Abstract
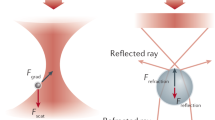
Single-molecule force spectroscopy has emerged as a powerful tool to investigate the forces and motions associated with biological molecules and enzymatic activity. The most common force spectroscopy techniques are optical tweezers, magnetic tweezers and atomic force microscopy. Here we describe these techniques and illustrate them with examples highlighting current capabilities and limitations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cluzel, P. et al. DNA: An extensible molecule. Science 271, 792–794 (1996).
Evans, E., Ritchie, K. & Merkel, R. Sensitive force technique to probe molecular adhesion and structural linkages at biological interfaces. Biophys. J. 68, 2580–2587 (1995).
Smith, S.B., Finzi, L. & Bustamante, C. Direct mechanical measurements of the elasticity of single DNA molecules by using magnetic beads. Science 258, 1122–1126 (1992).
Kim, S.J., Blainey, P.C., Schroeder, C.M. & Xie, X.S. Multiplexed single-molecule assay for enzymatic activity on flow-stretched DNA. Nat. Methods 4, 397–399 (2007).
Greenleaf, W.J., Woodside, M.T. & Block, S.M. High-resolution, single-molecule measurements of biomolecular motion. Annu. Rev. Biophys. Biomol. Struct. 36, 171–190 (2007).
Neuman, K.C. & Block, S.M. Optical trapping. Rev. Sci. Instrum. 75, 2787–2809 (2004). A detailed and thorough technical review of optical trapping.
Tanase, M., Biais, N. & Sheetz, M. Magnetic tweezers in cell biology. Methods Cell Biol. 83, 473–493 (2007).
Zlatanova, J., Lindsay, S.M. & Leuba, S.H. Single molecule force spectroscopy in biology using the atomic force microscope. Prog. Biophys. Mol. Biol. 74, 37–61 (2000).
Lee, C.K., Wang, Y.M., Huang, L.S. & Lin, S. Atomic force microscopy: determination of unbinding force, off rate and energy barrier for protein-ligand interaction. Micron 38, 446–461 (2007).
Lang, M.J., Asbury, C.L., Shaevitz, J.W. & Block, S.M. An automated two-dimensional optical force clamp for single molecule studies. Biophys. J. 83, 491–501 (2002).
Neuman, K.C., Lionnet, T. & Allemand, J.F. Single-molecule micromanipulation techniques. Annu. Rev. Mater. Res. 37, 33–67 (2007).
Abbondanzieri, E.A., Greenleaf, W.J., Shaevitz, J.W., Landick, R. & Block, S.M. Direct observation of base-pair stepping by RNA polymerase. Nature 438, 460–465 (2005). This is an experimental tour de force in which individual 0.34 nm base-pair steps of transcribing RNA polymerases were directly measured with optical tweezers.
Grandbois, M., Beyer, M., Rief, M., Clausen-Schaumann, H. & Gaub, H.E. How strong is a covalent bond? Science 283, 1727–1730 (1999).
Hohng, S. et al. Fluorescence-force spectroscopy maps two-dimensional reaction landscape of the holliday junction. Science 318, 279–283 (2007).
Litvinov, R.I., Shuman, H., Bennett, J.S. & Weisel, J.W. Binding strength and activation state of single fibrinogen-integrin pairs on living cells. Proc. Natl. Acad. Sci. USA 99, 7426–7431 (2002).
Prass, M., Jacobson, K., Mogilner, A. & Radmacher, M. Direct measurement of the lamellipodial protrusive force in a migrating cell. J. Cell Biol. 174, 767–772 (2006).
MacKintosh, F.C. & Schmidt, C.F. Microrheology. Curr. Opin. Colloid Interface Sci. 4, 300–307 (1999).
Daniels, B.R., Masi, B.C. & Wirtz, D. Probing single-cell micromechanics in vivo: the microrheology of C. elegans developing embryos. Biophys. J. 90, 4712–4719 (2006).
Block, S.M., Goldstein, L.S. & Schnapp, B.J. Bead movement by single kinesin molecules studied with optical tweezers. Nature 348, 348–352 (1990).
Svoboda, K. & Block, S.M. Force and velocity measured for single kinesin molecules. Cell 77, 773–784 (1994).
