Abstract
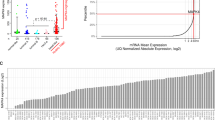
Triple-negative breast cancer (TNBC) is an aggressive subtype of breast cancer diagnosed in more than 200,000 women each year1 and is recalcitrant to targeted therapies2,3. Although TNBCs harbor multiple hyperactive receptor tyrosine kinases (RTKs)4,5,6,7,8, RTK inhibitors have been largely ineffective in TNBC patients thus far. We developed a broadly effective therapeutic strategy for TNBC that is based on combined inhibition of receptors that share the negative regulator PTPN12. Previously, we and others identified the tyrosine phosphatase PTPN12 as a tumor suppressor that is frequently inactivated in TNBC9,10. PTPN12 restrains several RTKs9,11,12,13,14,15,16,17, suggesting that PTPN12 deficiency leads to aberrant activation of multiple RTKs and a co-dependency on these receptors. This in turn leads to the therapeutic hypothesis that PTPN12-deficient TNBCs may be responsive to combined RTK inhibition. However, the repertoire of RTKs that are restrained by PTPN12 in human cells has not been systematically explored. By methodically identifying the suite of RTK substrates (MET, PDGFRβ, EGFR, and others) inhibited by PTPN12, we rationalized a combination RTK-inhibitor therapy that induced potent tumor regression across heterogeneous models of TNBC. Orthogonal approaches revealed that PTPN12 was recruited to and inhibited these receptors after ligand stimulation, thereby serving as a feedback mechanism to limit receptor signaling. Cancer-associated mutation of PTPN12 or reduced PTPN12 protein levels diminished this feedback mechanism, leading to aberrant activity of these receptors. Restoring PTPN12 protein levels restrained signaling from RTKs, including PDGFRβ and MET, and impaired TNBC survival. In contrast with single agents, combined inhibitors targeting the PDGFRβ and MET receptors induced the apoptosis in TNBC cells in vitro and in vivo. This therapeutic strategy resulted in tumor regressions in chemo-refractory patient-derived TNBC models. Notably, response correlated with PTPN12 deficiency, suggesting that impaired receptor feedback may establish a combined addiction to these proto-oncogenic receptors. Taken together, our data provide a rationale for combining RTK inhibitors in TNBC and other malignancies that lack receptor-activating mutations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Anders, C. & Carey, L.A. Understanding and treating triple-negative breast cancer. Oncology (Williston Park) 22, 1233–1239; discussion 1239–1240, 1243 (2008).
Anders, C.K., Zagar, T.M. & Carey, L.A. The management of early-stage and metastatic triple-negative breast cancer: a review. Hematol. Oncol. Clin. North. Am. 27, 737–749 viii (2013).
Bianchini, G., Balko, J.M., Mayer, I.A., Sanders, M.E. & Gianni, L. Triple-negative breast cancer: challenges and opportunities of a heterogeneous disease. Nat. Rev. Clin. Oncol. 13, 674–690 (2016).
Lehmann, B.D. et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Invest. 121, 2750–2767 (2011).
Hochgräfe, F. et al. Tyrosine phosphorylation profiling reveals the signaling network characteristics of Basal breast cancer cells. Cancer Res. 70, 9391–9401 (2010).
Duncan, J.S. et al. Dynamic reprogramming of the kinome in response to targeted MEK inhibition in triple-negative breast cancer. Cell 149, 307–321 (2012).
Nielsen, T.O. et al. Immunohistochemical and clinical characterization of the basal-like subtype of invasive breast carcinoma. Clin Cancer Res. 10, 5367–5374 (2004).
Mertins, P. et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 534, 55–62 (2016).
Sun, T. et al. Activation of multiple proto-oncogenic tyrosine kinases in breast cancer via loss of the PTPN12 phosphatase. Cell 144, 703–718 (2011).
Wu, M.Q. et al. Low expression of tyrosine-protein phosphatase nonreceptor type 12 is associated with lymph node metastasis and poor prognosis in operable triple-negative breast cancer. Asian Pac. J. Cancer Prev. 14, 287–292 (2013).
Villa-Moruzzi, E. PTPN12 controls PTEN and the AKT signalling to FAK and HER2 in migrating ovarian cancer cells. Mol. Cell. Biochem. 375, 151–157 (2013).
Zheng, Y. et al. Temporal regulation of EGF signalling networks by the scaffold protein Shc1. Nature 499, 166–171 (2013).
Charest, A., Wagner, J., Kwan, M. & Tremblay, M.L. Coupling of the murine protein tyrosine phosphatase PEST to the epidermal growth factor (EGF) receptor through a Src homology 3 (SH3) domain-mediated association with Grb2. Oncogene 14, 1643–1651 (1997).
Markova, B., Herrlich, P., Rönnstrand, L. & Böhmer, F.D. Identification of protein tyrosine phosphatases associating with the PDGF receptor. Biochemistry 42, 2691–2699 (2003).
Ambjørn, M. et al. A loss-of-function screen for phosphatases that regulate neurite outgrowth identifies PTPN12 as a negative regulator of TrkB tyrosine phosphorylation. PLoS One 8, e65371 (2013).
