Abstract
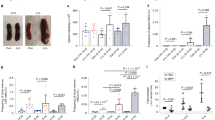
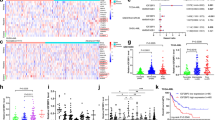
N6-methyladenosine (m6A) is an abundant nucleotide modification in mRNA that is required for the differentiation of mouse embryonic stem cells. However, it remains unknown whether the m6A modification controls the differentiation of normal and/or malignant myeloid hematopoietic cells. Here we show that shRNA-mediated depletion of the m6A-forming enzyme METTL3 in human hematopoietic stem/progenitor cells (HSPCs) promotes cell differentiation, coupled with reduced cell proliferation. Conversely, overexpression of wild-type METTL3, but not of a catalytically inactive form of METTL3, inhibits cell differentiation and increases cell growth. METTL3 mRNA and protein are expressed more abundantly in acute myeloid leukemia (AML) cells than in healthy HSPCs or other types of tumor cells. Furthermore, METTL3 depletion in human myeloid leukemia cell lines induces cell differentiation and apoptosis and delays leukemia progression in recipient mice in vivo. Single-nucleotide-resolution mapping of m6A coupled with ribosome profiling reveals that m6A promotes the translation of c-MYC, BCL2 and PTEN mRNAs in the human acute myeloid leukemia MOLM-13 cell line. Moreover, loss of METTL3 leads to increased levels of phosphorylated AKT, which contributes to the differentiation-promoting effects of METTL3 depletion. Overall, these results provide a rationale for the therapeutic targeting of METTL3 in myeloid leukemia.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Meyer, K.D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Geula, S. et al. Stem cells. m6A mRNA methylation facilitates resolution of naïve pluripotency toward differentiation. Science 347, 1002–1006 (2015).
Batista, P.J. et al. m6A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014).
Li, Z. et al. FTO plays an oncogenic role in acute myeloid leukemia as a N6-methyladenosine RNA demethylase. Cancer Cell 31, 127–141 (2017).
Mauer, J. et al. Reversible methylation of m6Am in the 5′ cap controls mRNA stability. Nature 541, 371–375 (2017).
Wang, X. et al. Corrigendum: Structural basis of N6-adenosine methylation by the METTL3-METTL14 complex. Nature 542, 260 (2017).
Lin, S., Choe, J., Du, P., Triboulet, R. & Gregory, R.I. The m6A methyltransferase METTL3 promotes translation in human cancer cells. Mol. Cell 62, 335–345 (2016).
Bagger, F.O. et al. BloodSpot: a database of gene expression profiles and transcriptional programs for healthy and malignant haematopoiesis. Nucleic Acids Res. 44 D1, D917–D924 (2016).
Śledź, P. & Jinek, M. Structural insights into the molecular mechanism of the m(6)A writer complex. eLife 5, e18434 (2016).
Wang, P., Doxtader, K.A. & Nam, Y. Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases. Mol. Cell 63, 306–317 (2016).
Wang, X. et al. Structural basis of N6-adenosine methylation by the METTL3-METTL14 complex. Nature 534, 575–578 (2016).
Shi, J. et al. Discovery of cancer drug targets by CRISPR-Cas9 screening of protein domains. Nat. Biotechnol. 33, 661–667 (2015).
Wang, T., et al. Gene essentiality profiling reveals gene networks and synthetic lethal interactions with oncogenic Ras. Cell 168, 890–903 (2017).
Linder, B. et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat. Methods 12, 767–772 (2015).
Sommer, S., Lavi, U. & Darnell, J.E. Jr . The absolute frequency of labeled N-6-methyladenosine in HeLa cell messenger RNA decreases with label time. J. Mol. Biol. 124, 487–499 (1978).
Ingolia, N.T., Lareau, L.F. & Weissman, J.S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Wang, X. et al. N6-methyladenosine modulates messenger RNA translation efficiency. Cell 161, 1388–1399 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Park, S.M. et al. Musashi-2 controls cell fate, lineage bias, and TGF-β signaling in HSCs. J. Exp. Med. 211, 71–87 (2014).
Li, S. et al. Distinct evolution and dynamics of epigenetic and genetic heterogeneity in acute myeloid leukemia. Nat. Med. 22, 792–799 (2016).
Sarry, J.E. et al. Human acute myelogenous leukemia stem cells are rare and heterogeneous when assayed in NOD/SCID/IL2Rgc-deficient mice. J. Clin. Invest. 121, 384–395 (2011).
Stoneley, M. et al. c-Myc protein synthesis is initiated from the internal ribosome entry segment during apoptosis. Mol. Cell. Biol. 20, 1162–1169 (2000).
Pandey, S. & Wang, E. Cells en route to apoptosis are characterized by the upregulation of c-fos, c-myc, c-jun, cdc2, and RB phosphorylation, resembling events of early cell-cycle traverse. J. Cell. Biochem. 58, 135–150 (1995).
