Abstract
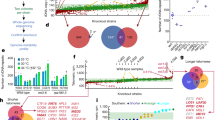
Functional genomic studies in Saccharomyces cerevisiae have contributed enormously to our understanding of cellular processes. Their full potential, however, has been hampered by the limited availability of reagents to systematically study essential genes and the inability to quantify the small effects of most gene deletions on growth. Here we describe the construction of a library of hypomorphic alleles of essential genes and a high-throughput growth competition assay to measure fitness with unprecedented sensitivity. These tools dramatically increase the breadth and precision with which quantitative genetic analysis can be performed in yeast. We illustrate the value of these approaches by using genetic interactions to reveal new relationships between chromatin-modifying factors and to create a functional map of the proteasome. Finally, by measuring the fitness of strains in the yeast deletion library, we addressed an enigma regarding the apparent prevalence of gene dispensability and found that most genes do contribute to growth.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Schuldiner, M. et al. Exploration of the function and organization of the yeast early secretory pathway through an epistatic miniarray profile. Cell 123, 507–519 (2005).
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Pan, X. et al. A robust toolkit for functional profiling of the yeast genome. Mol. Cell 16, 487–496 (2004).
Boone, C., Bussey, H. & Andrews, B.J. Exploring genetic interactions and networks with yeast. Nat. Rev. Genet. 8, 437–449 (2007).
Segre, D., Deluna, A., Church, G.M. & Kishony, R. Modular epistasis in yeast metabolism. Nat. Genet. 37, 77–83 (2005).
Mani, R., St. Onge, R.P., Hartman, J.L. IV, Giaever, G. & Roth, F.P. Defining genetic interaction. Proc. Natl. Acad. Sci. USA 105, 3461–3466 (2008).
Kelley, R. & Ideker, T. Systematic interpretation of genetic interactions using protein networks. Nat. Biotechnol. 23, 561–566 (2005).
Collins, S.R. et al. Functional dissection of protein complexes involved in yeast chromosome biology using a genetic interaction map. Nature 446, 806–810 (2007).
Warringer, J., Ericson, E., Fernandez, L., Nerman, O. & Blomberg, A. High-resolution yeast phenomics resolves different physiological features in the saline response. Proc. Natl. Acad. Sci. USA 100, 15724–15729 (2003).
Deutschbauer, A.M. et al. Mechanisms of haploinsufficiency revealed by genome-wide profiling in yeast. Genetics 169, 1915–1925 (2005).
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Nash, R. et al. Expanded protein information at SGD: new pages and proteome browser. Nucleic Acids Res. 35, D468–D471 (2007).
Mnaimneh, S. et al. Exploration of essential gene functions via titratable promoter alleles. Cell 118, 31–44 (2004).
Kanemaki, M., Sanchez-Diaz, A., Gambus, A. & Labib, K. Functional proteomic identification of DNA replication proteins by induced proteolysis in vivo. Nature 423, 720–724 (2003).
Gilon, T., Chomsky, O. & Kulka, R.G. Degradation signals for ubiquitin system proteolysis in Saccharomyces cerevisiae. EMBO J. 17, 2759–2766 (1998).
St. Onge, R.P. et al. Systematic pathway analysis using high-resolution fitness profiling of combinatorial gene deletions. Nat. Genet. 39, 199–206 (2007).
Newman, J.R. et al. Single-cell proteomic analysis of S. cerevisiae reveals the architecture of biological noise. Nature 441, 840–846 (2006).
Wang, Y. et al. Precision and functional specificity in mRNA decay. Proc. Natl. Acad. Sci. USA 99, 5860–5865 (2002).
Gasch, A.P. et al. Genomic expression programs in the response of yeast cells to environmental changes. Mol. Biol. Cell 11, 4241–4257 (2000).
Giaever, G. et al. Genomic profiling of drug sensitivities via induced haploinsufficiency. Nat. Genet. 21, 278–283 (1999).
Lum, P.Y. et al. Discovering modes of action for therapeutic compounds using a genome-wide screen of yeast heterozygotes. Cell 116, 121–137 (2004).
