Abstract
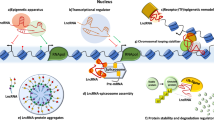
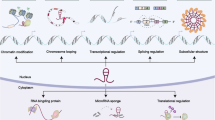
It is increasingly evident that many of the genomic mutations in cancer reside inside regions that do not encode proteins. However, these regions are often transcribed into long noncoding RNAs (lncRNAs). The recent application of next-generation sequencing to a growing number of cancer transcriptomes has indeed revealed thousands of lncRNAs whose aberrant expression is associated with different cancer types. Among the few that have been functionally characterized, several have been linked to malignant transformation. Notably, these lncRNAs have key roles in gene regulation and thus affect various aspects of cellular homeostasis, including proliferation, survival, migration or genomic stability. This review aims to summarize current knowledge of lncRNAs from the cancer perspective. It discusses the strategies that led to the identification of cancer-related lncRNAs and the methodologies and challenges involving the study of these molecules, as well as the imminent applications of these findings to the clinic.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Maurano, M.T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Huarte, M. et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419 (2010).
Zheng, G.X., Do, B.T., Webster, D.E., Khavari, P.A. & Chang, H.Y. Dicer-microRNA-Myc circuit promotes transcription of hundreds of long noncoding RNAs. Nat. Struct. Mol. Biol. 21, 585–590 (2014).
Gupta, R.A. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076 (2010).
Yu, W. et al. Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. Nature 451, 202–206 (2008).
Yap, K.L. et al. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol. Cell 38, 662–674 (2010).
Kotake, Y. et al. Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. Oncogene 30, 1956–1962 (2011).
Cunnington, M.S., Santibanez Koref, M., Mayosi, B.M., Burn, J. & Keavney, B. Chromosome 9p21 SNPs associated with multiple disease phenotypes correlate with ANRIL expression. PLoS Genet. 6, e1000899 (2010).
Zhang, E.B. et al. Long noncoding RNA ANRIL indicates a poor prognosis of gastric cancer and promotes tumor growth by epigenetically silencing of miR-99a/miR-449a. Oncotarget 5, 2276–2292 (2014).
Esteller, M. Non-coding RNAs in human disease. Nat. Rev. Genet. 12, 861–874 (2011).
Sanchez, Y. & Huarte, M. Long non-coding RNAs: challenges for diagnosis and therapies. Nucleic Acid Ther. 23, 15–20 (2013).
Ling, H. et al. Junk DNA and the long non-coding RNA twist in cancer genetics. Oncogene 34, 5003–5011 (2015).
Bussemakers, M.J. et al. DD3: a new prostate-specific gene, highly overexpressed in prostate cancer. Cancer Res. 59, 5975–5979 (1999).
Srikantan, V. et al. PCGEM1, a prostate-specific gene, is overexpressed in prostate cancer. Proc. Natl. Acad. Sci. USA 97, 12216–12221 (2000).
Hessels, D. et al. DD3(PCA3)-based molecular urine analysis for the diagnosis of prostate cancer. Eur. Urol. 44, 8–15 discussion, 15–16 (2003).
Yang, L. et al. lncRNA-dependent mechanisms of androgen-receptor–regulated gene activation programs. Nature 500, 598–602 (2013).
Hung, C.L. et al. A long noncoding RNA connects c-Myc to tumor metabolism. Proc. Natl. Acad. Sci. USA 111, 18697–18702 (2014).
Ji, P. et al. MALAT-1, a novel noncoding RNA, and thymosin β-4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 22, 8031–8041 (2003).
Gutschner, T., Hammerle, M. & Diederichs, S. MALAT1–a paradigm for long noncoding RNA function in cancer. J. Mol. Med. 91, 791–801 (2013).
Tripathi, V. et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell 39, 925–938 (2010).
Yang, L. et al. ncRNA- and Pc2 methylation–dependent gene relocation between nuclear structures mediates gene activation programs. Cell 147, 773–788 (2011).
West, J.A. et al. The long noncoding RNAs NEAT1 and MALAT1 bind active chromatin sites. Mol. Cell 55, 791–802 (2014).
Engreitz, J.M. et al. RNA-RNA interactions enable specific targeting of noncoding RNAs to nascent pre-mRNAs and chromatin sites. Cell 159, 188–199 (2014).
Iyer, M.K. et al. The landscape of long noncoding RNAs in the human transcriptome. Nat. Genet. 47, 199–208 (2015).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Hu, X. et al. A functional genomic approach identifies FAL1 as an oncogenic long noncoding RNA that associates with BMI1 and represses p21 expression in cancer. Cancer Cell 26, 344–357 (2014).
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Puente, X.S. et al. Non-coding recurrent mutations in chronic lymphocytic leukaemia. Nature doi:10.1038/nature14666 (22 July 2015).
Freedman, M.L. et al. Principles for the post-GWAS functional characterization of cancer risk loci. Nat. Genet. 43, 513–518 (2011).
Pandey, G.K. et al. The risk-associated long noncoding RNA NBAT-1 controls neuroblastoma progression by regulating cell proliferation and neuronal differentiation. Cancer Cell 26, 722–737 (2014).
