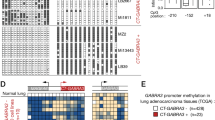
Abstract
For several human tumour types, allelic toss data suggest that one or more tumour suppressor genes reside telomeric to the p53 gene at chromosome 17p13.1. In the present study we have used a new strategy, involving molecular analysis of a DNA site hypermethylated in tumour DNA, to identify a candidate gene in this region (17p13.3). Our approach has led to identification of HIC-1 (hypermethylated in cancer), a new zinc-finger transcription factor gene which is ubiquitously expressed in normal tissues, but underexpressed in different tumour cells where it is hypermethylated. Multiple characteristics of this gene, including the presence of a p53 binding site in the 5′ flanking region, activation of the gene by expression of a wild-type p53 gene and suppression of C418 selectability of cultured brain, breast and colon cancer cells following insertion of the gene, make HIC-1 gene a strong candidate for a tumour suppressor gene in region 17p13.3.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Vogelstein, B. & Kinzler, K.W. . p53 function and dysfunction. Cell 70, 523–526 (1992).
Chen, L.-C. et al. Loss of heterozygosity on the short arm of chromosome 17 is associated with high proliferative capacity and DNA aneuploidy in primary human breast cancer. Proc. natn. Acad. Sci. U.S.A. 88, 3847–3851 (1991).
Takita, K.-I. et al. Correlation of loss of alleles on the short arm of chromosomes 11 and 17 with metastasis of primary breast cancer to lymph nodes. Cancer Res. 52, 3914–3917 (1992).
Deng, G. et al. Loss of heterozygosity and p53 gene mutations in breast cancer. Cancer Res. 54, 499–505 (1994).
Cornelis, R.S. et al. Evidence for a gene on 17p13.3, distal to TP53, as a target for allele loss in breast tumors without p53 mutations. Cancer Res. 54, 4200–4206 (1994).
Coles, C. et al. Evidence implicating at least two genes on chromosome 17p in breast carcinogenesis. Lancet 336, 761–763 (1990).
Makos, M. et al. Distinct hypeimethylation patterns occur at altered chromosome loci in human lung and colon cancer. Proc. natn. Acad. Sci. U.S.A. 89, 1929–1933 (1992).
Makos, M. et al. DNA hypermethylation is associated with 17p allelic loss in neural tumors. Cancer Res. 53, 2715–2718 (1993).
Makos, M. et al. Regional DNA hypermethylation at D17S5 precedes 17p structural changes in the progression of renal tumors. Cancer Res. 53, 2719–2722 (1993).
Baylin, S.B. et al. Abnormal patterns of DNA methylation in human neoplasia: Potential consequences fortumorprogression. Cancer Cells 3, 383–390 (1991).
Jones, P.A. & Buckley, J.D. The role of DNA methylation in cancer. Adv. Cancer Res. 54, 1–23 (1990).
Herman, J.G. et al. Silencing of the VHL tumor suppressor gene by DNA methylation in renal carcinoma. Proc. natn. Acad. Sci. 91, 9700–9704 (1994).
Ottaviano, Y.L. et al. Methylation of the estrogen receptor gene CpG island marks loss of estrogen receptor expression in human breast cancer cells. Cancer Res. 54, 2552–2555 (1994).
Issa, J.-P.J. et al. Methylation of the estrogen receptor CpG island links aging and neoplasia in human colon. Nature Genet. 7, 536–540 (1994).
Steenman, M.J.C. et al. Loss of imprinting of IGF2 is linked to reduced expression and abnormal methylation of H19 in Wilms' tumour. Nature Genet. 7, 433–439 (1994).
Ledbetter, D.H. et al. Molecular dissection of a contiguous gene syndrome: Frequent submicroscopic deletions, evolutionarily conserved sequences, and a hypomethylated “island” in the Miller-Dicker chromosome region. Proc. natn. Acad. Sci. U.S.A. 86, 5136–5140 (1989).
Gish, W. & States, D.J. Identification of protein coding regions by database similarity search. Nature Genet. 3, 266–272 (1993).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. molec. Biol. 215, 403–410 (1990).
Numoto, M. et al. Transcriptional represser ZF5 identifies a new conserved domain in zinc finger proteins. Nucleic Acids Res. 21, 3767–3775 (1993).
Harrison, S.D. & Travers, A.A. The tramtrack gene encodes a drosophila finger protein that interacts with the ftz transcriptional regulatory region and shows a novel embryonic expression pattern. EMBO J. 9, 207–216 (1990).
DiBello, P.R., Withers, D.A., Bayer, C.A., Fristrom, J.W. & Guild, G.M. The drosophila broad-complex encodes a family of related proteins containing zinc fingers. Genetics 129, 385–397 (1991).
Chardin, P., Courtois, G., Mattei, M.-G. & Gisselbrecht, S. The KUP gene, located on human chromosome 14, encodes a protein with two distant zinc fingers. Nucleic Acids Res. 19, 1431–1436 (1991).
Hromas, R. et al. A retinoic acid-responsive human zinc finger gene, MZF-1, preferentially expressed in myeloid cells. J. biol. Chem. 266, 14183–14187 (1991).
Chen, Z. et al. Fusion between a novel Kruppel-like zinc finger gene and the retinoic acid receptor-α locus due to a variant t(11;17) translocation associated with acute promyelocytic leukaemia. EMBO J. 12, 1161–11671 (1993).
