Abstract
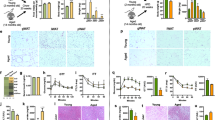
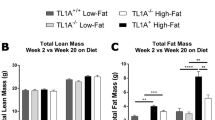
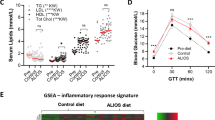
Nonalcoholic fatty liver disease (NAFLD), a common prelude to cirrhosis and hepatocellular carcinoma, is the most common chronic liver disease worldwide. Defining the molecular mechanisms underlying the pathogenesis of NAFLD has been hampered by a lack of animal models that closely recapitulate the severe end of the disease spectrum in humans, including bridging hepatic fibrosis. Here we demonstrate that a novel experimental model employing thermoneutral housing, as opposed to standard housing, resulted in lower stress-driven production of corticosterone, augmented mouse proinflammatory immune responses and markedly exacerbated high-fat diet (HFD)-induced NAFLD pathogenesis. Disease exacerbation at thermoneutrality was conserved across multiple mouse strains and was associated with augmented intestinal permeability, an altered microbiome and activation of inflammatory pathways that are associated with the disease in humans. Depletion of Gram-negative microbiota, hematopoietic cell deletion of Toll-like receptor 4 (TLR4) and inactivation of the IL-17 axis resulted in altered immune responsiveness and protection from thermoneutral-housing-driven NAFLD amplification. Finally, female mice, typically resistant to HFD-induced obesity and NAFLD, develop full disease characteristics at thermoneutrality. Thus, thermoneutral housing provides a sex-independent model of exacerbated NAFLD in mice and represents a novel approach for interrogation of the cellular and molecular mechanisms underlying disease pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Change history
21 June 2017
In the version of this article initially published online, a grant supporting the authors’ work was omitted from the Acknowledgments section. The grant “NIH T32AI118697 (associated with D.A.G.)” has now been added. The error has been corrected in the print, PDF and HTML versions of this article.
References
Ma, C. et al. NAFLD causes selective CD4+ T lymphocyte loss and promotes hepatocarcinogenesis. Nature 531, 253–257 (2016).
Rinella, M.E. Will the increased prevalence of nonalcoholic steatohepatitis (NASH) in the age of better hepatitis C virus therapy make NASH the deadlier disease? Hepatology 54, 1118–1120 (2011).
Tiniakos, D.G., Vos, M.B. & Brunt, E.M. Nonalcoholic fatty liver disease: pathology and pathogenesis. Annu. Rev. Pathol. 5, 145–171 (2010).
Rahman, K. et al. Loss of junctional adhesion molecule A promotes severe steatohepatitis in mice on a diet high in saturated fat, fructose, and cholesterol. Gastroenterology 151, 733–746 (2016).
Mehal, W.Z. The Gordian Knot of dysbiosis, obesity and NAFLD. Nat. Rev. Gastroenterol. Hepatol. 10, 637–644 (2013).
Zhu, L. et al. Characterization of gut microbiomes in nonalcoholic steatohepatitis (NASH) patients: a connection between endogenous alcohol and NASH. Hepatology 57, 601–609 (2013).
Miele, L. et al. Increased intestinal permeability and tight junction alterations in nonalcoholic fatty liver disease. Hepatology 49, 1877–1887 (2009).
Vespasiani-Gentilucci, U. et al. Hepatic Toll-like receptor 4 expression is associated with portal inflammation and fibrosis in patients with NAFLD. Liver Int. 35, 569–581 (2015).
Kiziltas, S. et al. TLR4 gene polymorphism in patients with nonalcoholic fatty liver disease in comparison to healthy controls. Metab. Syndr. Relat. Disord. 12, 165–170 (2014).
Mills, K.H. TLR-dependent T cell activation in autoimmunity. Nat. Rev. Immunol. 11, 807–822 (2011).
Harley, I.T. et al. IL-17 signaling accelerates the progression of nonalcoholic fatty liver disease in mice. Hepatology 59, 1830–1839 (2014).
Rau, M. et al. Progression from nonalcoholic fatty liver to nonalcoholic steatohepatitis is marked by a higher frequency of Th17 cells in the liver and an increased Th17/resting regulatory T cell ratio in peripheral blood and in the liver. J. Immunol. 196, 97–105 (2016).
Hebbard, L. & George, J. Animal models of nonalcoholic fatty liver disease. Nat. Rev. Gastroenterol. Hepatol. 8, 35–44 (2011).
Karp, C.L. Unstressing intemperate models: how cold stress undermines mouse modeling. J. Exp. Med. 209, 1069–1074 (2012).