Simmons, R.M. et al. Force on single actin filaments in a motility assay measured with an optical trap. Adv. Exp. Med. Biol. 332, 331–336 (1993).
Tinoco, I. Jr & Bustamante, C. The effect of force on thermodynamics and kinetics of single molecule reactions. Biophys. Chem. 101–102, 513–533 (2002). A detailed and comprehensive treatment of the effects of force on single-molecule reactions; the concepts and analysis presented form the underpinning of single-molecule force spectroscopy.
Tskhovrebova, L., Trinick, J., Sleep, J.A. & Simmons, R.M. Elasticity and unfolding of single molecules of the giant muscle protein titin. Nature 387, 308–312 (1997).
Kellermayer, M.S.Z., Smith, S.B., Granzier, H.L. & Bustamante, C. Folding-unfolding transitions in single titin molecules characterized with laser tweezers. Science 276, 1112–1116 (1997).
Rief, M., Gautel, M., Oesterhelt, F., Fernandez, J.M. & Gaub, H.E. Reversible unfolding of individual titin immunoglobulin domains by AFM. Science 276, 1109–1112 (1997). This paper and the one above (ref. 24 ) appeared together and described the first demonstrations of the mechanical unfolding of an individual protein.
Hummer, G. & Szabo, A. Free energy surfaces from single-molecule force spectroscopy. Acc. Chem. Res. 38, 504–513 (2005).
Woodside, M.T. et al. Nanomechanical measurements of the sequence-dependent folding landscapes of single nucleic acid hairpins. Proc. Natl. Acad. Sci. USA 103, 6190–6195 (2006).
Lim, C.T., Zhou, E.H., Li, A., Vedula, S.R.K. & Fu, H.X. Experimental techniques for single cell and single molecule biomechanics. Mater. Sci. Eng. C-Biomimetic Supramol. Syst. 26, 1278–1288 (2006).
Zhuang, X.W. & Rief, M. Single-molecule folding. Curr. Opin. Struct. Biol. 13, 88–97 (2003).
Evans, E. Probing the relation between force–lifetime–and chemistry in single molecular bonds. Annu. Rev. Biophys. Biomol. Struct. 30, 105–128 (2001).
Jarzynski, C. Nonequilibrium equality for free energy differences. Phys. Rev. Lett. 78, 2690–2693 (1997). This seminal paper presents the remarkable Jarzynski equality that relates the equilibrium free energy difference to non-equilibrium measurements of work. The Jarzynski equality and subsequent relations based on the equality permit the extraction of folding free energies from out-of-equilibrium mechanical unfolding experiments.
Lu, H.P., Xun, L. & Xie, X.S. Single-molecule enzymatic dynamics. Science 282, 1877–1882 (1998).
Xie, X.S. & Lu, H.P. Single-molecule enzymology. J. Biol. Chem. 274, 15967–15970 (1999).
Hua, W., Young, E.C., Fleming, M.L. & Gelles, J. Coupling of kinesin steps to ATP hydrolysis. Nature 388, 390–393 (1997).
Schnitzer, M.J. & Block, S.M. Kinesin hydrolyses one ATP per 8-nm step. Nature 388, 386–390 (1997).
Hermanson, G. Bioconjugate Techniques. (Academic Press, San Diego 1996).
Hinterdorfer, P. & Dufrene, Y.F. Detection and localization of single molecular recognition events using atomic force microscopy. Nat. Methods 3, 347–355 (2006).
Ashkin, A., Dziedzic, J.M., Bjorkholm, J.E. & Chu, S. Observation of a single-beam gradient force optical trap for dielectric particles. Opt. Lett. 11, 288–290 (1986). This classic paper is the first experimental demonstration of the single-beam gradient trap, or optical tweezers.
Neuman, K.C., Chadd, E.H., Liou, G.F., Bergman, K. & Block, S.M. Characterization of photodamage to Escherichia coli in optical traps. Biophys. J. 77, 2856–2863 (1999).
Sacconi, L., Tolic-Norrelykke, I.M., Stringari, C., Antolini, R. & Pavone, F.S. Optical micromanipulations inside yeast cells. Appl. Opt. 44, 2001–2007 (2005).
Cherney, D.P., Bridges, T.E. & Harris, J.M. Optical trapping of unilamellar phospholipid vesicles: Investigation of the effect of optical forces on the lipid membrane shape by confocal-raman microscopy. Anal. Chem. 76, 4920–4928 (2004).
