNA miR-8, in neuromuscular junction formation. Tissue-specific silencing revealed that postsynaptic activity of miR-8 is important for normal neuromuscular junction morphogenesis. Given that miR-SPs rely on a bipartite modular expression system, they could be used to elucidate the endogenous function of microRNAs in any species in which conditional expression can be achieved.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bushati, N. & Cohen, S.M. microRNA functions. Annu. Rev. Cell Dev. Biol. 23, 175–205 (2007).
Eulalio, A., Huntzinger, E. & Izaurralde, E. Getting to the root of miRNA-mediated gene silencing. Cell 132, 9–14 (2008).
Aboobaker, A.A., Tomancak, P., Patel, N., Rubin, G.M. & Lai, E.C. Drosophila microRNAs exhibit diverse spatial expression patterns during embryonic development. Proc. Natl. Acad. Sci. USA 102, 18017–18022 (2005).
Kawaji, H. & Hayashizaki, Y. Exploration of small RNAs. PLoS Genet. 4, e22 (2008).
Silver, S.J., Hagen, J.W., Okamura, K., Perrimon, N. & Lai, E.C. Functional screening identifies miR-315 as a potent activator of Wingless signaling. Proc. Natl. Acad. Sci. USA 104, 18151–18156 (2007).
Lim, L.P. et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433, 769–773 (2005).
Hutvagner, G., Simard, M.J., Mello, C.C. & Zamore, P.D. Sequence-specific inhibition of small RNA function. PLoS Biol. 2, E98 (2004).
Ebert, M.S., Neilson, J.R. & Sharp, P.A. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat. Methods 4, 721–726 (2007).
Brown, B.D. & Naldini, L. Exploiting and antagonizing microRNA regulation for therapeutic and experimental applications. Nat. Rev. Genet. 10, 578–585 (2009).
Stern, P. et al. A system for Cre-regulated RNA interference in vivo. Proc. Natl. Acad. Sci. USA 105, 13895–13900 (2008).
Elliott, D.A. & Brand, A.H. The GAL4 system: a versatile system for the expression of genes. Methods Mol. Biol. 420, 79–95 (2008).
Stark, A., Brennecke, J., Bushati, N., Russell, R.B. & Cohen, S.M. Animal microRNAs confer robustness to gene expression and have a significant impact on 3′UTR evolution. Cell 123, 1133–1146 (2005).
Kuzin, A., Kundu, M., Brody, T. & Odenwald, W.F. The Drosophila nerfin-1 mRNA requires multiple microRNAs to regulate its spatial and temporal translation dynamics in the developing nervous system. Dev. Biol. 310, 35–43 (2007).
Li, Y., Wang, F., Lee, J.A. & Gao, F.B. MicroRNA-9a ensures the precise specification of sensory organ precursors in Drosophila. Genes Dev. 20, 2793–2805 (2006).
Li, X. & Carthew, R.W. A microRNA mediates EGF receptor signaling and promotes photoreceptor differentiation in the Drosophila eye. Cell 123, 1267–1277 (2005).
Karres, J.S., Hilgers, V., Carrera, I., Treisman, J. & Cohen, S.M. The conserved microRNA miR-8 tunes atrophin levels to prevent neurodegeneration in Drosophila. Cell 131, 136–145 (2007).
Kosik, K.S. The neuronal microRNA system. Nat. Rev. Neurosci. 7, 911–920 (2006).
Lugli, G., Torvik, V.I., Larson, J. & Smalheiser, N.R. Expression of microRNAs and their precursors in synaptic fractions of adult mouse forebrain. J. Neurochem. 106, 650–661 (2008).
Collins, C.A. & DiAntonio, A. Synaptic development: insights from Drosophila. Curr. Opin. Neurobiol. 17, 35–42 (2007).
Keshishian, H. & Kim, Y.S. Orchestrating development and function: retrograde BMP signaling in the Drosophila nervous system. Trends Neurosci. 27, 143–147 (2004).
Shcherbata, H.R. et al. Stage-specific differences in the requirements for germline stem cell maintenance in the Drosophila ovary. Cell Stem Cell 1, 698–709 (2007).
Drees, F. & Gertler, F.B. Ena/VASP: proteins at the tip of the nervous system. Curr. Opin. Neurobiol. 18, 53–59 (2008).
