Abstract
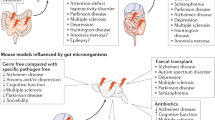
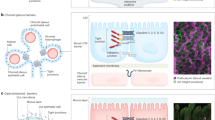
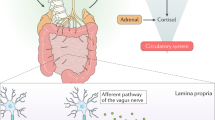
The ecosystem of the human gut consists of trillions of bacteria forming a bioreactor that is fueled by dietary macronutrients to produce bioactive compounds. These microbiota-derived metabolites signal to distant organs in the body, which enables the gut bacteria to connect to the immune and hormone system, to the brain (the gut–brain axis) and to host metabolism, as well as other functions of the host. This microbe–host communication is essential to maintain vital functions of the healthy host. Recently, however, the gut microbiota has been associated with a number of diseases, ranging from obesity and inflammatory diseases to behavioral and physiological abnormalities associated with neurodevelopmental disorders. In this Review, we will discuss microbiota–host cross-talk and intestinal microbiome signaling to extraintestinal organs. We will review mechanisms of how this communication might contribute to host physiology and discuss how misconfigured signaling might contribute to different diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sender, R., Fuchs, S. & Milo, R. Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell 164, 337–340 (2016).
Zoetendal, E.G., Akkermans, A.D. & De Vos, W.M. Temperature gradient gel electrophoresis analysis of 16S rRNA from human fecal samples reveals stable and host-specific communities of active bacteria. Appl. Environ. Microbiol. 64, 3854–3859 (1998).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Eckburg, P.B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638 (2005).
Hold, G.L., Pryde, S.E., Russell, V.J., Furrie, E. & Flint, H.J. Assessment of microbial diversity in human colonic samples by 16S rDNA sequence analysis. FEMS Microbiol. Ecol. 39, 33–39 (2002).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 32, 834–841 (2014).
Tamburini, S., Shen, N., Wu, H.C. & Clemente, J.C. The microbiome in early life: implications for health outcomes. Nat. Med. 22, 713–722 (2016).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Knights, D. et al. Rethinking “enterotypes”. Cell Host Microbe. 16, 433–437 (2014).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
Kovatcheva-Datchary, P. et al. Dietary fiber-induced improvement in glucose metabolism is associated with increased abundance of Prevotella. Cell Metab. 22, 971–982 (2015).
Kommineni, S. et al. Bacteriocin production augments niche competition by enterococci in the mammalian gastrointestinal tract. Nature 526, 719–722 (2015).
Pédron, T. et al. A crypt-specific core microbiota resides in the mouse colon. MBio 3, e00116–e00112 (2012).
Kaiko, G.E. et al. The colonic crypt protects stem cells from microbiota-derived metabolites. Cell 165, 1708–1720 (2016).
Haag, L.-M. & Siegmund, B. Intestinal microbiota and the innate immune system—a crosstalk in Crohn's disease pathogenesis. Front. Immunol. 6, 489 (2015).
Honda, K. & Littman, D.R. The microbiota in adaptive immune homeostasis and disease. Nature 535, 75–84 (2016).
Thaiss, C.A., Zmora, N., Levy, M. & Elinav, E. The microbiome and innate immunity. Nature 535, 65–74 (2016).
Medzhitov, R. Recognition of microorganisms and activation of the immune response. Nature 449, 819–826 (2007).
Cani, P.D. et al. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 56, 1761–1772 (2007).
Hersoug, L.-G., Møller, P. & Loft, S. Gut microbiota-derived lipopolysaccharide uptake and trafficking to adipose tissue: implications for inflammation and obesity. Obes. Rev. 17, 297–312 (2016).
Ghoshal, S., Witta, J., Zhong, J., de Villiers, W. & Eckhardt, E. Chylomicrons promote intestinal absorption of lipopolysaccharides. J. Lipid Res. 50, 90–97 (2009).
Bäckhed, F., Normark, S., Schweda, E.K.H., Oscarson, S. & Richter-Dahlfors, A. Structural requirements for TLR4-mediated LPS signalling: a biological role for LPS modifications. Microbes Infect. 5, 1057–1063 (2003).
Vatanen, T. et al. Variation in microbiome LPS mmunogenicity Contributes to Autoimmunity in Humans. Cell 165, 842–853 (2016).
Larsson, E. et al. Analysis of gut microbial regulation of host gene expression along the length of the gut and regulation of gut microbial ecology through MyD88. Gut. 61, 1124–1131 (2012).