Li, H. et al. Crystal structure and substrate specificity of PTPN12. Cell Rep. 15, 1345–1358 (2016).
Barr, A.J. et al. Large-scale structural analysis of the classical human protein tyrosine phosphatome. Cell 136, 352–363 (2009).
Li, J. et al. Loss of PTPN12 stimulates progression of ErbB2-dependent breast cancer by enhancing cell survival, migration, and epithelial-to-mesenchymal transition. Mol. Cell. Biol. 35, 4069–4082 (2015).
Yao, Z. et al. A global analysis of the receptor tyrosine kinase-protein phosphatase interactome. Mol. Cell 65, 347–360 (2017).
Hoadley, K.A. et al. EGFR associated expression profiles vary with breast tumor subtype. BMC Genomics 8, 258 (2007).
Prat, A. et al. Molecular characterization of basal-like and non-basal-like triple-negative breast cancer. Oncologist 18, 123–133 (2013).
Zagouri, F. et al. High MET expression is an adverse prognostic factor in patients with triple-negative breast cancer. Br. J. Cancer 108, 1100–1105 (2013).
Cancer Genome Atlas, N.. Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Forbes, S.A. et al. The catalogue of somatic mutations in cancer (COSMIC). Curr. Protoc. Hum. Genet. Chapter 10, Unit 10 11 (2008).
Katsonis, P. & Lichtarge, O. A formal perturbation equation between genotype and phenotype determines the evolutionary action of protein-coding variations on fitness. Genome Res. 24, 2050–2058 (2014).
Carey, L.A. et al. TBCRC 001: randomized phase II study of cetuximab in combination with carboplatin in stage IV triple-negative breast cancer. J. Clin. Oncol. 30, 2615–2623 (2012).
Miller, K. et al. Paclitaxel plus bevacizumab versus paclitaxel alone for metastatic breast cancer. N. Engl. J. Med. 357, 2666–2676 (2007).
Yardley, D.A. et al. Phase I/II trial of neoadjuvant sunitinib administered with weekly paclitaxel/carboplatin in patients with locally advanced triple-negative breast cancer. Breast Cancer Res. Treat. 152, 557–567 (2015).
Bianchi, G. et al. Phase II multicenter, uncontrolled trial of sorafenib in patients with metastatic breast cancer. Anticancer Drugs 20, 616–624 (2009).
Finn, R.S. et al. Estrogen receptor, progesterone receptor, human epidermal growth factor receptor 2 (HER2), and epidermal growth factor receptor expression and benefit from lapatinib in a randomized trial of paclitaxel with lapatinib or placebo as first-line treatment in HER2-negative or unknown metastatic breast cancer. J. Clin. Oncol. 27, 3908–3915 (2009).
Prenen, H. et al. Efficacy of the kinase inhibitor SU11248 against gastrointestinal stromal tumor mutants refractory to imatinib mesylate. Clin. Cancer Res. 12, 2622–2627 (2006).
Guida, T. et al. Sorafenib inhibits imatinib-resistant KIT and platelet-derived growth factor receptor beta gatekeeper mutants. Clin. Cancer Res. 13, 3363–3369 (2007).
Qi, J. et al. Multiple mutations and bypass mechanisms can contribute to development of acquired resistance to MET inhibitors. Cancer Res. 71, 1081–1091 (2011).
Sørlie, T. et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl. Acad. Sci. USA 98, 10869–10874 (2001).
Zhang, X. et al. A renewable tissue resource of phenotypically stable, biologically and ethnically diverse, patient-derived human breast cancer xenograft models. Cancer Res. 73, 4885–4897 (2013).
Xunyi, Y. et al. Clinicopathological significance of PTPN12 expression in human breast cancer. Braz. J. Med. Biol. Res. 45, 1334–1340 (2012).
Wu, P., Nielsen, T.E. & Clausen, M.H. Small-molecule kinase inhibitors: an analysis of FDA-approved drugs. Drug Discov. Today 21, 5–10 (2016).
Su, Z. et al. PTPN12 inhibits oral squamous epithelial carcinoma cell proliferation and invasion and can be used as a prognostic marker. Med. Oncol. 30, 618 (2013).
Cao, X. et al. Tyrosine-protein phosphatase non-receptor type 12 expression is a good prognostic factor in resectable non-small cell lung cancer. Oncotarget 6, 11704–11713 (2015).
Cao, X. et al. Tyrosine-protein phosphatase nonreceptor type 12 is a novel prognostic biomarker for esophageal squamous cell carcinoma. Ann. Thorac. Surg. 93, 1674–1680 (2012).
Luo, R.Z. et al. Decreased expression of PTPN12 correlates with tumor recurrence and poor survival of patients with hepatocellular carcinoma. PLoS One 9, e85592 (2014).
Zhang, X.K. et al. The prognostic significance of tyrosine-protein phosphatase nonreceptor type 12 expression in nasopharyngeal carcinoma. Tumour Biol. 36, 5201–5208 (2015).