Gordon, C.J. Temperature Regulation in Laboratory Rodents (Cambridge University Press, 1993).
Maloney, S.K., Fuller, A., Mitchell, D., Gordon, C. & Overton, J.M. Translating animal model research: does it matter that our rodents are cold? Physiology (Bethesda) 29, 413–420 (2014).
Stemmer, K. et al. Thermoneutral housing is a critical factor for immune function and diet-induced obesity in C57BL/6 nude mice. Int. J. Obes. (Lond) 39, 791–797 (2015).
Bowers, S.L., Bilbo, S.D., Dhabhar, F.S. & Nelson, R.J. Stressor-specific alterations in corticosterone and immune responses in mice. Brain Behav. Immun. 22, 105–113 (2008).
Giles, D.A. et al. Modulation of ambient temperature promotes inflammation and initiates atherosclerosis in wild type C57BL/6 mice. Mol. Metab. 5, 1121–1130 (2016).
Rudaya, A.Y., Steiner, A.A., Robbins, J.R., Dragic, A.S. & Romanovsky, A.A. Thermoregulatory responses to lipopolysaccharide in the mouse: dependence on the dose and ambient temperature. Am. J. Physiol. Regul. Integr. Comp. Physiol. 289, R1244–R1252 (2005).
Moragues, V. & Pinkerton, H. Variation in morbidity and mortality of murine typhus infection in mice with changes in the environmental temperature. J. Exp. Med. 79, 41–43 (1944).
Foxman, E.F. et al. Temperature-dependent innate defense against the common cold virus limits viral replication at warm temperature in mouse airway cells. Proc. Natl. Acad. Sci. USA 112, 827–832 (2015).
Eng, J.W. et al. Housing temperature–induced stress drives therapeutic resistance in murine tumour models through β2-adrenergic receptor activation. Nat. Commun. 6, 6426 (2015).
Tian, X.Y. et al. Thermoneutral housing accelerates metabolic inflammation to potentiate atherosclerosis but not insulin resistance. Cell Metab. 23, 165–178 (2016).
Athyros, V.G. et al. Cardiovascular risk across the histological spectrum and the clinical manifestations of non-alcoholic fatty liver disease: an update. World J. Gastroenterol. 21, 6820–6834 (2015).
Kox, M. et al. Voluntary activation of the sympathetic nervous system and attenuation of the innate immune response in humans. Proc. Natl. Acad. Sci. USA 111, 7379–7384 (2014).
Hanssen, M.J. et al. Short-term cold acclimation improves insulin sensitivity in patients with type 2 diabetes mellitus. Nat. Med. 21, 863–865 (2015).
Overton, J.M. Phenotyping small animals as models for the human metabolic syndrome: thermoneutrality matters. Int. J. Obes. (Lond) 34 (Suppl. 2), S53–S58 (2010).
Swoap, S.J. et al. Vagal tone dominates autonomic control of mouse heart rate at thermoneutrality. Am. J. Physiol. Heart Circ. Physiol. 294, H1581–H1588 (2008).
Emre, Y. & Nübel, T. Uncoupling protein UCP2: when mitochondrial activity meets immunity. FEBS Lett. 584, 1437–1442 (2010).
Bhattacharyya, S., Brown, D.E., Brewer, J.A., Vogt, S.K. & Muglia, L.J. Macrophage glucocorticoid receptors regulate Toll-like receptor 4–mediated inflammatory responses by selective inhibition of p38 MAP kinase. Blood 109, 4313–4319 (2007).
Izeboud, C.A., Mocking, J.A., Monshouwer, M., van Miert, A.S. & Witkamp, R.F. Participation of β-adrenergic receptors on macrophages in modulation of LPS-induced cytokine release. J. Recept. Signal Transduct. Res. 19, 191–202 (1999).
Emre, Y. et al. Mitochondria contribute to LPS-induced MAPK activation via uncoupling protein UCP2 in macrophages. Biochem. J. 402, 271–278 (2007).
Wahle, M. et al. β2-adrenergic receptors mediate the differential effects of catecholamines on cytokine production of PBMC. J. Interferon Cytokine Res. 25, 384–394 (2005).
Xu, H. et al. Chronic inflammation in fat plays a crucial role in the development of obesity-related insulin resistance. J. Clin. Invest. 112, 1821–1830 (2003).
Brunt, E.M., Kleiner, D.E., Wilson, L.A., Belt, P. & Neuschwander-Tetri, B.A. Nonalcoholic fatty liver disease (NAFLD) activity score and the histopathologic diagnosis in NAFLD: distinct clinicopathologic meanings. Hepatology 53, 810–820 (2011).
