Abstract
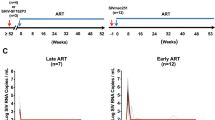
In the quest for a functional cure or the eradication of HIV infection, it is necessary to know the sizes of the reservoirs from which infection rebounds after treatment interruption. Thus, we quantified SIV and HIV tissue burdens in tissues of infected nonhuman primates and lymphoid tissue (LT) biopsies from infected humans. Before antiretroviral therapy (ART), LTs contained >98% of the SIV RNA+ and DNA+ cells. With ART, the numbers of virus (v) RNA+ cells substantially decreased but remained detectable, and their persistence was associated with relatively lower drug concentrations in LT than in peripheral blood. Prolonged ART also decreased the levels of SIV- and HIV-DNA+ cells, but the estimated size of the residual tissue burden of 108 vDNA+ cells potentially containing replication-competent proviruses, along with evidence of continuing virus production in LT despite ART, indicated two important sources for rebound following treatment interruption. The large sizes of these tissue reservoirs underscore challenges in developing 'HIV cure' strategies targeting multiple sources of virus production.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Yukl, S.A. et al. Challenges in detecting HIV persistence during potentially curative interventions: a study of the Berlin patient. PLoS Pathog. 9, e1003347 (2013).
Embretson, J. et al. Analysis of human immunodeficiency virus-infected tissues by amplification and in situ hybridization reveals latent and permissive infections at single-cell resolution. Proc. Natl. Acad. Sci. USA 90, 357–361 (1993).
Embretson, J. et al. Massive covert infection of helper T lymphocytes and macrophages by HIV during the incubation period of AIDS. Nature 362, 359–362 (1993).
Faust, R.A. et al. Outpatient biopsies of the palatine tonsil: access to lymphoid tissue for assessment of human immunodeficiency virus RNA titers. Otolaryngol. Head Neck Surg. 114, 593–598 (1996).
Fletcher, C.V. et al. Persistent HIV-1 replication is associated with lower antiretroviral drug concentrations in lymphatic tissues. Proc. Natl. Acad. Sci. USA 111, 2307–2312 (2014).
Haase, A.T. Population biology of HIV-1 infection: viral and CD4+ T cell demographics and dynamics in lymphatic tissues. Annu. Rev. Immunol. 17, 625–656 (1999).
Reinhart, T.A. et al. Simian immunodeficiency virus burden in tissues and cellular compartments during clinical latency and AIDS. J. Infect. Dis. 176, 1198–1208 (1997).
Reinhart, T.A. et al. A new approach to investigating the relationship between productive infection and cytopathicity in vivo. Nat. Med. 3, 218–221 (1997).
Chomont, N. et al. HIV reservoir size and persistence are driven by T cell survival and homeostatic proliferation. Nat. Med. 15, 893–900 (2009).
Chun, T.W. et al. Quantification of latent tissue reservoirs and total body viral load in HIV-1 infection. Nature 387, 183–188 (1997).
Chun, T.W. et al. Rebound of plasma viremia following cessation of antiretroviral therapy despite profoundly low levels of HIV reservoir: implications for eradication. AIDS 24, 2803–2808 (2010).
Chun, T.W. et al. HIV-infected individuals receiving effective antiviral therapy for extended periods of time continually replenish their viral reservoir. J. Clin. Invest. 115, 3250–3255 (2005).
Pantaleo, G. et al. Lymphoid organs function as major reservoirs for human immunodeficiency virus. Proc. Natl. Acad. Sci. USA 88, 9838–9842 (1991).
Schacker, T. et al. Rapid accumulation of human immunodeficiency virus (HIV) in lymphatic tissue reservoirs during acute and early HIV infection: implications for timing of antiretroviral therapy. J. Infect. Dis. 181, 354–357 (2000).
Schnittman, S.M. et al. The reservoir for HIV-1 in human peripheral blood is a T cell that maintains expression of CD4. Science 245, 305–308 (1989).
Haase, A.T. et al. Quantitative image analysis of HIV-1 infection in lymphoid tissue. Science 274, 985–989 (1996).
Li, Q. et al. Peak SIV replication in resting memory CD4+ T cells depletes gut lamina propria CD4+ T cells. Nature 434, 1148–1152 (2005).
Schacker, T. et al. Productive infection of T cells in lymphoid tissues during primary and early human immunodeficiency virus infection. J. Infect. Dis. 183, 555–562 (2001).
Rothenberger, M.K. et al. Large number of rebounding/founder HIV variants emerge from multifocal infection in lymphatic tissues after treatment interruption. Proc. Natl. Acad. Sci. USA 112, E1126–E1134 (2015).
Lorenzo-Redondo, R. et al. Persistent HIV-1 replication maintains the tissue reservoir during therapy. Nature 530, 51–56 (2016).
Ho, Y.C. et al. Replication-competent noninduced proviruses in the latent reservoir increase barrier to HIV-1 cure. Cell 155, 540–551 (2013).
Deleage, C. et al. Defining HIV and SIV reservoirs in lymphoid tissues. Pathog. Immun. 1, 68–106 (2016).
Reilly, C., Wietgrefe, S., Sedgewick, G. & Haase, A. Determination of simian immunodeficiency virus production by infected activated and resting cells. AIDS 21, 163–168 (2007).
Del Prete, G.Q. et al. Elevated plasma viral loads in romidepsin-treated simian immunodeficiency virus-infected rhesus macaques on suppressive combination antiretroviral therapy. Antimicrob. Agents Chemother. 60, 1560–1572 (2015).
Del Prete, G.Q. et al. Effect of suberoylanilide hydroxamic acid (SAHA) administration on the residual virus pool in a model of combination antiretroviral therapy-mediated suppression in SIVmac239-infected indian rhesus macaques. Antimicrob. Agents Chemother. 58, 6790–6806 (2014).
