Abstract
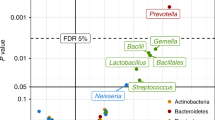
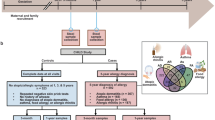
Gut microbiota bacterial depletions and altered metabolic activity at 3 months are implicated in childhood atopy and asthma1. We hypothesized that compositionally distinct human neonatal gut microbiota (NGM) exist, and are differentially related to relative risk (RR) of childhood atopy and asthma. Using stool samples (n = 298; aged 1–11 months) from a US birth cohort and 16S rRNA sequencing, neonates (median age, 35 d) were divisible into three microbiota composition states (NGM1–3). Each incurred a substantially different RR for multisensitized atopy at age 2 years and doctor-diagnosed asthma at age 4 years. The highest risk group, labeled NGM3, showed lower relative abundance of certain bacteria (for example, Bifidobacterium, Akkermansia and Faecalibacterium), higher relative abundance of particular fungi (Candida and Rhodotorula) and a distinct fecal metabolome enriched for pro-inflammatory metabolites. Ex vivo culture of human adult peripheral T cells with sterile fecal water from NGM3 subjects increased the proportion of CD4+ cells producing interleukin (IL)-4 and reduced the relative abundance of CD4+CD25+FOXP3+ cells. 12,13-DiHOME, enriched in NGM3 versus lower-risk NGM states, recapitulated the effect of NGM3 fecal water on relative CD4+CD25+FOXP3+ cell abundance. These findings suggest that neonatal gut microbiome dysbiosis might promote CD4+ T cell dysfunction associated with childhood atopy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Arrieta, M.C. et al. Early infancy microbial and metabolic alterations affect risk of childhood asthma. Sci. Transl. Med. 7, 307ra152 (2015).
Asher, M.I. et al. Worldwide time trends in the prevalence of symptoms of asthma, allergic rhinoconjunctivitis, and eczema in childhood. Lancet 368, 733–743 (2006).
Simpson, A. et al. Beyond atopy: multiple patterns of sensitization in relation to asthma in a birth cohort study. Am. J. Respir. Crit. Care Med. 181, 1200–1206 (2010).
Aichbhaumik, N. et al. Prenatal exposure to household pets influences fetal immunoglobulin E production. Clin. Exp. Allergy 38, 1787–1794 (2008).
Havstad, S. et al. Atopic phenotypes identified with latent class analyses at age 2 years. J. Allergy Clin. Immunol. 134, 722–727.e2 (2014).
Hoffmann, C. et al. Archaea and fungi of the human gut microbiome: correlations with diet and bacterial residents. PLoS One 8, e66019 (2013).
Holmes, I., Harris, K. & Quince, C. Dirichlet multinomial mixtures: generative models for microbial metagenomics. PLoS One 7, e30126 (2012).
Langille, M.G.I. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821 (2013).
Morin, C., Blier, P.U. & Fortin, S. Eicosapentaenoic acid and docosapentaenoic acid monoglycerides are more potent than docosahexaenoic acid monoglyceride to resolve inflammation in a rheumatoid arthritis model. Arthritis Res. Ther. 17, 142 (2015).
Amagai, Y. et al. Dihomo-γ-linolenic acid prevents the development of atopic dermatitis through prostaglandin D1 production in NC/Tnd mice. J. Dermatol. Sci. 79, 30–37 (2015).
Bode, L. Human milk oligosaccharides: every baby needs a sugar mama. Glycobiology 22, 1147–1162 (2012).
Weichert, S. et al. Bioengineered 2′-fucosyllactose and 3-fucosyllactose inhibit the adhesion of Pseudomonas aeruginosa and enteric pathogens to human intestinal and respiratory cell lines. Nutr. Res. 33, 831–838 (2013).
DeAngelis, K.M. et al. Selective progressive response of soil microbial community to wild oat roots. ISME J. 3, 168–178 (2009).
Caporaso, J.G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 108 (Suppl.), 4516–4522 (2011).
Caporaso, J.G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Magocč, T. & Salzberg, S.L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Caporaso, J.G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Edgar, R.C., Haas, B.J., Clemente, J.C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 6, 610–618 (2012).