Gaboriau-Routhiau, V. et al. The key role of segmented filamentous bacteria in the coordinated maturation of gut helper T cell responses. Immunity 31, 677–689 (2009).
Chung, H. et al. Gut immune maturation depends on colonization with a host-specific microbiota. Cell 149, 1578–1593 (2012).
Kashyap, P.C. et al. Complex interactions among diet, gastrointestinal transit, and gut microbiota in humanized mice. Gastroenterology 144, 967–977 (2013).
Zelante, T. et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 39, 372–385 (2013).
Lamas, B. et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 22, 598–605 (2016).
Bergman, E.N. Energy contributions of volatile fatty acids from the gastrointestinal tract in various species. Physiol. Rev. 70, 567–590 (1990).
Bäckhed, F. et al. The gut microbiota as an environmental factor that regulates fat storage. Proc. Natl. Acad. Sci. USA 101, 15718–15723 (2004).
Bäckhed, F., Manchester, J.K., Semenkovich, C.F. & Gordon, J.I. Mechanisms underlying the resistance to diet-induced obesity in germ-free mice. Proc. Natl. Acad. Sci. USA 104, 979–984 (2007).
Samuel, B.S. et al. Effects of the gut microbiota on host adiposity are modulated by the short-chain fatty-acid binding G protein-coupled receptor, Gpr41. Proc. Natl. Acad. Sci. USA 105, 16767–16772 (2008).
Tolhurst, G. et al. Short-chain fatty acids stimulate glucagon-like peptide-1 secretion via the G-protein-coupled receptor FFAR2. Diabetes 61, 364–371 (2012).
Fava, S. Glucagon-like peptide 1 and the cardiovascular system. Curr. Diabetes Rev. 10, 302–310 (2014).
Nøhr, M.K. et al. GPR41/FFAR3 and GPR43/FFAR2 as cosensors for short-chain fatty acids in enteroendocrine cells vs FFAR3 in enteric neurons and FFAR2 in enteric leukocytes. Endocrinology 154, 3552–3564 (2013).
De Vadder, F. et al. Microbiota-generated metabolites promote metabolic benefits via gut-brain neural circuits. Cell 156, 84–96 (2014).
Frost, G. et al. The short-chain fatty acid acetate reduces appetite via a central homeostatic mechanism. Nat. Commun. 5, 3611 (2014).
Perry, R.J. et al. Acetate mediates a microbiome-brain-β-cell axis to promote metabolic syndrome. Nature 534, 213–217 (2016).
Kimura, I. et al. The gut microbiota suppresses insulin-mediated fat accumulation via the short-chain fatty acid receptor GPR43. Nat. Commun. 4, 1829 (2013).
Midtvedt, T. Microbial bile acid transformation. Am. J. Clin. Nutr. 27, 1341–1347 (1974).
Swann, J.R. et al. Systemic gut microbial modulation of bile acid metabolism in host tissue compartments. Proc. Natl. Acad. Sci. USA 108 (Suppl. 1), 4523–4530 (2011).
Sayin, S.I. et al. Gut microbiota regulates bile acid metabolism by reducing the levels of tauro-beta-muricholic acid, a naturally occurring FXR antagonist. Cell Metab. 17, 225–235 (2013).
Islam, K.B.M.S. et al. Bile acid is a host factor that regulates the composition of the cecal microbiota in rats. Gastroenterology 141, 1773–1781 (2011).
Kurdi, P., Kawanishi, K., Mizutani, K. & Yokota, A. Mechanism of growth inhibition by free bile acids in lactobacilli and bifidobacteria. J. Bacteriol. 188, 1979–1986 (2006).
Li, F. et al. Microbiome remodelling leads to inhibition of intestinal farnesoid X receptor signalling and decreased obesity. Nat. Commun. 4, 2384 (2013).
Jiang, C. et al. Intestine-selective farnesoid X receptor inhibition improves obesity-related metabolic dysfunction. Nat. Commun. 6, 10166 (2015).
Parséus, A. et al. Microbiota-induced obesity requires farnesoid X receptor. Gut. http://dx.doi.org/10.1136/gutjnl-2015-310283 (2016).
Thomas, C. et al. TGR5-mediated bile acid sensing controls glucose homeostasis. Cell Metab. 10, 167–177 (2009).
Broeders, E.P.M. et al. The bile acid chenodeoxycholic acid increases human brown adipose tissue Activity. Cell Metab. 22, 418–426 (2015).