North, T.W. et al. Viral sanctuaries during highly active antiretroviral therapy in a nonhuman primate model for AIDS. J. Virol. 84, 2913–2922 (2010).
Clements, J.E. et al. The central nervous system as a reservoir for simian immunodeficiency virus (SIV): steady-state levels of SIV DNA in brain from acute through asymptomatic infection. J. Infect. Dis. 186, 905–913 (2002).
Clements, J.E. et al. The central nervous system is a viral reservoir in simian immunodeficiency virus–infected macaques on combined antiretroviral therapy: a model for human immunodeficiency virus patients on highly active antiretroviral therapy. J. Neurovirol. 11, 180–189 (2005).
Estes, J. et al. Collagen deposition limits immune reconstitution in the gut. J. Infect. Dis. 198, 456–464 (2008).
Estes, J.D., Haase, A.T. & Schacker, T.W. The role of collagen deposition in depleting CD4+ T cells and limiting reconstitution in HIV-1 and SIV infections through damage to the secondary lymphoid organ niche. Semin. Immunol. 20, 181–186 (2008).
Schacker, T.W. et al. Lymphatic tissue fibrosis is associated with reduced numbers of naive CD4+ T cells in human immunodeficiency virus type 1 infection. Clin. Vaccine Immunol. 13, 556–560 (2006).
Schacker, T.W. et al. Collagen deposition in HIV-1 infected lymphatic tissues and T cell homeostasis. J. Clin. Invest. 110, 1133–1139 (2002).
Schacker, T.W. et al. Amount of lymphatic tissue fibrosis in HIV infection predicts magnitude of HAART-associated change in peripheral CD4 cell count. AIDS 19, 2169–2171 (2005).
Bruner, K.M. et al. Defective proviruses rapidly accumulate during acute HIV-1 infection. Nat. Med. 22, 1043–1049 (2016).
Lee, E.M. et al. Molecular methods for evaluation of virological status of nonhuman primates challenged with simian immunodeficiency or simian-human immunodeficiency viruses. J. Virol. Methods 163, 287–294 (2010).
Zhang, Z.Q. et al. Roles of substrate availability and infection of resting and activated CD4+ T cells in transmission and acute simian immunodeficiency virus infection. Proc. Natl. Acad. Sci. USA 101, 5640–5645 (2004).
Cavert, W. et al. Kinetics of response in lymphoid tissues to antiretroviral therapy of HIV-1 infection. Science 276, 960–964 (1997).
Podany, A.T., Winchester, L.C., Robbins, B.L. & Fletcher, C.V. Quantification of cell-associated atazanavir, darunavir, lopinavir, ritonavir, and efavirenz concentrations in human mononuclear cell extracts. Antimicrob. Agents Chemother. 58, 2866–2870 (2014).
Guidance for Industry. Bioanalytical Method Validation (U.S. Department of Health and Human Services, 2001).
Acknowledgements
The authors thank S. Povolny and C. Olson for help with manuscript preparation and editing, and the Antiviral Pharmacology Laboratory at the University of Nebraska for quantification of ARV concentrations. This work was supported by grants UL1TR000114 (T.W.S.), AI096109 (J.D.E., L. Swainson, S.G.D., J.A., J.J., T.E.S., J.M.M., and T.W.S.), OD011107 (P.A.L.), AI124965 (C.V.F.), and AI074340 (J.G.C., G.J.B., T.H., A.K., J.A., J.J., T.E.S., M.H., S.P.C., J.S., C.V.F., A.T.H., and T.W.S.), and in part by federal funds from the National Cancer Institute, National Institutes of Health, under contract no. HHSN261200800001E (J.D.E., C.D., and J.D.L.). The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government.
Author information
Authors and Affiliations
Contributions
J.D.E. contributed to study design and oversight, sample analysis, and manuscript preparation. C.K. contributed to study design, oversight of the Uganda cohort, and manuscript preparation. F.S. contributed to management of the Ugandan cohort, including regulatory and sample management. L. Swainson, G.Q.D.P., J.A., C.D., J.J., T.E.S., M.H., S.P.C., H.P., T.R., J. Schuster, J. Schoephoerster, and K.P. contributed to sample analysis. P.S. contributed to sample and data analysis. K.N.M. contributed to sample collection and analysis, and data management. J.G.C., G.J.B., and T.H. performed surgery to collect LN and rectal samples from the Ugandan cohort. A.K. contributed to sample collection and study design. L. Shang performed the TSA/ELF assays and analysis. S.W.W. contributed to data analysis and provided technical support. C.V.F. contributed to drug-level data acquisition, sample analysis, and manuscript preparation. S.G.D., P.A.L., J.D.L., D.C.D., J.M.M., A.T.H., and T.W.S. contributed to study design, data analysis, and manuscript preparation
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–4. (PDF 3106 kb)
Rights and permissions
About this article
Cite this article
Estes, J., Kityo, C., Ssali, F. et al. Defining total-body AIDS-virus burden with implications for curative strategies. Nat Med 23, 1271–1276 (2017). https://doi.org/10.1038/nm.4411
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4411
This article is cited by
-
Immune targeting of HIV-1 reservoir cells: a path to elimination strategies and cure
Nature Reviews Microbiology (2024)
-
Mucosal T-cell responses to chronic viral infections: Implications for vaccine design
Cellular & Molecular Immunology (2024)
-
More than the Infinite Monkey Theorem: NHP Models in the Development of a Pediatric HIV Cure
Current HIV/AIDS Reports (2024)
-
HIV infection of non-classical cells in the brain
Retrovirology (2023)
-
Phenotypic signatures of immune selection in HIV-1 reservoir cells
Nature (2023)