Barnes, G., Hansen, W.J., Holcomb, C.L. & Rine, J. Asparagine-linked glycosylation in Saccharomyces cerevisiae: genetic analysis of an early step. Mol. Cell. Biol. 4, 2381–2388 (1984).
Ungar, D., Oka, T., Krieger, M. & Hughson, F.M. Retrograde transport on the COG railway. Trends Cell Biol. 16, 113–120 (2006).
Zlatanova, J. & Thakar, A. H2A.Z: view from the top. Structure 16, 166–179 (2008).
Yang, X.J. & Seto, E. The Rpd3/Hda1 family of lysine deacetylases: from bacteria and yeast to mice and men. Nat. Rev. Mol. Cell Biol. 9, 206–218 (2008).
Tompa, R. & Madhani, H.D. Histone H3 lysine 36 methylation antagonizes silencing in Saccharomyces cerevisiae independently of the Rpd3S histone deacetylase complex. Genetics 175, 585–593 (2007).
Hanna, J. & Finley, D. A proteasome for all occasions. FEBS Lett. 581, 2854–2861 (2007).
Collins, G.A. & Tansey, W.P. The proteasome: a utility tool for transcription? Curr. Opin. Genet. Dev. 16, 197–202 (2006).
Velichutina, I., Connerly, P.L., Arendt, C.S., Li, X. & Hochstrasser, M. Plasticity in eucaryotic 20S proteasome ring assembly revealed by a subunit deletion in yeast. EMBO J. 23, 500–510 (2004).
Gu, Z. et al. Role of duplicate genes in genetic robustness against null mutations. Nature 421, 63–66 (2003).
Papp, B., Pal, C. & Hurst, L.D. Metabolic network analysis of the causes and evolution of enzyme dispensability in yeast. Nature 429, 661–664 (2004).
Hillenmeyer, M.E. et al. The chemical genomic portrait of yeast: uncovering a phenotype for all genes. Science 320, 362–365 (2008).
Shaner, N.C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 22, 1567–1572 (2004).
Acknowledgements
We acknowledge J. Hill and E. Rodriguez for help with strain construction; C.-S. Chin for flow cytometry software development; J. Newman for an introduction to high-throughput flow cytometry; R. Tsien (University of California, San Diego) for a construct encoding dTomato; B. Toyama for assistance with graphics; C. Boone, G. Giaever and C. Nislow for communication of results before publication; and V. Denic, J. Hollien, J. Ihmels, N. Ingolia, J. Newman and members of the Weissman laboratory for helpful discussion and critical reading of the manuscript. This work was supported by funding from the Howard Hughes Medical Institute (J.S.W.), the Fannie and John Hertz Foundation (D.K.B.), the US National Science Foundation (D.K.B) and the Larry L. Hillblom Foundation (D.M.C.).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–2, Supplementary Note, Supplementary Methods (PDF 1848 kb)
Supplementary Data 1
List of primers used in this study. (XLS 425 kb)
Supplementary Data 2
Growth rate measurements for flow cytometry assay validation. (XLS 88 kb)
Supplementary Data 3
DAmP library growth rates measured by flow cytometry. (XLS 185 kb)
Supplementary Data 4
Deletion library growth rates measured by flow cytometry. (XLS 476 kb)
Supplementary Data 5
Growth rate measurements used for genetic interaction data. (XLS 85 kb)
Rights and permissions
About this article
Cite this article
Breslow, D., Cameron, D., Collins, S. et al. A comprehensive strategy enabling high-resolution functional analysis of the yeast genome. Nat Methods 5, 711–718 (2008). https://doi.org/10.1038/nmeth.1234
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1234
This article is cited by
-
From beer to breadboards: yeast as a force for biological innovation
Genome Biology (2024)
-
Plasma membrane damage limits replicative lifespan in yeast and induces premature senescence in human fibroblasts
Nature Aging (2024)
-
Evolutionary conservation of the fidelity of transcription
Nature Communications (2023)
-
Resolving Deleterious and Near-Neutral Effects Requires Different Pooled Fitness Assay Designs
Journal of Molecular Evolution (2023)
-
Epigenetics Identifier screens reveal regulators of chromatin acylation and limited specificity of acylation antibodies
Scientific Reports (2021)