Abstract
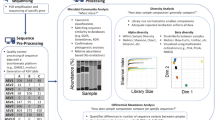
We constructed error-correcting DNA barcodes that allow one run of a massively parallel pyrosequencer to process up to 1,544 samples simultaneously. Using these barcodes we processed bacterial 16S rRNA gene sequences representing microbial communities in 286 environmental samples, corrected 92% of sample assignment errors, and thus characterized nearly as many 16S rRNA genes as have been sequenced to date by Sanger sequencing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Margulies, M. et al. Nature 437, 376–380 (2005).
Sogin, M.L. et al. Proc. Natl. Acad. Sci. USA 103, 12115–12120 (2006).
Huber, J.A. et al. Science 318, 97–100 (2007).
Roesch, L.F.W. et al. ISME J. 1, 283–290 (2007).
Pace, N.R. Science 276, 734–740 (1997).
Binladen, J. et al. PLoS ONE 2, e197 (2007).
Hoffmann, C. et al. Nucleic Acids Res. 35, e91 (2007).
Parameswaran, P. et al. Nucleic Acids Res. 35, e130 (2007).
Huse, S.M., Huber, J.A., Morrison, H.G., Sogin, M.L. & Welch, D.M. Genome Biol. 8, R143 (2007).
Morelos-Zaragoza, R.H. The Art of Error-Correcting Coding (John Wiley & Sons, Hoboken, New Jersey, 2006).
Liu, Z., Lozupone, C., Hamady, M., Bushman, F.D. & Knight, R. Nucleic Acids Res. 35, e120 (2007).
Dojka, M.A., Hugenholtz, P., Haack, S.K. & Pace, N.R. Appl. Environ. Microbiol. 64, 3869–3877 (1998).
Lozupone, C.A. & Knight, R. Proc. Natl. Acad. Sci. USA 104, 11436–11440 (2007).
Lozupone, C., Hamady, M. & Knight, R. BMC Bioinformatics 7, 371 (2006).
Lozupone, C. & Knight, R. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Acknowledgements
We thank N. Pace, L. Gold and F. Accurso for support and encouragement, J.I. Gordon and R. Bushman for helpful discussions, and R. Ley, C. Lozupone and D. McDonald for feedback on the manuscript. This work was supported in part by the US National Institutes of Health–University of Colorado at Boulder Molecular Biophysics Training Program (T32GM065103), and grants from the Cystic Fibrosis Foundation and National Institutes of Health (U01 HL081335-01, P01DK078669).
Author information
Authors and Affiliations
Contributions
M.H. and R.K. designed and implemented the analyses, and wrote the manuscript. J.J.W., J.K.H. and N.J.G. generated the 454 dataset.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1, Supplementary Data 1, Supplementary Methods (PDF 422 kb)
Supplementary Data 2
Decoding software example, Readme file and demo output (PDF 67 kb)
Rights and permissions
About this article
Cite this article
Hamady, M., Walker, J., Harris, J. et al. Error-correcting barcoded primers for pyrosequencing hundreds of samples in multiplex. Nat Methods 5, 235–237 (2008). https://doi.org/10.1038/nmeth.1184
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1184
This article is cited by
-
Core species and interactions prominent in fish-associated microbiome dynamics
Microbiome (2023)
-
Dynamics of species-rich predator–prey networks and seasonal alternations of core species
Nature Ecology & Evolution (2023)
-
Discovering covalent inhibitors of protein–protein interactions from trillions of sulfur(VI) fluoride exchange-modified oligonucleotides
Nature Chemistry (2023)
-
Associations of gut microbiome with endogenous estrogen levels in healthy postmenopausal women
Cancer Causes & Control (2023)
-
Vertical transmission of cellulolytic protists in termites is imperfect, but sufficient, due to biparental transmission
Symbiosis (2023)