Abstract
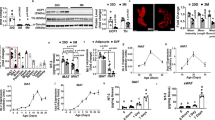
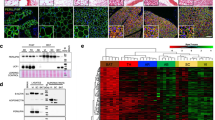
The cellular mechanism(s) linking macrophages to norepinephrine (NE)-mediated regulation of thermogenesis have been a topic of debate. Here we identify sympathetic neuron–associated macrophages (SAMs) as a population of cells that mediate clearance of NE via expression of solute carrier family 6 member 2 (SLC6A2), an NE transporter, and monoamine oxidase A (MAOA), a degradation enzyme. Optogenetic activation of the sympathetic nervous system (SNS) upregulates NE uptake by SAMs and shifts the SAM profile to a more proinflammatory state. NE uptake by SAMs is prevented by genetic deletion of Slc6a2 or inhibition of the encoded transporter. We also observed an increased proportion of SAMs in the SNS of two mouse models of obesity. Genetic ablation of Slc6a2 in SAMs increases brown adipose tissue (BAT) content, causes browning of white fat, increases thermogenesis, and leads to substantial and sustained weight loss in obese mice. We further show that this pathway is conserved, as human sympathetic ganglia also contain SAMs expressing the analogous molecular machinery for NE clearance, which thus constitutes a potential target for obesity treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Zeng, W. et al. Sympathetic neuro-adipose connections mediate leptin-driven lipolysis. Cell 163, 84–94 (2015).
Mathis, D. Immunological goings-on in visceral adipose tissue. Cell Metab. 17, 851–859 (2013).
Nguyen, K.D. et al. Alternatively activated macrophages produce catecholamines to sustain adaptive thermogenesis. Nature 480, 104–108 (2011).
Fischer, K. et al. Alternatively activated macrophages do not synthesize catecholamines or contribute to adipose tissue adaptive thermogenesis. Nat. Med. 23, 623–630 (2017).
Spadaro, O. et al. IGF1 shapes macrophage activation in response to immunometabolic challenge. Cell Rep. 19, 225–234 (2017).
Reitman, M.L. How does fat transition from white to beige? Cell Metab. 26, 14–16 (2017).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Anlauf, E. & Derouiche, A. Glutamine synthetase as an astrocytic marker: its cell type and vesicle localization. Front. Endocrinol. (Lausanne) 4, 144 (2013).
Bignami, A., Eng, L.F., Dahl, D. & Uyeda, C.T. Localization of the glial fibrillary acidic protein in astrocytes by immunofluorescence. Brain Res. 43, 429–435 (1972).
Chaudhry, F.A. et al. Glutamate transporters in glial plasma membranes: highly differentiated localizations revealed by quantitative ultrastructural immunocytochemistry. Neuron 15, 711–720 (1995).
Jessen, K.R. & Mirsky, R. The origin and development of glial cells in peripheral nerves. Nat. Rev. Neurosci. 6, 671–682 (2005).
Ludwin, S.K., Kosek, J.C. & Eng, L.F. The topographical distribution of S-100 and GFA proteins in the adult rat brain: an immunohistochemical study using horseradish peroxidase–labelled antibodies. J. Comp. Neurol. 165, 197–207 (1976).
Mearow, K.M., Mill, J.F. & Vitkovic, L. The ontogeny and localization of glutamine synthetase gene expression in rat brain. Brain Res. Mol. Brain Res. 6, 223–232 (1989).
Raff, M.C. et al. Galactocerebroside is a specific cell-surface antigenic marker for oligodendrocytes in culture. Nature 274, 813–816 (1978).
Regan, M.R. et al. Variations in promoter activity reveal a differential expression and physiology of glutamate transporters by glia in the developing and mature CNS. J. Neurosci. 27, 6607–6619 (2007).
Rusnakova, V. et al. Heterogeneity of astrocytes: from development to injury—single cell gene expression. PLoS One 8, e69734 (2013).