Ryan, K.K. et al. FXR is a molecular target for the effects of vertical sleeve gastrectomy. Nature 509, 183–188 (2014).
Tremaroli, V. et al. Roux-en-Y gastric bypass and vertical banded gastroplasty induce long-term changes on the human gut microbiome contributing to fat mass regulation. Cell Metab. 22, 228–238 (2015).
Schauer, P.R. et al. Bariatric surgery versus intensive medical therapy for diabetes—3-year outcomes. N. Engl. J. Med. 370, 2002–2013 (2014).
McGavigan, A.K. et al. TGR5 contributes to glucoregulatory improvements after vertical sleeve gastrectomy in mice. Gut. http://dx.doi.org/10.1136/gutjnl-2015-309871 (2015).
Fang, S. et al. Intestinal FXR agonism promotes adipose tissue browning and reduces obesity and insulin resistance. Nat. Med. 21, 159–165 (2015).
Downes, M. et al. A chemical, genetic, and structural analysis of the nuclear bile acid receptor FXR. Mol. Cell 11, 1079–1092 (2003).
Neuschwander-Tetri, B.A. et al. Farnesoid X nuclear receptor ligand obeticholic acid for non-cirrhotic, non-alcoholic steatohepatitis (FLINT): a multicentre, randomised, placebo-controlled trial. Lancet 385, 956–965 (2015).
Cypess, A.M. et al. Identification and importance of brown adipose tissue in adult humans. N. Engl. J. Med. 360, 1509–1517 (2009).
Wang, Q.A., Tao, C., Gupta, R.K. & Scherer, P.E. Tracking adipogenesis during white adipose tissue development, expansion and regeneration. Nat. Med. 19, 1338–1344 (2013).
Harms, M. & Seale, P. Brown and beige fat: development, function and therapeutic potential. Nat. Med. 19, 1252–1263 (2013).
Ziętak, M. et al. Altered microbiota contributes to reduced diet-induced obesity upon cold exposure. Cell Metab. 23, 1216–1223 (2016).
Chevalier, C. et al. Gut microbiota orchestrates energy homeostasis during Cold. Cell 163, 1360–1374 (2015).
Suárez-Zamorano, N. et al. Microbiota depletion promotes browning of white adipose tissue and reduces obesity. Nat. Med. 21, 1497–1501 (2015).
Collins, S. & Surwit, R.S. The beta-adrenergic receptors and the control of adipose tissue metabolism and thermogenesis. Recent Prog. Horm. Res. 56, 309–328 (2001).
Sommer, F. et al. The gut microbiota modulates energy metabolism in the hibernating brown bear Ursus arctos. Cell Rep. 14, 1655–1661 (2016).
Turnbaugh, P.J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Turnbaugh, P.J., Bäckhed, F., Fulton, L. & Gordon, J.I. Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe. 3, 213–223 (2008).
Turnbaugh, P.J. et al. The effect of diet on the human gut microbiome: A metagenomic analysis in humanized gnotobiotic mice. Sci. Transl. Med. 1, 6ra14 (2009).
Blanton, L.V. et al. Gut bacteria that prevent growth impairments transmitted by microbiota from malnourished children. Science 351, aad3311 (2016).
Smith, M.I. et al. Gut microbiomes of Malawian twin pairs discordant for kwashiorkor. Science 339, 548–554 (2013).
Engfer, M.B., Stahl, B., Finke, B., Sawatzki, G. & Daniel, H. Human milk oligosaccharides are resistant to enzymatic hydrolysis in the upper gastrointestinal tract. Am. J. Clin. Nutr. 71, 1589–1596 (2000).
Charbonneau, M.R. et al. Sialylated milk oligosaccharides promote microbiota-dependent growth in models of infant undernutrition. Cell 164, 859–871 (2016).
Subramanian, S. et al. Persistent gut microbiota immaturity in malnourished Bangladeshi children. Nature 510, 417–421 (2014).
Schwarzer, M. et al. Lactobacillus plantarum strain maintains growth of infant mice during chronic undernutrition. Science 351, 854–857 (2016).
Fleissner, C.K. et al. Absence of intestinal microbiota does not protect mice from diet-induced obesity. Br. J. Nutr. 104, 919–929 (2010).
Ding, S. et al. High-fat diet: bacteria interactions promote intestinal inflammation which precedes and correlates with obesity and insulin resistance in mouse. PLoS One 5, e12191 (2010).
Sonnenburg, E.D. et al. Diet-induced extinctions in the gut microbiota compound over generations. Nature 529, 212–215 (2016).
