Abstract
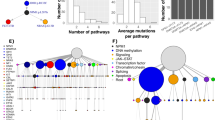
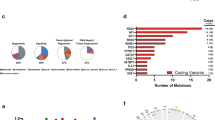
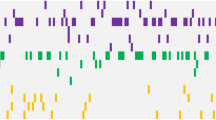
We present the molecular landscape of pediatric acute myeloid leukemia (AML) and characterize nearly 1,000 participants in Children's Oncology Group (COG) AML trials. The COG–National Cancer Institute (NCI) TARGET AML initiative assessed cases by whole-genome, targeted DNA, mRNA and microRNA sequencing and CpG methylation profiling. Validated DNA variants corresponded to diverse, infrequent mutations, with fewer than 40 genes mutated in >2% of cases. In contrast, somatic structural variants, including new gene fusions and focal deletions of MBNL1, ZEB2 and ELF1, were disproportionately prevalent in young individuals as compared to adults. Conversely, mutations in DNMT3A and TP53, which were common in adults, were conspicuously absent from virtually all pediatric cases. New mutations in GATA2, FLT3 and CBL and recurrent mutations in MYC-ITD, NRAS, KRAS and WT1 were frequent in pediatric AML. Deletions, mutations and promoter DNA hypermethylation convergently impacted Wnt signaling, Polycomb repression, innate immune cell interactions and a cluster of zinc finger–encoding genes associated with KMT2A rearrangements. These results highlight the need for and facilitate the development of age-tailored targeted therapies for the treatment of pediatric AML.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
01 January 2018
In the version of this article initially published online, Figure 1a has two black boxes in the key that are labeled as 'Unknown'; these boxes should be white, matching the segments in the donut charts shown below the key. The error has been corrected in the print, PDF and HTML versions of this article.
References
Steliarova-Foucher, E. et al. International incidence of childhood cancer, 2001–10: a population-based registry study. Lancet Oncol. 18, 719–731 (2017).
Li, S. et al. Distinct evolution and dynamics of epigenetic and genetic heterogeneity in acute myeloid leukemia. Nat. Med. 22, 792–799 (2016).
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Papaemmanuil, E. et al. Genomic classification and prognosis in acute myeloid leukemia. N. Engl. J. Med. 374, 2209–2221 (2016).
Patel, J.P. et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N. Engl. J. Med. 366, 1079–1089 (2012).
Ho, P.A. et al. Leukemic mutations in the methylation-associated genes DNMT3A and IDH2 are rare events in pediatric AML: a report from the Children's Oncology Group. Pediatr. Blood Cancer 57, 204–209 (2011).
Farrar, J.E. et al. Genomic profiling of pediatric acute myeloid leukemia reveals a changing mutational landscape from disease diagnosis to relapse. Cancer Res. 76, 2197–2205 (2016).
Lange, B.J. et al. Outcomes in CCG-2961, a Children's Oncology Group phase 3 trial for untreated pediatric acute myeloid leukemia: a report from the Children's Oncology Group. Blood 111, 1044–1053 (2008).
Cooper, T.M. et al. AAML03P1, a pilot study of the safety of gemtuzumab ozogamicin in combination with chemotherapy for newly diagnosed childhood acute myeloid leukemia: a report from the Children's Oncology Group. Cancer 118, 761–769 (2012).
Gamis, A.S. et al. Gemtuzumab ozogamicin in children and adolescents with denovo acute myeloid leukemia improves event-free survival by reducing relapse risk: results from the randomized phase III Children's Oncology Group trial AAML0531. J. Clin. Oncol. 32, 3021–3032 (2014).
Lavallée, V.P. et al. Identification of MYC mutations in acute myeloid leukemias with NUP98-NSD1 translocations. Leukemia 30, 1621–1624 (2016).
Faber, Z.J. et al. The genomic landscape of core-binding factor acute myeloid leukemias. Nat. Genet. 48, 1551–1556 (2016).
Tarlock, K. et al. Discovery and functional validation of novel pediatric specific FLT3 activating mutations in acute myeloid leukemia: results from the COG/NCI target initiative. Blood 126, 87 (2015).
Ley, T.J. et al. DNMT3A mutations in acute myeloid leukemia. N. Engl. J. Med. 363, 2424–2433 (2010).
Wang, Y.A. et al. DNA methyltransferase-3a interacts with p53 and represses p53-mediated gene expression. Cancer Biol. Ther. 4, 1138–1143 (2005).
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014).
Goossens, S. et al. The EMT regulator Zeb2/Sip1 is essential for murine embryonic hematopoietic stem/progenitor cell differentiation and mobilization. Blood 117, 5620–5630 (2011).