Lee, W.M., Reece, P.J., Marchington, R.F., Metzger, N.K. & Dholakia, K. Construction and calibration of an optical trap on a fluorescence optical microscope. Nat. Protocols 2, 3226–3238 (2007).
Neuman, K.C., Abbondanzieri, E.A. & Block, S.M. Measurement of the effective focal shift in an optical trap. Opt. Lett. 30, 1318–1320 (2005).
Fallman, E. & Axner, O. Influence of a glass-water interface on the on-axis trapping of micrometer-sized spherical objects by optical tweezers. Appl. Opt. 42, 3915–3926 (2003).
Viana, N.B., Rocha, M.S., Mesquita, O.N., Mazolli, A. & Neto, P.A.M. Characterization of objective transmittance for optical tweezers. Appl. Opt. 45, 4263–4269 (2006).
Misawa, H., Koshioka, M., Sasaki, K., Kitamura, N. & Masuhara, H. Three-dimensional optical trapping and laser ablation of a single polymer latex particle in water. J. Appl. Phys. 70, 3829–3836 (1991).
Rohrbach, A. & Stelzer, E.H.K. Three-dimensional position detection of optically trapped dielectric particles. J. Appl. Phys. 91, 5474–5488 (2002).
Gittes, F. & Schmidt, C.F. Interference model for back-focal-plane displacement detection in optical tweezers. Opt. Lett. 23, 7–9 (1998).
Peterman, E.J.G., van Dijk, M.A., Kapitein, L.C. & Schmidt, C.F. Extending the bandwidth of optical-tweezers interferometry. Rev. Sci. Instrum. 74, 3246–3249 (2003).
Greenleaf, W.J., Woodside, M.T., Abbondanzieri, E.A. & Block, S.M. Passive all-optical force clamp for high-resolution laser trapping. Phys. Rev. Lett. 95, 208102 (2005).
Carter, A.R., King, G.M. & Perkins, T.T. Back-scattered detection provides atomic-scale localization precision, stability, and registration in 3D. Opt. Express 15, 13434–13445 (2007).
Carter, A.R. et al. Stabilization of an optical microscope to 0.1 nm in three dimensions. Appl. Opt. 46, 421–427 (2007).
Vermeulen, K.C. et al. Calibrating bead displacements in optical tweezers using acousto-optic deflectors. Rev. Sci. Instrum. 77, 013704 (2006).
Tolic-Norrelykke, S.F. et al. Calibration of optical tweezers with positional detection in the back focal plane. Rev. Sci. Instrum. 77, 103101 (2006).
Svoboda, K., Schmidt, C.F., Schnapp, B.J. & Block, S.M. Direct observation of kinesin stepping by optical trapping interferometry. Nature 365, 721–727 (1993). The first direct measurement of the steps taken by individual kinesin.
Mammen, M. et al. Optically controlled collisions of biological objects to evaluate potent polyvalent inhibitors of virus-cell adhesion. Chem. Biol. 3, 757–763 (1996).
Kerssemakers, J.W.J. et al. Assembly dynamics of microtubules at molecular resolution. Nature 442, 709–712 (2006).
Wang, M.D. et al. Force and velocity measured for single molecules of RNA polymerase. Science 282, 902–907 (1998).
Herbert, K.M. et al. Sequence-resolved detecton of pausing by single RNA polymerase molecules. Cell 125, 1083–1094 (2006).
Neuman, K.C., Abbondanzieri, E.A., Landick, R., Gelles, J. & Block, S.M. Ubiquitous transcriptional pausing is independent of RNA polymerase backtracking. Cell 115, 437–447 (2003).
Shaevitz, J.W., Abbondanzieri, E.A., Landick, R. & Block, S.M. Backtracking by single RNA polymerase molecules observed at near-base-pair resolution. Nature 426, 684–687 (2003).
Moffitt, J.R., Chemla, Y.R., Izhaky, D. & Bustamante, C. Differential detection of dual traps improves the spatial resolution of optical tweezers. Proc. Natl. Acad. Sci. USA 103, 9006–9011 (2006).
Dame, R.T., Noom, M.C. & Wuite, G.J. Bacterial chromatin organization by H-NS protein unravelled using dual DNA manipulation. Nature 444, 387–390 (2006).