Tremaroli, V. & Bäckhed, F. Functional interactions between the gut microbiota and host metabolism. Nature 489, 242–249 (2012).
Caesar, R. et al. Gut-derived lipopolysaccharide augments adipose macrophage accumulation but is not essential for impaired glucose or insulin tolerance in mice. Gut 61, 1701–1707 (2012).
Caesar, R., Tremaroli, V., Kovatcheva-Datchary, P., Cani, P.D. & Bäckhed, F. Crosstalk between gut microbiota and dietary lipids aggravates WAT inflammation through TLR signaling. Cell Metab. 22, 658–668 (2015).
Luche, E. et al. Metabolic endotoxemia directly increases the proliferation of adipocyte precursors at the onset of metabolic diseases through a CD14-dependent mechanism. Mol. Metab. 2, 281–291 (2013).
Erridge, C., Attina, T., Spickett, C.M. & Webb, D.J. A high-fat meal induces low-grade endotoxemia: evidence of a novel mechanism of postprandial inflammation. Am. J. Clin. Nutr. 86, 1286–1292 (2007).
Amar, J. et al. Intestinal mucosal adherence and translocation of commensal bacteria at the early onset of type 2 diabetes: molecular mechanisms and probiotic treatment. EMBO Mol. Med. 3, 559–572 (2011).
Beyaz, S. et al. High-fat diet enhances stemness and tumorigenicity of intestinal progenitors. Nature 531, 53–58 (2016).
Everard, A. et al. Cross-talk between Akkermansia muciniphila and intestinal epithelium controls diet-induced obesity. Proc. Natl. Acad. Sci. USA 110, 9066–9071 (2013).
Henao-Mejia, J. et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 482, 179–185 (2012).
Elinav, E. et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145, 745–757 (2011).
Bajaj, J.S. et al. Linkage of gut microbiome with cognition in hepatic encephalopathy. Am. J. Physiol. Gastrointest. Liver Physiol. 302, G168–G175 (2012).
Makiura, N. et al. Relationship of Porphyromonas gingivalis with glycemic level in patients with type 2 diabetes following periodontal treatment. Oral Microbiol. Immunol. 23, 348–351 (2008).
Wang, Z. et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–63 (2011).
Koeth, R.A. et al. Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat. Med. 19, 576–585 (2013).
Zhu, W. et al. Gut microbial metabolite TMAO enhances platelet hyperreactivity and thrombosis risk. Cell 165, 111–124 (2016).
Wang, Z. et al. Non-lethal inhibition of gut microbial trimethylamine production for the treatment of atherosclerosis. Cell 163, 1585–1595 (2015).
Jonsson, A.L. & Bäckhed, F. Drug the bug! Cell 163, 1565–1566 (2015).
Luczynski, P. et al. Adult microbiota-deficient mice have distinct dendritic morphological changes: differential effects in the amygdala and hippocampus. Eur. J. Neurosci. http://dx.doi.org/10.1111/ejn.13291 (2016).
Ogbonnaya, E.S. et al. Adult hippocampal neurogenesis is regulated by the microbiome. Biol. Psychiatry 78, e7–e9 (2015).
Hoban, A.E. et al. Regulation of prefrontal cortex myelination by the microbiota. Transl. Psychiatry 6, e774 (2016).
Braniste, V. et al. The gut microbiota influences blood-brain barrier permeability in mice. Sci. Transl. Med. 6, 263ra158 (2014).
Erny, D. et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat. Neurosci. 18, 965–977 (2015).
Sudo, N. et al. Postnatal microbial colonization programs the hypothalamic-pituitary-adrenal system for stress response in mice. J. Physiol. (Lond.) 558, 263–275 (2004).
Bercik, P. et al. The anxiolytic effect of Bifidobacterium longum NCC3001 involves vagal pathways for gut-brain communication. Neurogastroenterol. Motil. 23, 1132–1139 (2011).
Bravo, J.A. et al. Ingestion of Lactobacillus strain regulates emotional behavior and central GABA receptor expression in a mouse via the vagus nerve. Proc. Natl. Acad. Sci. USA 108, 16050–16055 (2011).
Bercik, P. et al. The intestinal microbiota affect central levels of brain-derived neurotropic factor and behavior in mice. Gastroenterology 141, 599–609.e3 (2011).
Clarke, G. et al. The microbiome-gut-brain axis during early life regulates the hippocampal serotonergic system in a sex-dependent manner. Mol. Psychiatry 18, 666–673 (2013).