Caporaso, J.G. et al. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26, 266–267 (2010).
Price, M.N., Dehal, P.S. & Arkin, A.P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS One 5, e9490 (2010).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Edgar, R.C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Abarenkov, K. et al. The UNITE database for molecular identification of fungi–recent updates and future perspectives. New Phytol. 186, 281–285 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. Roy. Stat. Soc. B 57, 289–300 (1995).
Obermaier, B. et al. Development of a new protocol for 2-day generation of mature dendritic cells from human monocytes. Biol. Proced. Online 5, 197–203 (2003).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Vázquez-Baeza, Y., Pirrung, M., Gonzalez, A. & Knight, R. EMPeror: a tool for visualizing high-throughput microbial community data. Gigascience 2, 16 (2013).
Letunic, I. & Bork, P. Interactive Tree Of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res. 39, W475–W478 (2011).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks cytoscape. Genome Res. 13, 2498–2504 (2003).
Acknowledgements
This study was supported by the US National Institutes of Health and National Institute of Allergy and Infectious Diseases P01 AI089473-01 (C.C.J., D.R.O., H.A.B., N.W.L., S.V.L., G.W., and E.M.Z.) and the Alfred P. Sloan Foundation 2013-6-03 (S.V.L.). We thank C. Arrieta and B. Finlay for graciously sharing sequencing data from the CHILD study.
Author information
Authors and Affiliations
Contributions
S.V.L., C.C.J., D.R.O., H.A.B., N.W.L., G.W., and E.M.Z., designed research; C.C.J., D.R.O., K.E.F., D.F., B.L., D.L.L., S.L., A.R.P., E.R., and G.W., performed research; A.R.S., S.H., and A.M.L. contributed new analytic tools; K.E.F., A.R.S., S.H., S.L., A.M.L., and S.V.L. analyzed data; and K.E.F. and S.V.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Tables 1–2, 8–10; Supplementary Note (PDF 4354 kb)
Supplementary Table 3
Association between early life factors and IGMs (XLS 55 kb)
Supplementary Table 4
Association between early life factors and NGMs (XLS 46 kb)
Supplementary Table 5
Factors tested for possible confounding effect on the risk of developing PM atopy for NGM (XLS 50 kb)
Supplementary Table 6
Bacterial taxa exhibiting significantly increased relative abundance in low-risk NGM1 versus the high-risk NGM3 neonatal gut microbiota (XLSX 77 kb)
Supplementary Table 7
Bacterial taxa exhibiting significantly increased relative abundance in low-risk NGM2 versus the high-risk NGM3 neonatal gut microbiota (XLSX 78 kb)
Supplementary Table 11
Metabolites significantly enriched in low-risk NGM1 versus high-risk NGM3 neonatal gut microbiota. (XLS 40 kb)
Supplementary Table 12
Metabolites significantly enriched in low-risk NGM2 versus high-risk NGM3 neonatal gut microbiota (XLS 46 kb)
Rights and permissions
About this article
Cite this article
Fujimura, K., Sitarik, A., Havstad, S. et al. Neonatal gut microbiota associates with childhood multisensitized atopy and T cell differentiation. Nat Med 22, 1187–1191 (2016). https://doi.org/10.1038/nm.4176
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4176
This article is cited by
-
Divergent maturational patterns of the infant bacterial and fungal gut microbiome in the first year of life are associated with inter-kingdom community dynamics and infant nutrition
Microbiome (2024)
-
Microbiome dysbiosis: a modifiable state and target to prevent Staphylococcus aureus infections and other diseases in neonates
Journal of Perinatology (2024)
-
VESPA: an optimized protocol for accurate metabarcoding-based characterization of vertebrate eukaryotic endosymbiont and parasite assemblages
Nature Communications (2024)
-
A comprehensive guide to assess gut mycobiome and its role in pathogenesis and treatment of inflammatory bowel disease
Indian Journal of Gastroenterology (2024)
-
Fecal Microbiota Transplantation Alleviates Allergic Rhinitis via CD4+ T Cell Modulation Through Gut Microbiota Restoration
Inflammation (2024)