Smith, D.E. et al. The bacteriophage phi 29 portal motor can package DNA against a large internal force. Nature 413, 748–752 (2001).
Liphardt, J., Onoa, B., Smith, S.B., Tinoco, I. & Bustamante, C. Reversible unfolding of single RNA molecules by mechanical force. Science 292, 733–737 (2001). This paper was the first to show the mechanical unfolding of RNA structures, and it laid the groundwork for subsequent measurements of enzymatic unfolding of nucleic acid structures.
Dumont, S. et al. RNA translocation and unwinding mechanism of HCV NS3 helicase and its coordination by ATP. Nature 439, 105–108 (2006).
Friese, M.E.J., Nieminen, T.A., Heckenberg, N.R. & Rubinsztein-Dunlop, H. Optical alignment and spinning of laser-trapped microscopic particles. Nature 394, 348–350 (1998).
Deufel, C., Forth, S., Simmons, C.R., Dejgosha, S. & Wang, M.D. Nanofabricated quartz cylinders for angular trapping: DNA supercoiling torque detection. Nat. Methods 4, 223–225 (2007).
La Porta, A. & Wang, M.D. Optical torque wrench: angular trapping, rotation, and torque detection of quartz microparticles. Phys. Rev. Lett. 92, 190801 (2004).
Gross, S.P. Application of optical traps in vivo. Methods Enzymol. 361, 162–174 (2003).
Peterman, E.J.G., Gittes, F. & Schmidt, C.F. Laser-induced heating in optical traps. Biophys. J. 84, 1308–1316 (2003).
Abbondanzieri, E.A., Shaevitz, J.W. & Block, S.M. Picocalorimetry of transcription by RNA polymerase. Biophys. J. 89, L61–L63 (2005).
Seol, Y., Carpenter, A.E. & Perkins, T.T. Gold nanoparticles: enhanced optical trapping and sensitivity coupled with significant heating. Opt. Lett. 31, 2429–2431 (2006).
Liang, H. et al. Wavelength dependence of cell cloning efficiency after optical trapping. Biophys. J. 70, 1529–1533 (1996).
Strick, T., Allemand, J., Croquette, V. & Bensimon, D. Twisting and stretching single DNA molecules. Prog. Biophys. Mol. Biol. 74, 115–140 (2000).
Charvin, G., Strick, T.R., Bensimon, D. & Croquette, V. Tracking topoisomerase activity at the single-molecule level. Annu. Rev. Biophys. Biomol. Struct. 34, 201–219 (2005).
Strick, T.R., Croquette, V. & Bensimon, D. Single-molecule analysis of DNA uncoiling by a type II topoisomerase. Nature 404, 901–904 (2000). This was the first single-molecule measurement of topoisomerase activity using magnetic tweezers.
Gore, J. et al. Mechanochemical analysis of DNA gyrase using rotor bead tracking. Nature 439, 100–104 (2006).
Itoh, H. et al. Mechanically driven ATP synthesis by F-1-ATPase. Nature 427, 465–468 (2004).
Fisher, J.K. et al. Thin-foil magnetic force system for high-numerical-aperture microscopy. Rev. Sci. Instrum. 77, 023702 (2006).
Yan, J., Skoko, D. & Marko, J.F. Near-field-magnetic-tweezer manipulation of single DNA molecules. Phys. Rev. E 70, 011905 (2004).
Gosse, C. & Croquette, V. Magnetic tweezers: micromanipulation and force measurement at the molecular level. Biophys. J. 82, 3314–3329 (2002).
Lee, S.H. et al. Characterizing and tracking single colloidal particles with video holographic microscopy. Opt. Express 15, 18275–18282 (2007).
Lionnet, T., Spiering, M.M., Benkovic, S.J., Bensimon, D. & Croquette, V. Real-time observation of bacteriophage T4 gp41 helicase reveals an unwinding mechanism. Proc. Natl. Acad. Sci. USA 104, 19790–19795 (2007).
Strick, T.R., Allemand, J.F., Bensimon, D., Bensimon, A. & Croquette, V. The elasticity of a single supercoiled DNA molecule. Science 271, 1835–1837 (1996).
Strick, T.R., Allemand, J.F., Bensimon, D. & Croquette, V. Behavior of supercoiled DNA. Biophys. J. 74, 2016–2028 (1998).