Gates, J. et al. Enabled plays key roles in embryonic epithelial morphogenesis in Drosophila. Development 134, 2027–2039 (2007).
Asakawa, K. & Kawakami, K. Targeted gene expression by the Gal4-UAS system in zebrafish. Dev. Growth Differ. 50, 391–399 (2008).
Sauer, B. & Henderson, N. Cre-stimulated recombination at loxP-containing DNA sequences placed into the mammalian genome. Nucleic Acids Res. 17, 147–161 (1989).
Rebay, I. & Rubin, G.M. Yan functions as a general inhibitor of differentiation and is negatively regulated by activation of the Ras1/MAPK pathway. Cell 81, 857–866 (1995).
Comer, A.R., Ahern-Djamali, S.M., Juang, J.L., Jackson, P.D. & Hoffmann, F.M. Phosphorylation of Enabled by the Drosophila Abelson tyrosine kinase regulates the in vivo function and protein-protein interactions of Enabled. Mol. Cell. Biol. 18, 152–160 (1998).
Brand, A.H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Johnson, K.G. et al. The HSPGs Syndecan and Dallylike bind the receptor phosphatase LAR and exert distinct effects on synaptic development. Neuron 49, 517–531 (2006).
Grimson, A. et al. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol. Cell 27, 91–105 (2007).
Rehmsmeier, M., Steffen, P., Hochsmann, M. & Giegerich, R. Fast and effective prediction of microRNA/target duplexes. RNA 10, 1507–1517 (2004).
Acknowledgements
We thank M. Greenberg, N. Perrimon, T. Schwarz, M. Feany and J. Flanagan for critical feedback on the manuscript; M. Ebert for advice on designing the miR-SP constructs and for comments on the manuscript; V. Sridhar and members of the Harvard NeuroDiscovery Center Optical Imaging program for technical assistance; R.W. Carthew (Northwestern University) for GMR-YanAct; F.B. Gao (University of California San Francisco) for miR-9aJ22 and miR-9aE39; M. Peifer (University of North Carolina) for UAS-FP4-mitoEGFP; F.M. Hoffmann (University of Wisconsin-Madison) for ena210 and UAS-ena; H. Ruohola-Baker (University of Washington) for miR-8-GFP sensor; J. Brennecke (Institutes of Molecular Biotechnology of the Austrian Academy of Sciences, Vienna) for nerfin-1 3′ UTR reporter; S. Cohen (Temasek Life Sciences Laboratory) for miR-8Δ2; and members of the Developmental Studies Hybridoma Bank (University of Iowa, Department of Biological Sciences) for antibodies. C.M.L., C.S.L. and D.V.V. were supported by a grant from the National Institutes of Health (NS40043). T.A.F. was supported in part by a fellowship from the John Douglas French Alzheimer's Foundation.
Author information
Authors and Affiliations
Contributions
T.A.F. conceived and designed the miR-SP technology. C.M.L. and T.A.F. designed and performed the experiments, analyzed the data and interpreted the results. C.S.L. generated the miR-8 null mutant and independent of this study performed preliminary NMJ analysis and proposed Ena as a candidate miR-8 effector. D.V.V. supervised the study and provided material and salary support for C.M.L., C.S.L. and T.A.F. The text and figures were drafted by T.A.F., and T.A.F., D.V.V. and C.M.L. edited them.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–2 (PDF 1828 kb)
Rights and permissions
About this article
Cite this article
Loya, C., Lu, C., Van Vactor, D. et al. Transgenic microRNA inhibition with spatiotemporal specificity in intact organisms. Nat Methods 6, 897–903 (2009). https://doi.org/10.1038/nmeth.1402
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1402
This article is cited by
-
Drosophila enabled promotes synapse morphogenesis and regulates active zone form and function
Neural Development (2020)
-
The conserved microRNA miR-34 regulates synaptogenesis via coordination of distinct mechanisms in presynaptic and postsynaptic cells
Nature Communications (2020)
-
Emerging ways to treat breast cancer: will promises be met?
Cellular Oncology (2018)
-
Global identification of microRNAs associated with chlorantraniliprole resistance in diamondback moth Plutella xylostella (L.)
Scientific Reports (2017)
-
Analysis of secondary structural elements in human microRNA hairpin precursors
BMC Bioinformatics (2016)