Goossens, S. et al. ZEB2 drives immature T-cell lymphoblastic leukaemia development via enhanced tumour-initiating potential and IL-7 receptor signalling. Nat. Commun. 6, 5794 (2015).
Xiang, P. et al. Identification of E74-like factor 1 (ELF1) as a transcriptional regulator of the Hox cofactor MEIS1. Exp. Hematol. 38, 798–798 (2010).
Ostronoff, F. et al. NUP98/NSD1 and FLT3/ITD coexpression is more prevalent in younger AML patients and leads to induction failure: a COG and SWOG report. Blood 124, 2400–2407 (2014).
Maxson, J.E. et al. CSF3R mutations have a high degree of overlap with CEBPA mutations in pediatric AML. Blood 127, 3094–3098 (2016).
Quintana-Bustamante, O. et al. Overexpression of wild-type or mutants forms of CEBPA alter normal human hematopoiesis. Leukemia 26, 1537–1546 (2012).
Vicente, C., Conchillo, A., García-Sánchez, M.A. & Odero, M.D. The role of the GATA2 transcription factor in normal and malignant hematopoiesis. Crit. Rev. Oncol. Hematol. 82, 1–17 (2012).
Ng, K.P. et al. Runx1 deficiency permits granulocyte lineage commitment but impairs subsequent maturation. Oncogenesis 2, e78 (2013).
Wang, Y. et al. The Wnt/β-catenin pathway is required for the development of leukemia stem cells in AML. Science 327, 1650–1653 (2010).
Valencia, A. et al. Wnt signaling pathway is epigenetically regulated by methylation of Wnt antagonists in acute myeloid leukemia. Leukemia 23, 1658–1666 (2009).
Nanbakhsh, A. et al. c-Myc regulates expression of NKG2D ligands ULBP1/2/3 in AML and modulates their susceptibility to NK-mediated lysis. Blood 123, 3585–3595 (2014).
Marcucci, G. et al. MicroRNA expression in cytogenetically normal acute myeloid leukemia. N. Engl. J. Med. 358, 1919–1928 (2008).
Ramamurthy, R. et al. miR-155 expression and correlation with clinical outcome in pediatric AML: a report from Children's Oncology Group. Pediatr. Blood Cancer 63, 2096–2103 (2016).
Fooladinezhad, H., Khanahmad, H., Ganjalikhani-Hakemi, M. & Doosti, A. Negative regulation of TIM-3 expression in AML cell line (HL-60) using miR-330-5p. Br. J. Biomed. Sci. 73, 129–133 (2016).
Lim, E.L. et al. Comprehensive sequence analysis of relapse and refractory pediatric acute myeloid leukemia identifies miRNA and mRNA transcripts associated with treatment resistance—a report from the COG/NCI-target AML initiative. Blood 126, 687 (2015).
Gruber, T.A. et al. An Inv(16)(p13.3q24.3)-encoded CBFA2T3-GLIS2 fusion protein defines an aggressive subtype of pediatric acute megakaryoblastic leukemia. Cancer Cell 22, 683–697 (2012).
Liang, K. et al. Therapeutic targeting of MLL degradation pathways in MLL-rearranged leukemia. Cell 168, 59–72 (2017).
Itskovich, S.S. et al. MBNL1 as a new therapeutic target in MLL-fusion gene leukemia. Blood 126, 462 (2015).
Caudell, D. et al. Retroviral insertional mutagenesis identifies Zeb2 activation as a novel leukemogenic collaborating event in CALM-AF10 transgenic mice. Blood 115, 1194–1203 (2010).
Li, J. et al. The EMT transcription factor Zeb2 controls adult murine hematopoietic differentiation by regulating cytokine signaling. Blood 129, 460–472 (2017).
Shlush, L.I. et al. Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature 506, 328–333 (2014).
Guryanova, O.A. et al. DNMT3A mutations promote anthracycline resistance in acute myeloid leukemia via impaired nucleosome remodeling. Nat. Med. 22, 1488–1495 (2016).
Loghavi, S. et al. Clinical features of de novo acute myeloid leukemia with concurrent DNMT3A, FLT3 and NPM1 mutations. J. Hematol. Oncol. 7, 74 (2014).
Xu, B. et al. Tumor suppressor menin represses paired box gene 2 expression via Wilms tumor suppressor protein–Polycomb group complex. J. Biol. Chem. 286, 13937–13944 (2011).
Sinha, S. et al. Mutant WT1 is associated with DNA hypermethylation of PRC2 targets in AML and responds to EZH2 inhibition. Blood 125, 316–326 (2015).
Abdel-Wahab, O. et al. ASXL1 mutations promote myeloid transformation through loss of PRC2-mediated gene repression. Cancer Cell 22, 180–193 (2012).
