Abstract
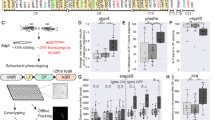
Mutations in MECP2 cause Rett syndrome (RTT), an X-linked neurological disorder characterized by regressive loss of neurodevelopmental milestones and acquired psychomotor deficits. However, the cellular heterogeneity of the brain impedes an understanding of how MECP2 mutations contribute to RTT. Here we developed a Cre-inducible method for cell-type-specific biotin tagging of MeCP2 in mice. Combining this approach with an allelic series of knock-in mice carrying frequent RTT-associated mutations (encoding T158M and R106W) enabled the selective profiling of RTT-associated nuclear transcriptomes in excitatory and inhibitory cortical neurons. We found that most gene-expression changes were largely specific to each RTT-associated mutation and cell type. Lowly expressed cell-type-enriched genes were preferentially disrupted by MeCP2 mutations, with upregulated and downregulated genes reflecting distinct functional categories. Subcellular RNA analysis in MeCP2-mutant neurons further revealed reductions in the nascent transcription of long genes and uncovered widespread post-transcriptional compensation at the cellular level. Finally, we overcame X-linked cellular mosaicism in female RTT models and identified distinct gene-expression changes between neighboring wild-type and mutant neurons, providing contextual insights into RTT etiology that support personalized therapeutic interventions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Chahrour, M. & Zoghbi, H.Y. The story of Rett syndrome: from clinic to neurobiology. Neuron 56, 422–437 (2007).
Amir, R.E. et al. Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat. Genet. 23, 185–188 (1999).
Shahbazian, M.D., Antalffy, B., Armstrong, D.L. & Zoghbi, H.Y. Insight into Rett syndrome: MeCP2 levels display tissue- and cell-specific differences and correlate with neuronal maturation. Hum. Mol. Genet. 11, 115–124 (2002).
Lewis, J.D. et al. Purification, sequence, and cellular localization of a novel chromosomal protein that binds to methylated DNA. Cell 69, 905–914 (1992).
Jones, P.L. et al. Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nat. Genet. 19, 187–191 (1998).
Lyst, M.J. et al. Rett syndrome mutations abolish the interaction of MeCP2 with the NCoR/SMRT co-repressor. Nat. Neurosci. 16, 898–902 (2013).
Nan, X. et al. Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. Nature 393, 386–389 (1998).
Skene, P.J. et al. Neuronal MeCP2 is expressed at near histone-octamer levels and globally alters the chromatin state. Mol. Cell 37, 457–468 (2010).
Chahrour, M. et al. MeCP2, a key contributor to neurological disease, activates and represses transcription. Science 320, 1224–1229 (2008).
Chen, L. et al. MeCP2 binds to non-CG methylated DNA as neurons mature, influencing transcription and the timing of onset for Rett syndrome. Proc. Natl. Acad. Sci. USA 112, 5509–5514 (2015).
Li, Y. et al. Global transcriptional and translational repression in human-embryonic-stem-cell-derived Rett syndrome neurons. Cell Stem Cell 13, 446–458 (2013).
Cuddapah, V.A. et al. Methyl-CpG-binding protein 2 (MECP2) mutation type is associated with disease severity in Rett syndrome. J. Med. Genet. 51, 152–158 (2014).
Ghosh, R.P., Horowitz-Scherer, R.A., Nikitina, T., Gierasch, L.M. & Woodcock, C.L. Rett syndrome–causing mutations in human MeCP2 result in diverse structural changes that impact folding and DNA interactions. J. Biol. Chem. 283, 20523–20534 (2008).
Ho, K.L. et al. MeCP2 binding to DNA depends upon hydration at methyl-CpG. Mol. Cell 29, 525–531 (2008).
Ballestar, E., Yusufzai, T.M. & Wolffe, A.P. Effects of Rett syndrome mutations of the methyl-CpG binding domain of the transcriptional repressor MeCP2 on selectivity for association with methylated DNA. Biochemistry 39, 7100–7106 (2000).
Brown, K. et al. The molecular basis of variable phenotypic severity among common missense mutations causing Rett syndrome. Hum. Mol. Genet. 25, 558–570 2016).
Goffin, D. et al. Rett syndrome mutation MeCP2 T158A disrupts DNA binding, protein stability and ERP responses. Nat. Neurosci. 15, 274–283 (2011).
Baker, S.A. et al. An AT-hook domain in MeCP2 determines the clinical course of Rett syndrome and related disorders. Cell 152, 984–996 (2013).
Katz, D.M. et al. Preclinical research in Rett syndrome: setting the foundation for translational success. Dis. Model. Mech. 5, 733–745 (2012).
Lamonica, J.M. et al. Elevating expression of MeCP2 T158M rescues DNA binding and Rett syndrome–like phenotypes. J. Clin. Invest. 127, 1889–1904 (2017).