Sensenbrenner, M., Lucas, M. & Deloulme, J.C. Expression of two neuronal markers, growth-associated protein 43 and neuron-specific enolase, in rat glial cells. J. Mol. Med. (Berl.) 75, 653–663 (1997).
Buttgereit, A. et al. Sall1 is a transcriptional regulator defining microglia identity and function. Nat. Immunol. 17, 1397–1406 (2016).
Wentworth, J.M. et al. Pro-inflammatory CD11c+CD206+ adipose tissue macrophages are associated with insulin resistance in human obesity. Diabetes 59, 1648–1656 (2010).
Amano, S.U. et al. Local proliferation of macrophages contributes to obesity-associated adipose tissue inflammation. Cell Metab. 19, 162–171 (2014).
Wolf, Y. et al. Brown-adipose-tissue macrophages control tissue innervation and homeostatic energy expenditure. Nat. Immunol. 18, 665–674 (2017).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, eaal3222 (2017).
Clausen, B.E., Burkhardt, C., Reith, W., Renkawitz, R. & Förster, I. Conditional gene targeting in macrophages and granulocytes using LysMcre mice. Transgenic Res. 8, 265–277 (1999).
Merad, M. et al. Langerhans cells renew in the skin throughout life under steady-state conditions. Nat. Immunol. 3, 1135–1141 (2002).
Shirey-Rice, J.K. et al. Norepinephrine transporter variant A457P knock-in mice display key features of human postural orthostatic tachycardia syndrome. Dis. Model. Mech. 6, 1001–1011 (2013).
Aderem, A. & Underhill, D.M. Mechanisms of phagocytosis in macrophages. Annu. Rev. Immunol. 17, 593–623 (1999).
Stjärne, L. Basic mechanisms and local modulation of nerve impulse–induced secretion of neurotransmitters from individual sympathetic nerve varicosities. Rev. Physiol. Biochem. Pharmacol. 112, 1–137 (1989).
Schroeder, C. & Jordan, J. Norepinephrine transporter function and human cardiovascular disease. Am. J. Physiol. Heart Circ. Physiol. 303, H1273–H1282 (2012).
Filiano, A.J. et al. Unexpected role of interferon-γ in regulating neuronal connectivity and social behaviour. Nature 535, 425–429 (2016).
Galle-Treger, L. et al. Nicotinic acetylcholine receptor agonist attenuates ILC2-dependent airway hyperreactivity. Nat. Commun. 7, 13202 (2016).
Ibiza, S. et al. Glial-cell-derived neuroregulators control type 3 innate lymphoid cells and gut defence. Nature 535, 440–443 (2016).
Kipnis, J. Multifaceted interactions between adaptive immunity and the central nervous system. Science 353, 766–771 (2016).
Louveau, A. et al. Structural and functional features of central nervous system lymphatic vessels. Nature 523, 337–341 (2015).
Rosas-Ballina, M. et al. Acetylcholine-synthesizing T cells relay neural signals in a vagus nerve circuit. Science 334, 98–101 (2011).
Gabanyi, I. et al. Neuro-immune interactions drive tissue programming in intestinal macrophages. Cell 164, 378–391 (2016).
Gautier, E.L. et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat. Immunol. 13, 1118–1128 (2012).
Okabe, Y. & Medzhitov, R. Tissue-specific signals control reversible program of localization and functional polarization of macrophages. Cell 157, 832–844 (2014).
Crotti, A. & Ransohoff, R.M. Microglial physiology and pathophysiology: insights from genome-wide transcriptional profiling. Immunity 44, 505–515 (2016).
Prinz, M. & Priller, J. Microglia and brain macrophages in the molecular age: from origin to neuropsychiatric disease. Nat. Rev. Neurosci. 15, 300–312 (2014).
Kong, Y., Ruan, L., Qian, L., Liu, X. & Le, Y. Norepinephrine promotes microglia to uptake and degrade amyloid β peptide through upregulation of mouse formyl peptide receptor 2 and induction of insulin-degrading enzyme. J. Neurosci. 30, 11848–11857 (2010).