Nagaya, T. et al. Down-regulation of SREBP-1c is associated with the development of burned-out NASH. J. Hepatol. 53, 724–731 (2010).
Rizki, G. et al. Mice fed a lipogenic methionine-choline-deficient diet develop hypermetabolism coincident with hepatic suppression of SCD-1. J. Lipid Res. 47, 2280–2290 (2006).
Teufel, A. et al. Comparison of gene expression patterns between mouse models of nonalcoholic fatty liver disease and liver tissues from patients. Gastroenterology 151, 513–525 (2016).
Brown, M.P. et al. Knowledge-based analysis of microarray gene expression data by using support vector machines. Proc. Natl. Acad. Sci. USA 97, 262–267 (2000).
Alexander, J., Chang, G.Q., Dourmashkin, J.T. & Leibowitz, S.F. Distinct phenotypes of obesity-prone AKR/J, DBA2J and C57BL/6J mice compared to control strains. Int. J. Obes. (Lond) 30, 50–59 (2006).
Li, L., Chen, L. & Hu, L. Nuclear factor high-mobility group box1 mediating the activation of Toll-like receptor 4 signaling in hepatocytes in the early stage of non-alcoholic fatty liver disease in mice. J. Clin. Exp. Hepatol. 1, 123–124 (2011).
Wieckowska, A. et al. Increased hepatic and circulating interleukin-6 levels in human nonalcoholic steatohepatitis. Am. J. Gastroenterol. 103, 1372–1379 (2008).
Vandanmagsar, B. et al. The NLRP3 inflammasome instigates obesity-induced inflammation and insulin resistance. Nat. Med. 17, 179–188 (2011).
Langrish, C.L. et al. IL-23 drives a pathogenic T cell population that induces autoimmune inflammation. J. Exp. Med. 201, 233–240 (2005).
Ramesh, R. et al. Pro-inflammatory human Th17 cells selectively express P-glycoprotein and are refractory to glucocorticoids. J. Exp. Med. 211, 89–104 (2014).
Sano, T. et al. An IL-23R/IL-22 circuit regulates epithelial serum amyloid A to promote local effector Th17 responses. Cell 163, 381–393 (2015).
Pan, J.J. & Fallon, M.B. Gender and racial differences in nonalcoholic fatty liver disease. World J. Hepatol. 6, 274–283 (2014).
Kanuri, G. & Bergheim, I. In vitro and in vivo models of non-alcoholic fatty liver disease (NAFLD). Int. J. Mol. Sci. 14, 11963–11980 (2013).
Bäckhed, F., Manchester, J.K., Semenkovich, C.F. & Gordon, J.I. Mechanisms underlying the resistance to diet-induced obesity in germ-free mice. Proc. Natl. Acad. Sci. USA 104, 979–984 (2007).
Puddu, A., Sanguineti, R., Montecucco, F. & Viviani, G.L. Evidence for the gut microbiota short-chain fatty acids as key pathophysiological molecules improving diabetes. Mediators Inflamm. 2014, 162021 (2014).
Inra, C.N. et al. A perisinusoidal niche for extramedullary haematopoiesis in the spleen. Nature 527, 466–471 (2015).
Alisi, A. et al. Plasma high mobility group box 1 protein reflects fibrosis in pediatric nonalcoholic fatty liver disease. Expert Rev. Mol. Diagn. 14, 763–771 (2014).
do Nascimento, J.H., Epifanio, M., Soder, R.B. & Baldisserotto, M. MRI-diagnosed nonalcoholic fatty liver disease is correlated to insulin resistance in adolescents. Acad. Radiol. 20, 1436–1442 (2013).
Sorrentino, P. et al. Predicting fibrosis worsening in obese patients with NASH through parenchymal fibronectin, HOMA-IR, and hypertension. Am. J. Gastroenterol. 105, 336–344 (2010).
Jin, W. & Dong, C. IL-17 cytokines in immunity and inflammation. Emerg. Microbes Infect. 2, e60 (2013).
Meng, F. et al. Interleukin-17 signaling in inflammatory, Kupffer cells, and hepatic stellate cells exacerbates liver fibrosis in mice. Gastroenterology 143, 765–776 (2012).
Giles, D.A., Moreno-Fernandez, M.E. & Divanovic, S. IL-17 axis driven inflammation in non-alcoholic fatty liver disease progression. Curr. Drug Targets 16, 1315–1323 (2015).