Dekker, N.H. et al. The mechanism of type IA topoisomerases. Proc. Natl. Acad. Sci. USA 99, 12126–12131 (2002).
Charvin, G., Bensimon, D. & Croquette, V. Single-molecule study of DNA unlinking by eukaryotic and prokaryotic type-II topoisomerases. Proc. Natl. Acad. Sci. USA 100, 9820–9825 (2003).
Conroy, R.S. Force spectroscopy with optical and magnetic tweezers. in Handbook of Molecular Force Spectroscopy (ed. Noy, A.) 23–96 (Springer US, New York, 2008).
Bausch, A.R., Ziemann, F., Boulbitch, A.A., Jacobson, K. & Sackmann, E. Local measurements of viscoelastic parameters of adherent cell surfaces by magnetic bead microrheometry. Biophys. J. 75, 2038–2049 (1998).
de Vries, A.H., Krenn, B.E., van Driel, R. & Kanger, J.S. Micro magnetic tweezers for nanomanipulation inside live cells. Biophys. J. 88, 2137–2144 (2005).
Bausch, A.R., Moller, W. & Sackmann, E. Measurement of local viscoelasticity and forces in living cells by magnetic tweezers. Biophys. J. 76, 573–579 (1999).
Keller, M., Schilling, J. & Sackmann, E. Oscillatory magnetic bead rheometer for complex fluid microrheometry. Rev. Sci. Instrum. 72, 3626–3634 (2001).
Danilowicz, C., Greenfield, D. & Prentiss, M. Dissociation of ligand-receptor complexes using magnetic tweezers. Anal. Chem. 77, 3023–3028 (2005).
Koster, D.A., Croquette, V., Dekker, C., Shuman, S. & Dekker, N.H. Friction and torque govern the relaxation of DNA supercoils by eukaryotic topoisomerase IB. Nature 434, 671–674 (2005).
Koster, D.A., Palle, K., Bot, E.S., Bjornsti, M.A. & Dekker, N.H. Antitumour drugs impede DNA uncoiling by topoisomerase I. Nature 448, 213–217 (2007). The mechanism of action of a type I topoisomerase inhibitor measured at the single-molecule level was shown to be directly related to its effects in vivo.
Revyakin, A., Ebright, R.H. & Strick, T.R. Single-molecule DNA nanomanipulation: improved resolution through use of shorter DNA fragments. Nat. Methods 2, 127–138 (2005).
Revyakin, A., Liu, C., Ebright, R.H. & Strick, T.R. Abortive initiation and productive initiation by RNA polymerase involve DNA scrunching. Science 314, 1139–1143 (2006).
Binnig, G., Quate, C.F. & Gerber, C. Atomic force microscope. Phys. Rev. Lett. 56, 930–933 (1986).
Lee, G.U., Chrisey, L.A. & Colton, R.J. Direct measurement of the forces between complementary strands of DNA. Science 266, 771–773 (1994).
Dai, P. et al. X-ray-diffraction and scanning-tunneling-microscopy studies of a liquid-crystal film adsorbed on single-crystal graphite. Phys. Rev. B Condens. Matter 47, 7401–7407 (1993).
Binnig, G., Garcia, N. & Rohrer, H. Conductivity sensitivity of inelastic scanning tunneling microscopy. Phys. Rev. B Condens. Matter 32, 1336–1338 (1985).
Marti, O. et al. Scanning probe microscopy of biological samples and other surfaces. J. Microsc. 152, 803–809 (1988).
Engel, A. Biological applications of scanning probe microscopes. Annu. Rev. Biophys. Biophys. Chem. 20, 79–108 (1991).
Lindsay, S.M. Biological scanning probe microscopy comes of age. Biophys. J. 67, 2134–2135 (1994).
Bustamante, J.O., Liepins, A., Prendergast, R.A., Hanover, J.A. & Oberleithner, H. Patch clamp and atomic force microscopy demonstrate TATA-binding protein (TBP) interactions with the nuclear pore complex. J. Membr. Biol. 146, 263–272 (1995).
Shao, Z., Yang, J. & Somlyo, A.P. Biological atomic force microscopy: from microns to nanometers and beyond. Annu. Rev. Cell Dev. Biol. 11, 241–265 (1995).
Drake, B. et al. Imaging crystals, polymers, and processes in water with the atomic force microscope. Science 243, 1586–1589 (1989).