Göllner, S. et al. Loss of the histone methyltransferase EZH2 induces resistance to multiple drugs in acute myeloid leukemia. Nat. Med. 23, 69–78 (2017).
Drmanac, R. et al. Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 327, 78–81 (2010).
Zhang, J. et al. Germline mutations in predisposition genes in pediatric cancer. N. Engl. J. Med. 373, 2336–2346 (2015).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Cingolani, P. et al. Using Drosophila melanogaster as a model for genotoxic chemical mutational studies with a new program, SnpSift. Front. Genet. 3, 35 (2012).
Saunders, C.T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor–normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Ye, K., Schulz, M.H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Zhou, X. et al. Exploring genomic alteration in pediatric cancer using ProteinPaint. Nat. Genet. 48, 4–6 (2016).
Lawrence, M.S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Zhang, Y., Zhou, H., Zhou, J. & Sun, W. Regression models for multivariate count data. J. Comput. Graph. Stat. 26, 1–13 (2017).
Leiserson, M.D., Wu, H.-T., Vandin, F. & Raphael, B.J. CoMEt: a statistical approach to identify combinations of mutually exclusive alterations in cancer. Genome Biol. 16, 160 (2015).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
van Borkulo, C.D. et al. A new method for constructing networks from binary data. Sci. Rep. 4, 5918 (2014).
Zeileis, A. & Grothendieck, G. zoo: S3 infrastructure for regular and irregular time series. J. Stat. Softw. 14, 1–27 (2005).
Mayakonda, A. & Koeffler, H.P. Maftools: Efficient analysis, visualization and summarization of MAF files from large-scale cohort based cancer studies. Preprint at https://www.biorxiv.org/content/early/2016/05/11/052662/ (2016).
Miller, C.A. et al. SciClone: inferring clonal architecture and tracking the spatial and temporal patterns of tumor evolution. PLOS Comput. Biol. 10, e1003665 (2014).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Chun, H.J. et al. Genome-wide profiles of extra-cranial malignant rhabdoid tumors reveal heterogeneity and dysregulated developmental pathways. Cancer Cell 29, 394–406 (2016).
Robertson, G. et al. De novo assembly and analysis of RNA-seq data. Nat. Methods 7, 909–912 (2010).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows—Wheeler transform. Bioinformatics 26, 589–595 (2010).
de Hoon, M.J. et al. Cross-mapping and the identification of editing sites in mature microRNAs in high-throughput sequencing libraries. Genome res. 20, 257–264 (2010).
Leek, J.T., Johnson, W.E., Parker, H.S., Jaffe, A.E. & Storey, J.D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Triche, T.J. Jr., Weisenberger, D.J., Van Den Berg, D., Laird, P.W. & Siegmund, K.D. Low-level processing of Illumina Infinium DNA methylation BeadArrays. Nucleic Acids Res. 41, e90 (2013).
Aryee, M.J. et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics 30, 1363–1369 (2014).
Fortin, J.P. et al. Functional normalization of 450k methylation array data improves replication in large cancer studies. Genome Biol. 15, 503 (2014).
Lee, D.D. & Seung, H.S. Learning the parts of objects by non-negative matrix factorization. Nature 401, 788–791 (1999).
Abrahamsson, J. et al. Response-guided induction therapy in pediatric acute myeloid leukemia with excellent remission rate. J. Clin. Oncol. 29, 310–315 (2011).
Burnett, A.K. et al. Identification of patients with acute myeloblastic leukemia who benefit from the addition of gemtuzumab ozogamicin: results of the MRC AML15 trial. J. Clin. Oncol. 29, 369–377 (2011).
Creutzig, U. et al. Less toxicity by optimizing chemotherapy, but not by addition of granulocyte colony-stimulating factor in children and adolescents with acute myeloid leukemia: results of AML-BFM 98. J. Clin. Oncol. 24, 4499–4506 (2006).
Creutzig, U. et al. Treatment strategies and long-term results in paediatric patients treated in four consecutive AML-BFM trials. Leukemia 19, 2030–2042 (2005).
Gibson, B.E. et al. Treatment strategy and long-term results in paediatric patients treated in consecutive UK AML trials. Leukemia 19, 2130–2138 (2005).
Kardos, G. et al. Treatment strategy and results in children treated on three Dutch Childhood Oncology Group acute myeloid leukemia trials. Leukemia 19, 2063–2071 (2005).
Perel, Y. et al. Impact of addition of maintenance therapy to intensive induction and consolidation chemotherapy for childhood acute myeloblastic leukemia: results of a prospective randomized trial, LAME 89/91. Leucámie Aiqüe Myéloïde Enfant. J. Clin. Oncol. 20, 2774–2782 (2002).