Meerbrey, K.L. et al. The pINDUCER lentiviral toolkit for inducible RNA interference in vitro and in vivo. Proc. Natl. Acad. Sci. USA 108, 3665–3670 (2011).
Lichtarge, O., Bourne, H.R. & Cohen, F.E. An evolutionary trace method defines binding surfaces common to protein families. J. Mol. Biol. 257, 342–358 (1996).
Mihalek, I., Res, I. & Lichtarge, O. A family of evolution-entropy hybrid methods for ranking protein residues by importance. J. Mol. Biol. 336, 1265–1282 (2004).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Mihalek, I., Res, I., Yao, H. & Lichtarge, O. Combining inference from evolution and geometric probability in protein structure evaluation. J. Mol. Biol. 331, 263–279 (2003).
Baameur, F. et al. Role for the regulator of G-protein signaling homology domain of G protein-coupled receptor kinases 5 and 6 in beta 2-adrenergic receptor and rhodopsin phosphorylation. Mol. Pharmacol. 77, 405–415 (2010).
Lichtarge, O., Yamamoto, K.R. & Cohen, F.E. Identification of functional surfaces of the zinc binding domains of intracellular receptors. J. Mol. Biol. 274, 325–337 (1997).
Ribes-Zamora, A., Mihalek, I., Lichtarge, O. & Bertuch, A.A. Distinct faces of the Ku heterodimer mediate DNA repair and telomeric functions. Nat. Struct. Mol. Biol. 14, 301–307 (2007).
Lua, R.C. & Lichtarge, O. PyETV: a PyMOL evolutionary trace viewer to analyze functional site predictions in protein complexes. Bioinformatics 26, 2981–2982 (2010).
Söderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006).
Acknowledgements
We would like to thank members of T.F.W. laboratory for helpful comments. The authors also acknowledge the joint participation of the Adrienne Helis Melvin Medical Research Foundation through its direct engagement in the continuous active conduct of medical research in conjunction with Baylor College of Medicine for cancer research. The Dan L. Duncan Cancer Center Shared Resources was supported by the NCI P30CA125123 Center Grant and provided technical assistance including Cell-Based Assay Screening Service (D. Liu), Biostatistics & Informatics Shared Resource (S. Hilsenbeck), and Cytometry and Cell Sorting (J. Sederstrom; P30 AI036211 and S10 RR024574). M.T.L. was supported by the Susan G. Komen Foundation (KG120001), a NCI/NIH SPORE Supplement Award (P50 CA186784), and the Patient-derived Xenograft and Advance In vivo Models Core at Baylor College of Medicine and P30 Cancer Center Support Grant (NCI-CA125123). T.F.W. was supported by CPRIT (RP120583), the Susan G. Komen Foundation (KG090355), the NIH (1R01CA178039-01) and the DOD Breast Cancer Research Program (BC120604).
Author information
Authors and Affiliations
Contributions
A.N., H.-C.C., T.S., S.T., L.E.D., R.D.-V., S.J.K., M.O., D.W.C., S.M., I.P. and E.S. performed the experiments. A.R., R.D.-V. and D.M.H. performed statistical analyses. R.D.-V. and P.K. performed the analysis shown in Figure 2a. P.K. performed the analysis shown in Figure 2b,c. H.L. performed the experiments shown in Figure 2d and Supplementary Figure 6a. C.J.C. performed the TCGA analysis shown in Figure 1e. J.D. and F.S. performed the image analysis shown in Figure 1h,i. A.N., C.G., J.W.T., O.L., C.Y.L., B.Z., K.L.S., S.G.H., J.S., X.Y., C.K.O., R.S., J.G.C., D.J.S., M.F.R., M.J.E., C.A.S., M.T.L. and T.F.W. devised or supervised experiments. A.N. and T.F.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.J.S. is an employee of Pfizer. J.G.C. is an employee of Mirati Therapeutics and former employee of Pfizer.
Supplementary information
Supplementary Figures & Tables
Supplementary Figures 1–14 & Supplementary Tables 1–2 (PDF 1534 kb)
Rights and permissions
About this article
Cite this article
Nair, A., Chung, HC., Sun, T. et al. Combinatorial inhibition of PTPN12-regulated receptors leads to a broadly effective therapeutic strategy in triple-negative breast cancer. Nat Med 24, 505–511 (2018). https://doi.org/10.1038/nm.4507
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4507
This article is cited by
-
HER3 functions as an effective therapeutic target in triple negative breast cancer to potentiate the antitumor activity of gefitinib and paclitaxel
Cancer Cell International (2023)
-
Syntenin-1-mediated small extracellular vesicles promotes cell growth, migration, and angiogenesis by increasing onco-miRNAs secretion in lung cancer cells
Cell Death & Disease (2022)
-
In Vivo Modeling of Human Breast Cancer Using Cell Line and Patient-Derived Xenografts
Journal of Mammary Gland Biology and Neoplasia (2022)
-
Protein tyrosine phosphatase non-receptor type 12 (PTPN12), negatively regulated by miR-106a-5p, suppresses the progression of hepatocellular carcinoma
Human Cell (2022)
-
Genome interpretation using in silico predictors of variant impact
Human Genetics (2022)