Kettenmann, H. & Ransom, B.R. Neuroglia (Oxford University Press, 2013).
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Hanani, M. Satellite glial cells in sensory ganglia: from form to function. Brain Res. Brain Res. Rev. 48, 457–476 (2005).
Hanani, M. Satellite glial cells in sympathetic and parasympathetic ganglia: in search of function. Brain Res. Rev. 64, 304–327 (2010).
Camell, C.D. et al. Inflammasome-driven catecholamine catabolism in macrophages blunts lipolysis during ageing. Nature http://dx.doi.org/10.1038/nature24022 (2017).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Acknowledgements
We would like to thank the Unit for Imaging and Cytometry at the Instituto Gulbenkian de Ciência (IGC) for assistance with flow cytometry, cell sorting, and multiphoton microscopy. We also want to thank the Antibody Service at the IGC for the antibodies produced in house and the Histopathology facility at the IGC for tissue processing and histological assessment. This work was supported by the Fundação para a Ciéncia e Tecnologia (FCT), the European Molecular Biology Organization (EMBO), the Human Frontier Science Program (HFSP), Maratona da Saúde, and the US National Institutes of Health (NIH). R.M.P. was supported by FCT (SFRH/BD/88454/2012), J.S.S. was supported by the American Heart Association (16PRE30980030) and a training grant (T32DK007541), B.A.A. was supported by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), and N.M.-S. was supported by Xunta de Galicia (ED481B 2016/168-0). We thank M. Aouadi for helpful discussions.
Author information
Authors and Affiliations
Contributions
A.I.D. conceptualized the study. R.M.P. performed two-photon and confocal microscopy. E.S. and R.M.P. performed flow cytometry. J.S.S. and R.M.P. performed low-input RNA-seq. V.M.L., J.S.S., and R.M.P. analyzed the RNA-seq data. M.I., A.L.S., S.A., and E.T. performed electron microscopy. E.S., R.M.P., N.M.S., I. Mahú, B.A.A., and C.M.L. performed functional tests. N.K., I. Morris, R.M., and V.G. performed related mouse husbandry and genotyping. F.T. and M.V. processed human ganglia. M.K.H. provided the Slc6a2−/− mice. N.J.S. developed the low-input RNA-seq protocols. A.I.D., C.K.G., and R.M.P. wrote the original draft of the manuscript. A.I.D., C.K.G., R.M.P., and C.M.L. reviewed and edited the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 and Supplementary Table 1 (PDF 20527 kb)
In vivo visualization of SAMs in the neuro-adipose connection.
Intra-vital multi-photon visualization of a neuro-adipose connection in the inguinal fat pad of a live Cx3cr1GFP/+ mouse; LipidTOX (blue) labels adipocytes. Images are representative of 3 similar experiments. (MPG 192 kb)
In vivo visualization of ATMs in the subcutaneous adipose tissue
Intra-vital multi-photon visualization of the inguinal fat pad of a live Cx3cr1GFP/+ mouse; LipidTOX (blue) labels adipocytes. Images are representative of 3 similar experiments. (MPG 56 kb)
Rights and permissions
About this article
Cite this article
Pirzgalska, R., Seixas, E., Seidman, J. et al. Sympathetic neuron–associated macrophages contribute to obesity by importing and metabolizing norepinephrine. Nat Med 23, 1309–1318 (2017). https://doi.org/10.1038/nm.4422
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4422
This article is cited by
-
M2 macrophages independently promote beige adipogenesis via blocking adipocyte Ets1
Nature Communications (2024)
-
Macrophage and T cell networks in adipose tissue
Nature Reviews Endocrinology (2024)
-
The cancer-immune dialogue in the context of stress
Nature Reviews Immunology (2024)
-
Unraveling the complex roles of macrophages in obese adipose tissue: an overview
Frontiers of Medicine (2024)
-
Regional differences in the ultrastructure of mucosal macrophages in the rat large intestine
Cell and Tissue Research (2024)