Lyst, M.J. & Bird, A. Rett syndrome: a complex disorder with simple roots. Nat. Rev. Genet. 16, 261–275 (2015).
Fishell, G. & Heintz, N. The neuron identity problem: form meets function. Neuron 80, 602–612 (2013).
Molyneaux, B.J. et al. DeCoN: genome-wide analysis of in vivo transcriptional dynamics during pyramidal neuron fate selection in neocortex. Neuron 85, 275–288 (2015).
Mo, A. et al. Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86, 1369–1384 (2015).
Zhao, Y.-T., Goffin, D., Johnson, B.S. & Zhou, Z. Loss of MeCP2 function is associated with distinct gene expression changes in the striatum. Neurobiol. Dis. 59, 257–266 (2013).
Gabel, H.W. et al. Disruption of DNA-methylation-dependent long gene repression in Rett syndrome. Nature 522, 89–93 (2015).
Guo, J.U. et al. Distribution, recognition and regulation of non-CpG methylation in the adult mammalian brain. Nat. Neurosci. 17, 215–222 (2014).
Rube, H.T. et al. Sequence features accurately predict genome-wide MeCP2 binding in vivo. Nat. Commun. 7, 11025 (2016).
Deal, R.B. & Henikoff, S. A simple method for gene expression and chromatin profiling of individual cell types within a tissue. Dev. Cell 18, 1030–1040 (2010).
Lakso, M. et al. Efficient in vivo manipulation of mouse genomic sequences at the zygote stage. Proc. Natl. Acad. Sci. USA 93, 5860–5865 (1996).
Samaco, R.C. et al. A partial loss of function allele of methyl-CpG-binding protein 2 predicts a human neurodevelopmental syndrome. Hum. Mol. Genet. 17, 1718–1727 (2008).
Guy, J., Gan, J., Selfridge, J., Cobb, S. & Bird, A. Reversal of neurological defects in a mouse model of Rett syndrome. Science 315, 1143–1147 (2007).
Kumar, A. et al. Analysis of protein domains and Rett syndrome mutations indicate that multiple regions influence chromatin-binding dynamics of the chromatin-associated protein MECP2 in vivo. J. Cell Sci. 121, 1128–1137 (2008).
Goebbels, S. et al. Genetic targeting of principal neurons in neocortex and hippocampus of NEX-Cre mice. Genes 44, 611–621 (2006).
Monory, K. et al. The endocannabinoid system controls key epileptogenic circuits in the hippocampus. Neuron 51, 455–466 (2006).
Bhatt, D.M. et al. Transcript dynamics of proinflammatory genes revealed by sequence analysis of subcellular RNA fractions. Cell 150, 279–290 (2012).
Ameur, A. et al. Total RNA sequencing reveals nascent transcription and widespread co-transcriptional splicing in the human brain. Nat. Struct. Mol. Biol. 18, 1435–1440 (2011).
Sugino, K. et al. Cell-type-specific repression by methyl-CpG-binding protein 2 is biased toward long genes. J. Neurosci. 34, 12877–12883 (2014).
Maniatis, T. & Reed, R. An extensive network of coupling among gene expression machines. Nature 416, 499–506 (2002).
Core, L.J., Waterfall, J.J. & Lis, J.T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Brennan, C.M. & Steitz, J.A. HuR and mRNA stability. Cell. Mol. Life Sci. 58, 266–277 (2001).
Höck, J. & Meister, G. The Argonaute protein family. Genome Biol. 9, 210 (2008).
Flavell, S.W. & Greenberg, M.E. Signaling mechanisms linking neuronal activity to gene expression and plasticity of the nervous system. Annu. Rev. Neurosci. 31, 563–590 (2008).
Yang, H. et al. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154, 1370–1379 (2013).
Müller, M. & Can, K. Aberrant redox homoeostasis and mitochondrial dysfunction in Rett syndrome. Biochem. Soc. Trans. 42, 959–964 (2014).
Zylka, M.J., Simon, J.M. & Philpot, B.D. Gene length matters in neurons. Neuron 86, 353–355 (2015).
Linhoff, M.W., Garg, S.K. & Mandel, G. A high-resolution imaging approach to investigate chromatin architecture in complex tissues. Cell 163, 246–255 (2015).
King, I.F. et al. Topoisomerases facilitate transcription of long genes linked to autism. Nature 501, 58–62 (2013).
Nott, A. et al. Histone deacetylase 3 associates with MeCP2 to regulate FOXO and social behavior. Nat. Neurosci. 19, 1497–1505; advance online publication (2016).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Buxbaum, A.R., Yoon, Y.J., Singer, R.H. & Park, H.Y. Single-molecule insights into mRNA dynamics in neurons. Trends Cell Biol. 25, 468–475 (2015).