Sherrill, K.W., Byrd, M.P., Van Eden, M.E. & Lloyd, R.E. BCL-2 translation is mediated via internal ribosome entry during cell stress. J. Biol. Chem. 279, 29066–29074 (2004).
Ping, X.L. et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 24, 177–189 (2014).
Kharas, M.G. et al. Constitutively active AKT depletes hematopoietic stem cells and induces leukemia in mice. Blood 115, 1406–1415 (2010).
Sykes, S.M. et al. AKT/FOXO signaling enforces reversible differentiation blockade in myeloid leukemias. Cell 146, 697–708 (2011).
Cui, Q. et al. m6A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Reports 18, 2622–2634 (2017).
Zhang, S., et al. m6A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1 expression and cell proliferation program. Cancer Cell 31, 591–606 (2017).
Kwok, C.T., Marshall, A.D., Rasko, J.E. & Wong, J.J. Genetic alterations of m(6)A regulators predict poorer survival in acute myeloid leukemia. J. Hematol. Oncol. 10, 39 (2017).
Patil, D.P. et al. m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537, 369–373 (2016).
Olarerin-George, A.O. & Jaffrey, S.R. MetaPlotR: a Perl/R pipeline for plotting metagenes of nucleotide modifications and other transcriptomic sites. Bioinformatics 33, 1563–1564 (2017).
Bailey, T.L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–208 (2009).
Kruse, S. et al. A novel synthesis and detection method for cap-associated adenosine modifications in mouse mRNA. Sci. Rep. 1, 126 (2011).
Reid, D.W., Shenolikar, S. & Nicchitta, C.V. Simple and inexpensive ribosome profiling analysis of mRNA translation. Methods 91, 69–74 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank D. Bachovchin (MSKCC, New York) for the MOLM-13 constitutively expressing Cas9 cell line, and C. Vakoc (Cold Spring Harbor Laboratory, New York) for the constitutively expressing Cas9-RN2c cell line. We would like to thank the members of the Scaltriti laboratory (MSKCC, New York) for providing us with PI3K/AKT inhibitors. We thank the Weill Cornell Medicine Epigenomics Core for their assistance with sequencing. M.G.K. was supported by the US National Institutes of Health National Institute of Diabetes Digestive and Kidney Diseases Career Development Award, NIDDK NIH R01-DK101989-01A1, NCI 1R01CA193842-01, Kimmel Scholar Award, V-Scholar Award, Geoffrey Beene Award, Leukemia Lymphoma Society Career Development Award and Alex's Lemonade Stand A Award. This work was also supported by a Tri-Institutional Stem Cell Award (M.G.K. and S.R.J.), R01CA186702 (S.R.J.), T32CA062948 (B.F.P.), Ruth L. Kirschstein National Research Service Award 1F32CA22104-01 (B.F.P.), a Damon Runyon-Sohn Pediatric Cancer Fellowship Award DRSG10-14 (L.P.V.), and the American-Italian Cancer Foundation (S.Z.). The research was funded in part through the NIH/NCI Cancer Support Core Grant P30 CA08748 MGK. The RPPA core facility is funded by NCI #CA16672.
Author information
Authors and Affiliations
Contributions
L.P.V. led the project, performed experiments, analyzed data and wrote the manuscript. B.F.P., Y.C., D.N. and S.Z. performed experiments, analyzed data and edited the manuscript. G.M., T.C. and A.C. provided experimental supports. C.E.M., Y.S. and M.M. performed and analyzed MeRIP-seq experiments on patient-derived samples. J.S., C.F., M.P., F.E.G.-B., A.M., V.M.K. and M.C. provided patient samples. S.R.J. supervised the project and wrote the manuscript. M.G.K. directed the project, analyzed data and wrote the manuscript.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
Supplementary Figures 1–6 (PDF 1925 kb)
Supplementary Dataset
Supplementary Tables 1–13 (XLSX 9967 kb)
Rights and permissions
About this article
Cite this article
Vu, L., Pickering, B., Cheng, Y. et al. The N6-methyladenosine (m6A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat Med 23, 1369–1376 (2017). https://doi.org/10.1038/nm.4416
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4416
This article is cited by
-
METTL1 mediated tRNA m7G modification promotes leukaemogenesis of AML via tRNA regulated translational control
Experimental Hematology & Oncology (2024)
-
Ribosome profiling: a powerful tool in oncological research
Biomarker Research (2024)
-
METTL3 recruiting M2-type immunosuppressed macrophages by targeting m6A-SNAIL-CXCL2 axis to promote colorectal cancer pulmonary metastasis
Journal of Experimental & Clinical Cancer Research (2024)
-
Identification of a novel m6A-related lncRNAs signature and immunotherapeutic drug sensitivity in pancreatic adenocarcinoma
BMC Cancer (2024)
-
A novel AML1-ETO/FTO positive feedback loop promotes leukemogenesis and Ara-C resistance via stabilizing IGFBP2 in t(8;21) acute myeloid leukemia
Experimental Hematology & Oncology (2024)