Hsiao, E.Y. et al. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell 155, 1451–1463 (2013).
Niehus, R. & Lord, C. Early medical history of children with autism spectrum disorders. J. Dev. Behav. Pediatr. JDBP 27 (Suppl.), S120–S127 (2006).
Finegold, S.M. et al. Pyrosequencing study of fecal microflora of autistic and control children. Anaerobe 16, 444–453 (2010).
Wang, L. et al. Elevated fecal short chain fatty acid and ammonia concentrations in children with autism spectrum disorder. Dig. Dis. Sci. 57, 2096–2102 (2012).
Mayer, E.A., Padua, D. & Tillisch, K. Altered brain-gut axis in autism: comorbidity or causative mechanisms? BioEssays 36, 933–939 (2014).
MacFabe, D.F. et al. Neurobiological effects of intraventricular propionic acid in rats: possible role of short chain fatty acids on the pathogenesis and characteristics of autism spectrum disorders. Behav. Brain Res. 176, 149–169 (2007).
Buffington, S.A. et al. Microbial reconstitution reverses maternal diet-induced social and synaptic deficits in offspring. Cell 165, 1762–1775 (2016).
Sullivan, E.L., Nousen, E.K. & Chamlou, K.A. Maternal high fat diet consumption during the perinatal period programs offspring behavior. Physiol. Behav. 123, 236–242 (2014).
Connolly, N. et al. Maternal metabolic risk factors for autism spectrum disorder-An analysis of electronic medical records and linked birth data. Autism Res. 9, 829–837 (2016).
Benakis, C. et al. Commensal microbiota affects ischemic stroke outcome by regulating intestinal γδ T cells. Nat. Med. 22, 516–523; advance online publication (2016).
Niess, J.H. et al. CX3CR1-mediated dendritic cell access to the intestinal lumen and bacterial clearance. Science 307, 254–258 (2005).
Houlden, A. et al. Brain injury induces specific changes in the caecal microbiota of mice via altered autonomic activity and mucoprotein production. Brain Behav. Immun. http://dx.doi.org/10.1016/j.bbi.2016.04.003 (2016).
Arrieta, M.-C., Stiemsma, L.T., Amenyogbe, N., Brown, E.M. & Finlay, B. The intestinal microbiome in early life: health and disease. Front Immunol. 5, 427 (2014).
Russell, S.L. et al. Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma. EMBO Rep. 13, 440–447 (2012).
Arrieta, M.-C. et al. Early infancy microbial and metabolic alterations affect risk of childhood asthma. Sci. Transl. Med. 7, 307ra152 (2015).
Trompette, A. et al. Gut microbiota metabolism of dietary fiber influences allergic airway disease and hematopoiesis. Nat. Med. 20, 159–166 (2014).
Thorburn, A.N. et al. Evidence that asthma is a developmental origin disease influenced by maternal diet and bacterial metabolites. Nat. Commun. 6, 7320 (2015).
Zeevi, D. et al. Personalized nutrition by prediction of Glycemic Responses. Cell 163, 1079–1094 (2015).
Vrieze, A. et al. Transfer of intestinal microbiota from lean donors increases insulin sensitivity in individuals with metabolic syndrome. Gastroenterology 143, 913–916.e7 (2012).
Hohmann, E.L., Ananthakrishnan, A.N. & Deshpande, V. Case records of the Massachusetts General Hospital. Case 25-2014. A 37-year-old man with ulcerative colitis and bloody diarrhea. N. Engl. J. Med. 371, 668–675 (2014).
Alang, N. & Kelly, C.R. Weight gain after fecal microbiota transplantation. Open Forum Infect. Dis. 2, ofv004 (2015).
Abrahamsson, T.R. et al. Low diversity of the gut microbiota in infants with atopic eczema. J. Allergy Clin. Immunol. 129, 434–440.e2 (2012).
Song, H., Yoo, Y., Hwang, J., Na, Y.-C. & Kim, H.S. Faecalibacterium prausnitzii subspecies-level dysbiosis in the human gut microbiome underlying atopic dermatitis. J. Allergy Clin. Immunol. 137, 852–860 (2016).
Hevia, A. et al. Intestinal dysbiosis associated with systemic lupus erythematosus. MBio 5, e01548–14 (2014).
Lepage, P. et al. Twin study indicates loss of interaction between microbiota and mucosa of patients with ulcerative colitis. Gastroenterology 141, 227–236 (2011).
