Abstract
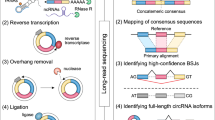
Describing the 'ORFeome' of an organism, including all major isoforms, is essential for a system-level understanding of any species; however, conventional cloning and sequencing approaches are prohibitively costly and labor-intensive. We describe a potentially genome-wide methodology for efficiently capturing new coding isoforms using reverse transcriptase (RT)-PCR recombinational cloning, 'deep-well' pooling and a next-generation sequencing platform. This ORFeome discovery pipeline will be applicable to any eukaryotic species with a sequenced genome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Johnson, J.M. et al. Science 302, 2141–2144 (2003).
Zahler, A.M. WormBook 2005, 1–13 (2005).
Schuster, S.C. Nat. Methods 5, 16–18 (2008).
Margulies, M. et al. Nature 437, 376–380 (2005).
Shendure, J. et al. Science 309, 1728–1732 (2005).
Moore, M.J. et al. BMC Plant Biol. 6, 17 (2006).
Oh, J.D. et al. Proc. Natl. Acad. Sci. USA 103, 9999–10004 (2006).
Torres, T.T., Metta, M., Ottenwalder, B. & Schlotterer, C. Genome Res. 18, 172–177 (2008).
Porreca, G.J. et al. Nat. Methods 4, 931–936 (2007).
Emrich, S.J., Barbazuk, W.B., Li, L. & Schnable, P.S. Genome Res. 17, 69–73 (2007).
Wicker, T. et al. BMC Genomics 7, 275 (2006).
Walhout, A.J. et al. Methods Enzymol. 328, 575–592 (2000).
The MGC Project Team. Genome Res. 14, 2121–2127 (2004).
Rual, J.F. et al. Genome Res. 14, 2128–2135 (2004).
Lamesch, P. et al. Genomics 89, 307–315 (2007).
Hamosh, A., Scott, A.F., Amberger, J.S., Bocchini, C.A. & McKusick, V.A. Nucleic Acids Res. 33, D514–D517 (2005).
Clamp, M. et al. Proc. Natl. Acad. Sci. USA 104, 19428–19433 (2007).
Yamasaki, C. et al. Nucleic Acids Res. 36, D793–D799 (2008).
Acknowledgements
This work was funded in part by a grant from the Ellison Foundation (awarded to M.V.) and in part by the Dana Farber Cancer Institute Strategic Initiative in support of Center for Cancer Systems Biology. F.P.R. acknowledges support from US National Institutes of Health grants NS054052, HG003224, HL081341. W.T. was supported in part by National Institutes of Health grant DK070078. We thank the West Quad Computing Group at Harvard Medical School as well as Research Computing at Massachusetts General Hospital for assistance with computational resources. We thank G. Temple for helpful comments on the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
K.M., L.S., N.T. and D.R.S. are employed by Agencourt Corp.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Note, Supplementary Methods (PDF 1421 kb)
Rights and permissions
About this article
Cite this article
Salehi-Ashtiani, K., Yang, X., Derti, A. et al. Isoform discovery by targeted cloning, 'deep-well' pooling and parallel sequencing. Nat Methods 5, 597–600 (2008). https://doi.org/10.1038/nmeth.1224
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1224
This article is cited by
-
ORF Capture-Seq as a versatile method for targeted identification of full-length isoforms
Nature Communications (2020)
-
Protein interaction network of alternatively spliced isoforms from brain links genetic risk factors for autism
Nature Communications (2014)
-
Incorporation of non-natural nucleotides into template-switching oligonucleotides reduces background and improves cDNA synthesis from very small RNA samples
BMC Genomics (2010)
-
Towards reliable isoform quantification using RNA-SEQ data
BMC Bioinformatics (2010)
-
Transcriptome sequencing of the Microarray Quality Control (MAQC) RNA reference samples using next generation sequencing
BMC Genomics (2009)