Mauger, O., Lemoine, F. & Scheiffele, P. Targeted intron retention and excision for rapid gene regulation in response to neuronal activity. Neuron 92, 1266–1278 (2016).
Khwaja, O.S. et al. Safety, pharmacokinetics, and preliminary assessment of efficacy of mecasermin (recombinant human IGF-1) for the treatment of Rett syndrome. Proc. Natl. Acad. Sci. USA 111, 4596–4601 (2014).
Lombardi, L.M., Baker, S.A. & Zoghbi, H.Y. MECP2 disorders: from the clinic to mice and back. J. Clin. Invest. 125, 2914–2923 (2015).
Xiao, C. et al. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell 131, 146–159 (2007).
de Boer, E. et al. Efficient biotinylation and single-step purification of tagged transcription factors in mammalian cells and transgenic mice. Proc. Natl. Acad. Sci. USA 100, 7480–7485 (2003).
Driegen, S. et al. A generic tool for biotinylation of tagged proteins in transgenic mice. Transgenic Res. 14, 477–482 (2005).
Greer, C.B. et al. Histone deacetylases positively regulate transcription through the elongation machinery. Cell Rep. 13, 1444–1455 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Merico, D., Isserlin, R., Stueker, O., Emili, A. & Bader, G.D. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS One 5, e13984 (2010).
Smoot, M.E., Ono, K., Ruscheinski, J., Wang, P.-L. & Ideker, T. Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27, 431–432 (2011).
Hart, T., Komori, H.K., LaMere, S., Podshivalova, K. & Salomon, D.R. Finding the active genes in deep RNA-seq gene expression studies. BMC Genomics 14, 778 (2013).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2014).
Acknowledgements
We would like to thank the IDDRC Mouse Gene Manipulation Core at Children's Hospital Boston (U54HD090255, M. Thompson), the Gene Targeting Core (P01DK049210, K. Kaestner) and the Transgenic and Chimeric Mouse Facility (J. Richa) at University of Pennsylvania for help in generating transgenic mice, the Flow Cytometry and Cell Sorting Resource Laboratory (H. Pletcher, W. DeMuth), and the Next Generation Sequencing Core (J. Schug) for technical assistance. B.S.J. is supported by a Cell and Molecular Biology Training Grant (TG32GM072290) and the UNCF/Merck Graduate Research Dissertation Fellowship. This work is supported by NIH grants K22AI112570 (G.V.), R21AI107067 and R01CA140485 (T.H.K.), R01MH091850 and R01NS081054 (Z.Z.), and a basic research grant from Rettsyndrome.org (Z.Z.). Z.Z. is a Pew Scholar in the Biomedical Sciences.
Author information
Authors and Affiliations
Contributions
Conceptualization, B.S.J. and Z.Z.; methodology, B.S.J., J.M.L., D.G. and Z.Z.; investigation, B.S.J., Y.-T.Z., M.F., J.M.L., K.H.W., Y.J.K. and D.B.; formal analyses, B.S.J., Y.-T.Z., G.G. and T.H.K.; validation, B.S.J., M.F., J.M.L. and G.V.; resources, B.S.J., Y.-T.Z. and Y.C.; data curation, Y.-T.Z.; writing manuscript, B.S.J.; review and editing, B.S.J., Y.-T.Z., M.F. and Z.Z.; visualization, B.S.J.; project administration and funding acquisition, Z.Z.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 (PDF 29209 kb)
Supplementary Table 1
Summary of RNA-seq experimental conditions used in this study (XLSX 14 kb)
Supplementary Table 2
RT-PCR primers used in this study (XLSX 65 kb)
Supplementary Table 3
List of HITS-CLIP data sets used for RBP analysis (XLSX 55 kb)
Rights and permissions
About this article
Cite this article
Johnson, B., Zhao, YT., Fasolino, M. et al. Biotin tagging of MeCP2 in mice reveals contextual insights into the Rett syndrome transcriptome. Nat Med 23, 1203–1214 (2017). https://doi.org/10.1038/nm.4406
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4406
This article is cited by
-
Overcoming genetic and cellular complexity to study the pathophysiology of X-linked intellectual disabilities
Journal of Neurodevelopmental Disorders (2024)
-
Buffering of transcription rate by mRNA half-life is a conserved feature of Rett syndrome models
Nature Communications (2023)
-
Joint single-cell profiling resolves 5mC and 5hmC and reveals their distinct gene regulatory effects
Nature Biotechnology (2023)
-
Neuronal Yin Yang1 in the prefrontal cortex regulates transcriptional and behavioral responses to chronic stress in mice
Nature Communications (2022)
-
Rett syndrome linked to defects in forming the MeCP2/Rbfox/LASR complex in mouse models
Nature Communications (2021)