Pession, A. et al. Results of the AIEOP AML 2002/01 multicenter prospective trial for the treatment of children with acute myeloid leukemia. Blood 122, 170–178 (2013).
Balgobind, B.V. et al. Integrative analysis of type-I and type-II aberrations underscores the genetic heterogeneity of pediatric acute myeloid leukemia. Haematologica 96, 1478–1487 (2011).
Hollink, I.H. et al. NUP98/NSD1 characterizes a novel poor prognostic group in acute myeloid leukemia with a distinct HOX gene expression pattern. Blood 118, 3645–3656 (2011).
Hollink, I.H. et al. Clinical relevance of Wilms tumor 1 gene mutations in childhood acute myeloid leukemia. Blood 113, 5951–5960 (2009).
Hollink, I.H. et al. Favorable prognostic impact of NPM1 gene mutations in childhood acute myeloid leukemia, with emphasis on cytogenetically normal AML. Leukemia 23, 262–270 (2009).
Camp, R.L., Dolled-Filhart, M. & Rimm, D.L. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin. Cancer Res. 10, 7252–7259 (2004).
Acknowledgements
Dedicated to the memory of our colleague, mentor and friend, Dr. Robert Arceci, whose vision and perseverance set this effort in motion: “I may not have gone where I intended to go, but I think I have ended up where I needed to be” (Douglas Adams, The Long Dark Tea-Time of the Soul). The results published here are based on data generated by the TARGET initiative and TCGA. The TARGET initiative is supported by NCI Grant U10CA98543. Work performed under contracts from the NCI, US National Institutes of Health within HHSN261200800001E includes specimen processing (Children's Oncology Group Biopathology Center), WGS (Complete Genomics), and RNA-seq and TCS (British Columbia Cancer Agency). The content of this publication does not necessarily reflect the views or policies of the US Department of Health and Human Services, nor does mention of trade names, commercial products or organizations imply endorsement by the US government. Computation for the work described in this paper was supported in part by Fred Hutchinson Scientific Computing, University of Southern California's Center for High-Performance Computing and National Science Foundation (NSF) award ACI-1341935. This work was additionally supported by COG Chairs U10CA180886 and U10CA98543; COG Statistics and Data Center U10CA098413 and U10CA180899; COG Specimen Banking U24CA114766; R01CA114563 (S.M.); St. Baldrick's Foundation (J.E.F., T.T. and S.M.); Alex's Lemonade Stand (S.M.); Target Pediatric AML (TpAML), P20GM121293 (J.E.F.); the Arkansas Biosciences Institute (J.E.F.); and the Jane Anne Nohl Hematology Research Fund (T.T.).
Author information
Authors and Affiliations
Contributions
M.A.S., D.S.G., S.M. and R.A. conceived and led the project. R.E.R., M.A.M., J.M.G.A., T.M.D., P.G., L.C.H., D.S.G. and S.M. managed the project. H.B., J.E.F., T.T., R.E.R., E.L.L., T.A.A., Y.M., R.M., A.J.M., M.A.M., J.Z., X.M., Yu Liu, Yanling Liu, T.M.D., A.C.H., B.S. and S.R.P. generated, processed and analyzed the data. S.C., G.R., C.M.Z., S.N., E.A.K. and A.S.G. shared critical data and reagents. H.B., J.E.F., T.T., R.E.R., E.L.L. and S.M. drafted the manuscript. All authors edited and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–28 (PDF 5723 kb)
Supplementary Tables
Supplementary Tables 1–11 (XLSX 1244 kb)
Rights and permissions
About this article
Cite this article
Bolouri, H., Farrar, J., Triche, T. et al. The molecular landscape of pediatric acute myeloid leukemia reveals recurrent structural alterations and age-specific mutational interactions. Nat Med 24, 103–112 (2018). https://doi.org/10.1038/nm.4439
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4439
This article is cited by
-
Development and validation of a promising 5-gene prognostic model for pediatric acute myeloid leukemia
Molecular Biomedicine (2024)
-
GOLGA7 is essential for NRAS trafficking from the Golgi to the plasma membrane but not for its palmitoylation
Cell Communication and Signaling (2024)
-
LIN28B promotes differentiation of fully transformed AML cells but is dispensable for fetal leukemia suppression
Leukemia (2024)
-
New genomic classification of pediatric acute myeloid leukemia
Nature Genetics (2024)
-
TP-0184 inhibits FLT3/ACVR1 to overcome FLT3 inhibitor resistance and hinder AML growth synergistically with venetoclax
Leukemia (2024)