Abstract
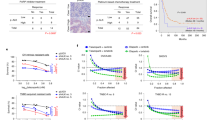
Poly (ADP-ribose) polymerase (PARP) inhibitors have emerged as promising therapeutics for many diseases, including cancer, in clinical trials1. One PARP inhibitor, olaparib (Lynparza, AstraZeneca), was recently approved by the FDA to treat ovarian cancer with mutations in BRCA genes. BRCA1 and BRCA2 have essential roles in repairing DNA double-strand breaks, and a deficiency of BRCA proteins sensitizes cancer cells to PARP inhibition2,3. Here we show that the receptor tyrosine kinase c-Met associates with and phosphorylates PARP1 at Tyr907 (PARP1 pTyr907 or pY907). PARP1 pY907 increases PARP1 enzymatic activity and reduces binding to a PARP inhibitor, thereby rendering cancer cells resistant to PARP inhibition. The combination of c-Met and PARP1 inhibitors synergized to suppress the growth of breast cancer cells in vitro and xenograft tumor models, and we observed similar synergistic effects in a lung cancer xenograft tumor model. These results suggest that the abundance of PARP1 pY907 may predict tumor resistance to PARP inhibitors, and that treatment with a combination of c-Met and PARP inhibitors may benefit patients whose tumors show high c-Met expression and who do not respond to PARP inhibition alone.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
25 August 2016
In the version of this article initially published, the concentrations of H2O2 were incorrectly labeled as micromolar (μM) instead of millimolar (mM) in the legends of Figures 1h, 3c–f and Supplementary Figures 3, 8f–h, 10g,j and 12c,d. The error has been corrected in the HTML and PDF versions of the article.
References
Feng, F.Y., de Bono, J.S., Rubin, M.A. & Knudsen, K.E. Chromatin to clinic: The molecular rationale for PARP1 inhibitor function. Mol. Cell 58, 925–934 (2015).
Bryant, H.E. et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434, 913–917 (2005).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Irani, K. et al. Mitogenic signaling mediated by oxidants in Ras-transformed fibroblasts. Science 275, 1649–1652 (1997).
Trachootham, D., Alexandre, J. & Huang, P. Targeting cancer cells by ROS-mediated mechanisms: a radical therapeutic approach? Nat. Rev. Drug Discov. 8, 579–591 (2009).
Radisky, D.C. et al. Rac1b and reactive oxygen species mediate MMP-3-induced EMT and genomic instability. Nature 436, 123–127 (2005).
Lindahl, T. Instability and decay of the primary structure of DNA. Nature 362, 709–715 (1993).
Schreiber, V., Dantzer, F., Ame, J.C. & de Murcia, G. Poly(ADP-ribose): novel functions for an old molecule. Nat. Rev. Mol. Cell Biol. 7, 517–528 (2006).
Narod, S.A. & Foulkes, W.D. BRCA1 and BRCA2: 1994 and beyond. Nat. Rev. Cancer 4, 665–676 (2004).
Tutt, A. et al. Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and advanced breast cancer: a proof-of-concept trial. Lancet 376, 235–244 (2010).
Lips, E.H. et al. Triple-negative breast cancer: BRCAness and concordance of clinical features with BRCA1 mutation carriers. Br. J. Cancer 108, 2172–2177 (2013).
O'Connor, M.J. Targeting the DNA Damage Response in Cancer. Mol. Cell 60, 547–560 (2015).
Badve, S. et al. Basal-like and triple-negative breast cancers: a critical review with an emphasis on the implications for pathologists and oncologists. Mod. Pathol. 24, 157–167 (2011).
Lehmann, B.D. et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Invest. 121, 2750–2767 (2011).
Gelmon, K.A. et al. Olaparib in patients with recurrent high-grade serous or poorly differentiated ovarian carcinoma or triple-negative breast cancer: a phase 2, multicentre, open-label, non-randomised study. Lancet Oncol. 12, 852–861 (2011).
El-Khamisy, S.F., Masutani, M., Suzuki, H. & Caldecott, K.W. A requirement for PARP-1 for the assembly or stability of XRCC1 nuclear foci at sites of oxidative DNA damage. Nucleic Acids Res. 31, 5526–5533 (2003).