Bustamante, C., Rivetti, C. & Keller, D.J. Scanning force microscopy under aqueous solutions. Curr. Opin. Struct. Biol. 7, 709–716 (1997).
Rief, M., Oesterhelt, F., Heymann, B. & Gaub, H.E. Single molecule force spectroscopy on polysaccharides by atomic force microscopy. Science 275, 1295–1297 (1997).
Cumpson, P.J., Zhdan, P. & Hedley, J. Calibration of AFM cantilever stiffness: a microfabricated array of reflective springs. Ultramicroscopy 100, 241–251 (2004).
Sarkar, A., Robertson, R.B. & Fernandez, J.M. Simultaneous atomic force microscope and fluorescence measurements of protein unfolding using a calibrated evanescent wave. Proc. Natl. Acad. Sci. USA 101, 12882–12886 (2004).
Rief, M., Gautel, M., Oesterhelt, F., Fernandez, J.M. & Gaub, H.E. Reversible unfolding of individual titin immunoglobulin domains by AFM. Science 276, 1109–1112 (1997).
Leake, M.C., Wilson, D., Gautel, M. & Simmons, R.M. The elasticity of single titin molecules using a two-bead optical tweezers assay. Biophys. J. 87, 1112–1135 (2004).
Kellermayer, M.S. Delayed dissociation of in vitro moving actin filaments from heavy meromyosin induced by low concentrations of Triton X-100. Biophys. Chem. 67, 199–210 (1997).
Smith, S.B., Cui, Y. & Bustamante, C. Overstretching B-DNA: the elastic response of individual double-stranded and single-stranded DNA molecules. Science 271, 795–799 (1996).
Florin, E.L., Moy, V.T. & Gaub, H.E. Adhesion forces between individual ligand-receptor pairs. Science 264, 415–417 (1994).
Yasuda, R., Noji, H., Kinosita, K. Jr & Yoshida, M. F1-ATPase is a highly efficient molecular motor that rotates with discrete 120 degree steps. Cell 93, 1117–1124 (1998).
Rief, M., Gautel, M., Schemmel, A. & Gaub, H.E. The mechanical stability of immunoglobulin and fibronectin III domains in the muscle protein titin measured by atomic force microscopy. Biophys. J. 75, 3008–3014 (1998).
Baumgartner, W. et al. Cadherin interaction probed by atomic force microscopy. Proc. Natl. Acad. Sci. USA 97, 4005–4010 (2000).
Marszalek, P.E. et al. Mechanical unfolding intermediates in titin modules. Nature 402, 100–103 (1999).
Carrion-Vazquez, M. et al. Mechanical design of proteins studied by single-molecule force spectroscopy and protein engineering. Prog. Biophys. Mol. Biol. 74, 63–91 (2000).
Steward, A., Toca-Herrera, J.L. & Clarke, J. Versatile cloning system for construction of multimeric proteins for use in atomic force microscopy. Protein Sci. 11, 2179–2183 (2002).
Katz, E. & Willner, I. Biomolecule-functionalized carbon nanotubes: applications in nanobioelectronics. ChemPhysChem 5, 1084–1104 (2004).
Bustamante, C., Marko, J.F., Siggia, E.D. & Smith, S. Entropic elasticity of lambda-phage DNA. Science 265, 1599–1600 (1994). In this work the non-linear elasticity of DNA was measured and fit with an analytical expression. These results underpin the subsequent single-molecule force spectroscopy measurements of DNA and DNA processing enzymes.
Fernandez, J.M. & Li, H.B. Force-clamp spectroscopy monitors the folding trajectory of a single protein. Science 303, 1674–1678 (2004).
Schwaiger, I., Kardinal, A., Schleicher, M., Noegel, A.A. & Rief, M. A mechanical unfolding intermediate in an actin-crosslinking protein. Nat. Struct. Mol. Biol. 11, 81–85 (2004).
Carrion-Vazquez, M. et al. The mechanical stability of ubiquitin is linkage dependent. Nat. Struct. Biol. 10, 738–743 (2003).
Dietz, H., Berkemeier, F., Bertz, M. & Rief, M. Anisotropic deformation response of single protein molecules. Proc. Natl. Acad. Sci. USA 103, 12724–12728 (2006).