Takahashi, K. et al. Reduced abundance of butyrate-producing bacteria species in the fecal microbial community in Crohn's disease. Digestion 93, 59–65 (2016).
Kostic, A.D. et al. The dynamics of the human infant gut microbiome in development and in progression toward type 1 diabetes. Cell Host Microbe. 17, 260–273 (2015).
Alkanani, A.K. et al. Alterations in intestinal microbiota correlate with susceptibility to type 1 diabetes. Diabetes 64, 3510–3520 (2015).
Chen, J. et al. Multiple sclerosis patients have a distinct gut microbiota compared to healthy controls. Sci. Rep. 6, 28484 (2016).
Tims, S. et al. Microbiota conservation and BMI signatures in adult monozygotic twins. ISME J. 7, 707–717 (2013).
Goodrich, J.K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799 (2014).
Huurre, A. et al. Mode of delivery—effects on gut microbiota and humoral immunity. Neonatology 93, 236–240 (2008).
Dominguez-Bello, M.G. et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc. Natl. Acad. Sci. USA 107, 11971–11975 (2010).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe. 17, 690–703 (2015).
Azad, M.B. et al. Impact of maternal intrapartum antibiotics, method of birth and breastfeeding on gut microbiota during the first year of life: a prospective cohort study. BJOG 123, 983–993 (2016).
Koleva, P.T., Kim, J.-S., Scott, J.A. & Kozyrskyj, A.L. Microbial programming of health and disease starts during fetal life. Birth Defects Res. C Embryo Today 105, 265–277 (2015).
Ley, R.E., Turnbaugh, P.J., Klein, S. & Gordon, J.I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Duncan, S.H. et al. Reduced dietary intake of carbohydrates by obese subjects results in decreased concentrations of butyrate and butyrate-producing bacteria in feces. Appl. Environ. Microbiol. 73, 1073–1078 (2007).
Dethlefsen, L., Huse, S., Sogin, M.L. & Relman, D.A. The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing. PLoS Biol. 6, e280 (2008).
Cotillard, A. et al. Dietary intervention impact on gut microbial gene richness. Nature 500, 585–588 (2013).
David, L.A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014).
David, L.A. et al. Host lifestyle affects human microbiota on daily timescales. Genome Biol. 15, R89 (2014).
Hofstra, J.J. et al. Changes in microbiota during experimental human Rhinovirus infection. BMC Infect. Dis. 15, 336 (2015).
Dethlefsen, L. & Relman, D.A. Incomplete recovery and individualized responses of the human distal gut microbiota to repeated antibiotic perturbation. Proc. Natl. Acad. Sci. USA 108 (Suppl. 1), 4554–4561 (2011).
Acknowledgements
We thank A. Hallén for her assistance with figures and artwork. Work in the authors' laboratory is supported by the Swedish Research Council, the NovoNordisk foundation, Torsten Söderberg's foundation, Swedish Heart Lung Foundation, Göran Gustafsson's foundation, IngaBritt och Arne Lundbergs foundation, Knut and Alice Wallenberg foundation, the FP7-sponsored program METACARDIS, the regional agreement on medical training and clinical research (ALF) between Region Västra Götaland and Sahlgrenska University Hospital. F.B. is a recipient of an ERC Consolidator Grant (European Research Council, Consolidator grant 615362 - METABASE). B.O.S. is a recipient of an FP7 Marie Curie IEF Fellowship (622909 MUCUS AND METABOLISM) from the European Union and a Human Frontier Science Program Long-Term Fellowship (LT000109/2014-l).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Schroeder, B., Bäckhed, F. Signals from the gut microbiota to distant organs in physiology and disease. Nat Med 22, 1079–1089 (2016). https://doi.org/10.1038/nm.4185
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4185
This article is cited by
-
Effect of a multi-strain probiotic mixture consumption on anxiety and depression symptoms induced in adult mice by postnatal maternal separation
Microbiome (2024)
-
Regulation of polysaccharide in Wu‐tou decoction on intestinal microflora and pharmacokinetics of small molecular compounds in AIA rats
Chinese Medicine (2024)
-
Microbes little helpers and suppliers for therapeutic asthma approaches
Respiratory Research (2024)
-
Combined transcriptome and metabolome analysis reveals breed-specific regulatory mechanisms in Dorper and Tan sheep
BMC Genomics (2024)
-
Composition of the colon microbiota in the individuals with inflammatory bowel disease and colon cancer
Folia Microbiologica (2024)