Javle, M. & Curtin, N.J. The role of PARP in DNA repair and its therapeutic exploitation. Br. J. Cancer 105, 1114–1122 (2011).
Swindall, A.F., Stanley, J.A. & Yang, E.S. PARP-1: friend or foe of DNA damage and repair in tumorigenesis? Cancers (Basel) 5, 943–958 (2013).
Maynard, S., Schurman, S.H., Harboe, C., de Souza-Pinto, N.C. & Bohr, V.A. Base excision repair of oxidative DNA damage and association with cancer and aging. Carcinogenesis 30, 2–10 (2009).
Robaszkiewicz, A. et al. Hydrogen peroxide-induced poly(ADP-ribosyl)ation regulates osteogenic differentiation-associated cell death. Free Radic. Biol. Med. 53, 1552–1564 (2012).
Chiarugi, P. & Cirri, P. Redox regulation of protein tyrosine phosphatases during receptor tyrosine kinase signal transduction. Trends Biochem. Sci. 28, 509–514 (2003).
Casaletto, J.B. & McClatchey, A.I. Spatial regulation of receptor tyrosine kinases in development and cancer. Nat. Rev. Cancer 12, 387–400 (2012).
Speers, C. et al. Identification of novel kinase targets for the treatment of estrogen receptor-negative breast cancer. Clin. Cancer Res. 15, 6327–6340 (2009).
Hsu, Y.H. et al. Definition of PKC-α, CDK6, and MET as therapeutic targets in triple-negative breast cancer. Cancer Res. 74, 4822–4835 (2014).
Zagouri, F. et al. High MET expression is an adverse prognostic factor in patients with triple-negative breast cancer. Br. J. Cancer 108, 1100–1105 (2013).
Matteucci, E., Bendinelli, P. & Desiderio, M.A. Nuclear localization of active HGF receptor Met in aggressive MDA-MB231 breast carcinoma cells. Carcinogenesis 30, 937–945 (2009).
Gomes, D.A. et al. c-Met must translocate to the nucleus to initiate calcium signals. J. Biol. Chem. 283, 4344–4351 (2008).
Wang, Y.N. et al. Membrane-bound trafficking regulates nuclear transport of integral epidermal growth factor receptor (EGFR) and ErbB-2. J. Biol. Chem. 287, 16869–16879 (2012).
Wang, Y.N. et al. COPI-mediated retrograde trafficking from the Golgi to the ER regulates EGFR nuclear transport. Biochem. Biophys. Res. Commun. 399, 498–504 (2010).
Wang, Y.N., Yamaguchi, H., Hsu, J.M. & Hung, M.C. Nuclear trafficking of the epidermal growth factor receptor family membrane proteins. Oncogene 29, 3997–4006 (2010).
Du, Y. et al. Syntaxin 6–mediated Golgi translocation plays an important role in nuclear functions of EGFR through microtubule-dependent trafficking. Oncogene 33, 756–770 (2014).
Sonnenblick, A., de Azambuja, E., Azim, H.A. Jr. & Piccart, M. An update on PARP inhibitors--moving to the adjuvant setting. Nat. Rev. Clin. Oncol. 12, 27–41 (2015).
Jagadeeswaran, R., Jagadeeswaran, S., Bindokas, V.P. & Salgia, R. Activation of HGF/c-Met pathway contributes to the reactive oxygen species generation and motility of small cell lung cancer cells. Am. J. Physiol. Lung Cell. Mol. Physiol. 292, L1488–L1494 (2007).
Anders, C.K. et al. Pharmacokinetics and efficacy of PEGylated liposomal doxorubicin in an intracranial model of breast cancer. PLoS One 8, e61359 (2013).
Linding, R. et al. Systematic discovery of in vivo phosphorylation networks. Cell 129, 1415–1426 (2007).
Ruf, A., Mennissier de Murcia, J., de Murcia, G. & Schulz, G.E. Structure of the catalytic fragment of poly(AD-ribose) polymerase from chicken. Proc. Natl. Acad. Sci. USA 93, 7481–7485 (1996).