Kerckaert, J.-P., Deweindt, C., Tilly, H., Quief, S., Lecocq, G. & Bastard, C. LAZ3, a novel zinc-finger encoding gene, is disrupted by recurring chromosome 3q27 translocations in human lymphomas. Nature Genet. 5, 66–70 (1993).
Ye, B.H. et al. Alterations of a zinc finger-encoding gene, BCL-6, in diffuse large-cell lymphoma. Science 262, 747–750 (1993).
Soeller, W.C., Oh, C.E. & Kornberg, T.B. Isolation of cDNAs encoding the drosophila GAGA transcription factor. Molec. cell. Biol. 13, 7961–7970 (1993).
Ruppert, J.M. et al. The GLI-Kruppel family of human genes. Molec. cell. Biol. 8, 3104–3113 (1988).
El-Deiry, W.S., Kern, S.E., Pietenpol, J.A., Kinzler, K.W. & Vogelstein, B. Definition of a consensus binding site for p53. Nature Genet. 1, 45–49 (1992).
Kern, S.E. et al. Identification of p53 as a sequence-specific DNA-binding protein. Science 252, 1708–1711 (1991).
Funk, W.D., Pak, D.T., Karas, R.H., Wright, W.E. & Shay, J.W. A transcriptionally active DNA-binding site for human p53 protein complexes. Molec. cell. Biol. 12, 2866–2871 (1992).
El-Deiry, W.S. et al. WAF1, a potential mediator of p53 tumor suppression. Cell 75, 817–825 (1993).
Baker, S.J., Markowitz, S., Fearon, E.R., Willson, J.K.V. & Vogelstein, B. Suppression of human colorectal carcinoma cell growth by wild-type p53. Science 249, 912–915 (1990).
Riggs, A.D. & Pfeifer, G.D. X-chromosome inactivation and cell memory. Trends Genet. 8, 169–174 (1992).
Li, E., Beard, C. & Jaenisch, R. Role for DNA methylation in genomic imprinting. Nature 366, 362–365 (1993).
Yisraeli, J. et al. Muscle specific activation of a methylated chimeric actin gene. Cell 46, 409–416 (1986).
Runnebaum, I.B., Nagarajan, M., Bowman, M., Soto, D. & Sukumar, S. Mutations in p53 as potential molecular markers for human breast cancer. Proc. natn. Acad. Sci. U.S.A. 88, 10657–10661 (1991).
Negrini, M. et al. Tumor and growth suppression of breast cancer cells by chromosome 17-associated functions. Cancer Res. 54, 1818–1824 (1994).
Chen, P., Ellmore, N. & Weissman, B.E. Functional evidence for a second tumor suppressor gene on human chromosome 17. Molec. cell. Biol. 14, 534–542 (1994).
Hensel, C.H., Xiang, R.H., Sakaguchi, A.Y. & Naylor, S.L. Use of the single strand conformation polymorphism technique and PCR to detect p53 gene mutations in small cell lung cancer. Oncogene 6, 1067–1071 (1991).
Baker, S.J. et al. p53 gene mutations occur in combination with 17p allelic deletions as late events in colorectal tumorigenesis. Cancer Res. 50, 7717–7722 (1990).
Van Meir, E.G. et al. Analysis of the p53 gene and its expression in human glioblastoma cells. Cancer Res. 54, 649–652 (1994).
Chen, C.Y. et al. Interactions between p53 and MDM2 in a mammalian cell cycle checkpoint pathway. Proc. natn. Acad. Sci. U.S.A. 91, 2684–2688 (1994).
Kastan, M.B. et al. A mammalian cell cycle checkpoint pathway utilizing p53 and GADD45 is defective in ataxia-telangiectasia. Cell 71, 587–597 (1992).
Miyashita, T. et al. Tumor suppressor p53 is a regulator of bcl-2 and bax gene expression in vitro and in vivo. Oncogene 9, 1799–1805 (1994).
Tsukiyama, T., Becker, P.B. & Wu, C. ATP-dependent nucleosome disruption at a heat-shock promoter mediated by binding of GAGA transcription factor. Nature 367, 525–532 (1994).
Bardwell, V.J. & Treisman, R. The POZ domain: A conserved protein-protein interaction motif. Genes Dev. 8, 1664–1677 (1994).
Miki, T., Kawamata, N., Hirosawa, S. & Aoki, N. Gene involved in the 3q27 translocation associated with B-cell lymphoma, BCL5, encodes a Kruppel-like zinc-finger protein. Blood 83, 26–32 (1994).
Carney, D.N., Bepler, G. & Gazdar, A.F. The serum-free establishment and in vitro growth properties of classic and variant small cell lung cancer cell lines. Recent Results Cancer Res. 99, 157–166 (1985).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wales, M., Biel, M., Deiry, W. et al. p53 activates expression of HIC-1, a new candidate tumour suppressor gene on 17p13.3. Nat Med 1, 570–577 (1995). https://doi.org/10.1038/nm0695-570
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm0695-570
This article is cited by
-
COX-2/PGE2 upregulation contributes to the chromosome 17p-deleted lymphoma
Oncogenesis (2023)
-
Cell diversity and plasticity during atrioventricular heart valve EMTs
Nature Communications (2023)
-
Cellular taxonomy of Hic1+ mesenchymal progenitor derivatives in the limb: from embryo to adult
Nature Communications (2022)
-
FBXW11 contributes to stem-cell-like features and liver metastasis through regulating HIC1-mediated SIRT1 transcription in colorectal cancer
Cell Death & Disease (2021)
-
Proline oxidase silencing inhibits p53-dependent apoptosis in MCF-7 breast cancer cells
Amino Acids (2021)