Abstract
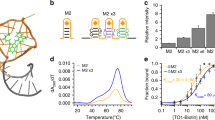
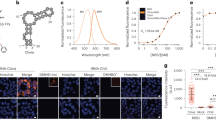
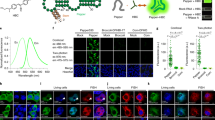
To visualize native or non-engineered RNA in live cells with single-molecule sensitivity, we developed multiply labeled tetravalent RNA imaging probes (MTRIPs). When delivered with streptolysin O into living human epithelial cancer cells and primary chicken fibroblasts, MTRIPs allowed the accurate imaging of native mRNAs and a non-engineered viral RNA, of RNA co-localization with known RNA-binding proteins, and of RNA dynamics and interactions with stress granules.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Fusco, D. et al. Curr. Biol. 13, 161–167 (2003).
Shav-Tal, Y. et al. Science 304, 1797–1800 (2004).
Vargas, D.Y., Raj, A., Marras, S.A., Kramer, F.R. & Tyagi, S. Proc. Natl. Acad. Sci. USA 102, 17008–17013 (2005).
Santangelo, P.J. & Bao, G. Nucleic Acids Res. 35, 3602–3611 (2007).
Utley, T.J. et al. Proc. Natl. Acad. Sci. USA 105, 10209–10214 (2008).
Femino, A.M., Fay, F.S., Fogarty, K. & Singer, R.H. Science 280, 585–590 (1998).
Latham, V.M. Jr, Kislauskis, E.H., Singer, R.H. & Ross, A.F. J. Cell Biol. 126, 1211–1219 (1994).
Ross, A.F., Oleynikov, Y., Kislauskis, E.H., Taneja, K.L. & Singer, R.H. Mol. Cell. Biol. 17, 2158–2165 (1997).
Iseni, F. et al. EMBO J. 21, 5141–5150 (2002).
Kedersha, N. et al. Mol. Biol. Cell 13, 195–210 (2002).
Zeitelhofer, M. et al. J. Neurosci. 28, 7555–7562 (2008).
Kedersha, N. et al. J. Cell Biol. 169, 871–884 (2005).
Mollet, S. et al. Mol. Biol. Cell 19, 4469–4479 (2008).
Conover, W.J. Practical Nonparametric Statistics 3rd edn. (Wiley, New York, 1999).
Kedersha, N. et al. J. Cell Biol. 151, 1257–1268 (2000).
Acknowledgements
This study was supported by the Georgia Institute of Technology administration (P.J.S.).
Author information
Authors and Affiliations
Contributions
P.J.S. designed and performed experiments, analyzed data and wrote manuscript, A.W.L. performed experiments and analyzed data, P.C. performed experiments, Y.S. and G.J.B. provided novel antibody, M.E.L. and J.E.C. provided virus and antibody and edited manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1-10, Supplementary Table 1 and Supplementary Results (PDF 5705 kb)
Supplementary Video 1
Time-lapse images of β-actin mRNA granules in live cell in one optical plane taken at 1 Hz for 3.0 minutes; first 1 minute is played back at 3 frames a second for 20 s. (MOV 2584 kb)
Supplementary Video 2
Time-lapse images of the trajectory of a single β-actin mRNA granule in a live cell in one optical plane taken at 1 Hz but played back at 3 frames a second. This movie represents only 70 s of the 3 minutes of total data taken in this cell. (MOV 3319 kb)
Supplementary Video 3
Time-lapse images of the trajectory of a single β-actin mRNA granule in a live cell in one optical plane taken at 5 Hz but played back at 3 frames a second for 30 s. (MOV 2375 kb)
Supplementary Video 4
Time-lapse images of stress granule (green) collision and penetration of a viral RNA granule (red). Images were taken every 5 seconds, and playback is at 3 frames per second. (MOV 529 kb)
Supplementary Video 5
Time-lapse images of the separation of the stress granule (green) with the viral RNA granule (red). Images were taken every 5 seconds, and playback is at 3 frames per second. (MOV 429 kb)
Supplementary Video 6
Images of the docking of a stress granule (green) and viral RNA granule (red). Images were taken every 5 seconds, and playback is at 3 frames per second. (MOV 610 kb)
Supplementary Video 7
Movie showing an example of granule fusion, where both viral RNA (red) and TIA-1 (green) are imaged. Images were taken every 5 seconds, and playback is at 3 frames per second. (MOV 1174 kb)
Supplementary Video 8
Movie showing an example of granule splitting, where both viral RNA (red) and TIA-1 (green) are imaged. Images were taken every 5 seconds, and playback is at 3 frames per second. (MOV 928 kb)
Rights and permissions
About this article
Cite this article
Santangelo, P., Lifland, A., Curt, P. et al. Single molecule–sensitive probes for imaging RNA in live cells. Nat Methods 6, 347–349 (2009). https://doi.org/10.1038/nmeth.1316
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1316
This article is cited by
-
Engineered mRNA-expressed antibodies prevent respiratory syncytial virus infection
Nature Communications (2018)
-
RSV glycoprotein and genomic RNA dynamics reveal filament assembly prior to the plasma membrane
Nature Communications (2017)
-
Computing in mammalian cells with nucleic acid strand exchange
Nature Nanotechnology (2016)
-
DNA nanotechnology from the test tube to the cell
Nature Nanotechnology (2015)
-
Tandem Spinach Array for mRNA Imaging in Living Bacterial Cells
Scientific Reports (2015)