McKee, C. et al. Propranolol, a β-adrenoceptor antagonist, worsens liver injury in a model of non-alcoholic steatohepatitis. Biochem. Biophys. Res. Commun. 437, 597–602 (2013).
McAlees, J.W. et al. Distinct Tlr4-expressing cell compartments control neutrophilic and eosinophilic airway inflammation. Mucosal Immunol. 8, 863–873 (2015).
Komiyama, Y. et al. IL-17 plays an important role in the development of experimental autoimmune encephalomyelitis. J. Immunol. 177, 566–573 (2006).
Giles, D.A. et al. Regulation of inflammation by IL-17A and IL-17F modulates non-alcoholic fatty liver disease pathogenesis. PLoS One 11, e0149783 (2016).
Chen, J., Bardes, E.E., Aronow, B.J. & Jegga, A.G. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 37, W305–W311 (2009).
Divanovic, S., Trompette, A., Ashworth, J.I., Rao, M.B. & Karp, C.L. Therapeutic enhancement of protective immunity during experimental leishmaniasis. PLoS Negl. Trop. Dis. 5, e1316 (2011).
Finkelman, F., Morris, S., Orekhova, T. & Sehy, D. The in vivo cytokine capture assay for measurement of cytokine production in the mouse. Curr. Protoc. Immunol. 54, 6.28 (2003).
Divanovic, S. et al. Negative regulation of Toll-like receptor 4 signaling by the Toll-like receptor homolog RP105. Nat. Immunol. 6, 571–578 (2005).
Castañeda, T.R. et al. Metabolic control by S6 kinases depends on dietary lipids. PLoS One 7, e32631 (2012).
Wu, D. et al. Interleukin-13 (IL-13)/IL-13 receptor alpha1 (IL-13Rα1) signaling regulates intestinal epithelial cystic fibrosis transmembrane conductance regulator channel-dependent Cl− secretion. J. Biol. Chem. 286, 13357–13369 (2011).
Schloss, P.D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Caporaso, J.G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Acknowledgements
This work was supported in part by NIH R01DK099222 and R01DK099222-S1 (to S.D.), the CCHMC Pediatric Diabetes and Obesity Center initiative (to S.D.), R01DK033201 (to C.R.K.), K12-HD000850 (to S.S.), NIH T32AI118697 (associated with D.A.G.), NIEHS Grant P30 ES006096 University of Cincinnati Center for Environmental Genomics (associated with D.A.G.), PHS Grant P30 DK078392 Pathology of the Digestive Disease Research Core Center at CCHMC (associated with S.D.) and German Research Foundation IRTG 1911 (projects A6 and B8 to C.S. and J.R.). We would also like to acknowledge C. Chougnet (CCHMC) for providing access to human PBMC samples and C. Woods (CCHMC) for technical assistance.
Author information
Authors and Affiliations
Contributions
D.A.G., M.E.M.-F., T.E.S., S.G., M.C., D.W., R.M., C.C.C., M.J.L., J.K., S.S., A.S. and D.R. participated in data generation. D.A.G., M.E.M.-F., S.G., D.R., R.K., B.J.A., S.K.S., R.S., K.A.S., D.B.H., J.R., S.P.H. and S.D. participated in data analysis and interpretation. S.S., K.S., Y.I., C.R.K., B.J.A., C.S. and C.L.K. provided materials and technical support and participated in critical review of the manuscript. D.A.G., S.S., C.R.K., J.R., C.S. and S.D. obtained the funding. D.A.G. and S.D. participated in the conception and design of the study, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures and Table
Supplementary Figures 1–12 and Supplementary Table 1 (PDF 2090 kb)
Rights and permissions
About this article
Cite this article
Giles, D., Moreno-Fernandez, M., Stankiewicz, T. et al. Thermoneutral housing exacerbates nonalcoholic fatty liver disease in mice and allows for sex-independent disease modeling. Nat Med 23, 829–838 (2017). https://doi.org/10.1038/nm.4346
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4346
This article is cited by
-
Sex differences in energy metabolism: natural selection, mechanisms and consequences
Nature Reviews Nephrology (2024)
-
TREM-2 Drives Development of Multiple Sclerosis by Promoting Pathogenic Th17 Polarization
Neuroscience Bulletin (2024)
-
An overview of mouse models of hepatocellular carcinoma
Infectious Agents and Cancer (2023)
-
Discrimination of cell-intrinsic and environment-dependent effects of natural genetic variation on Kupffer cell epigenomes and transcriptomes
Nature Immunology (2023)
-
Microbially produced vitamin B12 contributes to the lipid-lowering effect of silymarin
Nature Communications (2023)