Wright, C.F., Lindorff-Larsen, K., Randles, L.G. & Clarke, J. Parallel protein-unfolding pathways revealed and mapped. Nat. Struct. Biol. 10, 658–662 (2003).
Wiita, A.P. et al. Probing the chemistry of thioredoxin catalysis with force. Nature 450, 124–127 (2007).
Szoszkiewicz, R. et al. Dwell time analysis of a single-molecule mechanochemical reaction. Langmuir 24, 1356–1364 (2008).
Greene, D. et al. Single-molecule force spectroscopy reveals a stepwise unfolding of C. elegans giant protein kinase domains. Biophys. J. (in the press) (2008).
Brown, A.E., Litvinov, R.I., Discher, D.E. & Weisel, J.W. Forced unfolding of coiled-coils in fibrinogen by single-molecule AFM. Biophys. J. 92, L39–L41 (2007).
Kellermayer, M.S. et al. Reversible mechanical unzipping of amyloid beta-fibrils. J. Biol. Chem. 280, 8464–8470 (2005).
Oesterhelt, F. et al. Unfolding pathways of individual bacteriorhodopsins. Science 288, 143–146 (2000).
Ando, T. et al. A high-speed atomic force microscope for studying biological macromolecules. Proc. Natl. Acad. Sci. USA 98, 12468–12472 (2001).
Wen, J.D. et al. Following translation by single ribosomes one codon at a time. Nature 452, 598–603 (2008).
Tanner, N.A. et al. Single-molecule studies of fork dynamics in Escherichia coli DNA replication. Nat. Struct. Mol. Biol. 15, 170–176 (2008).
Hamdan, S.M. et al. Dynamic DNA helicase-DNA polymerase interactions assure processive replication fork movement. Mol. Cell 27, 539–549 (2007).
van Oijen, A.M. Single-molecule studies of complex systems: the replisome. Mol. Biosyst. 3, 117–125 (2007).
Lee, J.B. et al. DNA primase acts as a molecular brake in DNA replication. Nature 439, 621–624 (2006).
Ishijima, A. et al. Simultaneous observation of individual ATPase and mechanical events by a single myosin molecule during interaction with actin. Cell 92, 161–171 (1998).
Lang, M.J., Fordyce, P.M., Engh, A.M., Neuman, K.C. & Block, S.M. Simultaneous, coincident optical trapping and single-molecule fluorescence. Nat. Methods 1, 133–139 (2004).
Li, P.T., Bustamante, C. & Tinoco, I. Jr. Real-time control of the energy landscape by force directs the folding of RNA molecules. Proc. Natl. Acad. Sci. USA 104, 7039–7044 (2007).
Chiou, P.Y., Ohta, A.T. & Wu, M.C. Massively parallel manipulation of single cells and microparticles using optical images. Nature 436, 370–372 (2005).
Cohen, A.E. & Moerner, W.E. Suppressing Brownian motion of individual biomolecules in solution. Proc. Natl. Acad. Sci. USA 103, 4362–4365 (2006).
Berg-Sorensen, K. & Flyvbjerg, H. Power spectrum analysis for optical tweezers. Rev. Sci. Instrum. 75, 594–612 (2004). This is an invaluable reference for the treatment and fitting of Brownian noise and power spectra in single-molecule force spectroscopy measurements.
Acknowledgements
K.C.N. and A.N. are supported by the Intramural Program of the National Heart, Lung, and Blood Institute, National Institutes of Health. We thank G. Liou, R. Neuman and Y. Takagi for critical reading of the manuscript. K.C.N. acknowledges D. Bensimon, V. Croquette and S. Block, in addition to members of their labs.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Neuman, K., Nagy, A. Single-molecule force spectroscopy: optical tweezers, magnetic tweezers and atomic force microscopy. Nat Methods 5, 491–505 (2008). https://doi.org/10.1038/nmeth.1218
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1218
This article is cited by
-
Modelling and targeting mechanical forces in organ fibrosis
Nature Reviews Bioengineering (2024)
-
Polymeric protagonists for biological processes
Nature Chemistry (2023)
-
Phosphorescent extensophores expose elastic nonuniformity in polymer networks
Nature Communications (2023)
-
An external speed control for nanopore reads
Nature Nanotechnology (2023)
-
Interrogating imaginary optical force by the complex Maxwell stress tensor theorem
Light: Science & Applications (2023)