Knight, J.F. et al. Met synergizes with p53 loss to induce mammary tumors that possess features of claudin-low breast cancer. Proc. Natl. Acad. Sci. USA 110, E1301–E1310 (2013).
Ho-Yen, C.M., Jones, J.L. & Kermorgant, S. The clinical and functional significance of c-Met in breast cancer: a review. Breast Cancer Res. 17, 52 (2015).
Nowsheen, S., Cooper, T., Stanley, J.A. & Yang, E.S. Synthetic-lethal interactions between EGFR and PARP inhibition in human triple negative breast cancer cells. PLoS One 7, e46614 (2012).
Boccaccio, C. & Comoglio, P.M. Invasive growth: a MET-driven genetic programme for cancer and stem cells. Nat. Rev. Cancer 6, 637–645 (2006).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Sokal, R.R. & Michener, C.D. A Statistical Method for Evaluating Systematic Relationships (University of Kansas, 1958).
Chou, T.C. Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res. 70, 440–446 (2010).
Lee, H.J. et al. Tyrosine 370 phosphorylation of ATM positively regulates DNA damage response. Cell Res. 25, 225–236 (2015).
Acknowledgements
This study was funded in part by the following: the US National Institutes of Health grants (nos. CA109311 (M.-C. Hung) and CA099031 (M.-C. Hung), and a Cancer Center Support Grant CA16672); the Susan G. Komen Foundation (SAC100016 (M.-C. Hung)); the Breast Cancer Research Foundation; the Patel Memorial Breast Cancer Endowment Fund; The University of Texas MD Anderson–China Medical University and Hospital Sister Institution Fund; the Ministry of Science and Technology, International Research-intensive Centers of Excellence in Taiwan (I-RiCE; MOST 104-2911-I-002-302 (M.-C. Hung)); the Ministry of Health and Welfare, China Medical University Hospital Cancer Research Center of Excellence (MOHW104-TDU-B-212-124-002 (M.-C. Hung)); the Center for Biological Pathways at The University of Texas MD Anderson Cancer Center; and the Ting Tsung and Wei Fong Chao Research Fund. We also thank S. Patterson of the Department of Scientific Publications at The University of Texas MD Anderson Cancer Center for her editorial assistance.
Author information
Authors and Affiliations
Contributions
Y.D. designed and performed the experiments, analyzed data and wrote the manuscript; H.Y., Y.W., Y.-H.H., W.-C.L., W.-H.Y., P.G.L., G.R.L., W.-C.C., C.-H.C. and M.-K.C. performed experiments and analyzed data; H.-L.W. generated the antibody; M.-C. Hsu, C.-T.C., K.N. and Y.S. performed experiments; Y.W., W-.C.H., C.-L.L. and Y.-C.C. provided patient tissue samples; H.Y. and J.L.H. provided scientific input and wrote the manuscript; M.P. provided transgenic mouse cell lines. G.N.H. provided scientific and clinical input. P.J. provided scientific input. M.-C. Hung supervised the entire project, designed the experiments, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 and Supplementary Tables 1–3 (PDF 2748 kb)
Rights and permissions
About this article
Cite this article
Du, Y., Yamaguchi, H., Wei, Y. et al. Blocking c-Met–mediated PARP1 phosphorylation enhances anti-tumor effects of PARP inhibitors. Nat Med 22, 194–201 (2016). https://doi.org/10.1038/nm.4032
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4032
This article is cited by
-
MUC20 regulated by extrachromosomal circular DNA attenuates proteasome inhibitor resistance of multiple myeloma by modulating cuproptosis
Journal of Experimental & Clinical Cancer Research (2024)
-
PARP inhibitors: enhancing efficacy through rational combinations
British Journal of Cancer (2023)
-
PIN1 and CDK1 cooperatively govern pVHL stability and suppressive functions
Cell Death & Differentiation (2023)
-
Biomarkers beyond BRCA: promising combinatorial treatment strategies in overcoming resistance to PARP inhibitors
Journal of Biomedical Science (2022)
-
PBK drives PARP inhibitor resistance through the TRIM37/NFκB axis in ovarian cancer
Experimental & Molecular Medicine (2022)