Verhaegh, G.W. et al. Polymorphisms in the H19 gene and the risk of bladder cancer. Eur. Urol. 54, 1118–1126 (2008).
Matouk, I.J. et al. The H19 non-coding RNA is essential for human tumor growth. PLoS ONE 2, e845 (2007).
Angrand, P.O., Vennin, C., Le Bourhis, X. & Adriaenssens, E. The role of long non-coding RNAs in genome formatting and expression. Front. Genet. 6, 165 (2015).
Yang, C. et al. Tag SNPs in long non-coding RNA H19 contribute to susceptibility to gastric cancer in the Chinese Han population. Oncotarget 6, 15311–15320 (2015).
Perez-Llamas, C. & Lopez-Bigas, N. Gitools: analysis and visualisation of genomic data using interactive heat-maps. PLoS ONE 6, e19541 (2011).
Marín-Béjar, O. et al. Pint lincRNA connects the p53 pathway with epigenetic silencing by the Polycomb repressive complex 2. Genome Biol. 14, R104 (2013).
Huarte, M. & Rinn, J.L. Large non-coding RNAs: missing links in cancer? Hum. Mol. Genet. 19, R152–R161 (2010).
Sánchez, Y. et al. Genome-wide analysis of the human p53 transcriptional network unveils a lncRNA tumour suppressor signature. Nat. Commun. 5, 5812 (2014).
Hung, T. et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat. Genet. 43, 621–629 (2011).
Feldstein, O. et al. The long non-coding RNA ERIC is regulated by E2F and modulates the cellular response to DNA damage. Mol. Cancer 12, 131 (2013).
Hart, J.R., Roberts, T.C., Weinberg, M.S., Morris, K.V. & Vogt, P.K. MYC regulates the non-coding transcriptome. Oncotarget 5, 12543–12554 (2014).
Kim, T. et al. Role of MYC-regulated long noncoding RNAs in cell cycle regulation and tumorigenesis. J. Natl. Cancer Inst. 107, dju505 (2015).
Louro, R. et al. Androgen responsive intronic non-coding RNAs. BMC Biol. 5, 4 (2007).
Wang, L.G. et al. Androgen receptor overexpression in prostate cancer linked to Pur alpha loss from a novel repressor complex. Cancer Res. 68, 2678–2688 (2008).
Takayama, K. et al. Androgen-responsive long noncoding RNA CTBP1-AS promotes prostate cancer. EMBO J. 32, 1665–1680 (2013).
Chakravarty, D. et al. The oestrogen receptor α-regulated lncRNA NEAT1 is a critical modulator of prostate cancer. Nat. Commun. 5, 5383 (2014).
Guttman, M. et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 477, 295–300 (2011).
Bennett, C.F. & Swayze, E.E. RNA targeting therapeutics: molecular mechanisms of antisense oligonucleotides as a therapeutic platform. Annu. Rev. Pharmacol. Toxicol. 50, 259–293 (2010).
Kole, R., Krainer, A.R. & Altman, S. RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nat. Rev. Drug Discov. 11, 125–140 (2012).
Miller, J.C. et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nat. Biotechnol. 25, 778–785 (2007).
Zhang, F. et al. Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nat. Biotechnol. 29, 149–153 (2011).
Wiedenheft, B., Sternberg, S.H. & Doudna, J.A. RNA-guided genetic silencing systems in bacteria and archaea. Nature 482, 331–338 (2012).
Ho, T.T. et al. Targeting non-coding RNAs with the CRISPR/Cas9 system in human cell lines. Nucleic Acids Res. 43, e17 (2015).
Han, J. et al. Efficient in vivo deletion of a large imprinted lncRNA by CRISPR/Cas9. RNA Biol. 11, 829–835 (2014).
Eißmann, M. et al. Loss of the abundant nuclear non-coding RNA MALAT1 is compatible with life and development. RNA Biol. 9, 1076–1087 (2012).
Xiang, J.F. et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 24, 513–531 (2014).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517, 583–588 (2015).
Gutschner, T. & Diederichs, S. The hallmarks of cancer: a long non-coding RNA point of view. RNA Biol. 9, 703–719 (2012).
Yang, G., Lu, X. & Yuan, L. LncRNA: a link between RNA and cancer. Biochim. Biophys. Acta 1839, 1097–1109 (2014).
Keniry, A. et al. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat. Cell Biol. 14, 659–665 (2012).
Yang, F. et al. Repression of the long noncoding RNA-LET by histone deacetylase 3 contributes to hypoxia-mediated metastasis. Mol. Cell 49, 1083–1096 (2013).
Li, L. & Chang, H.Y. Physiological roles of long noncoding RNAs: insight from knockout mice. Trends Cell Biol. 24, 594–602 (2014).
Sauvageau, M. et al. Multiple knockout mouse models reveal lincRNAs are required for life and brain development. Elife 2, e01749 (2013).
Grote, P. et al. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev. Cell 24, 206–214 (2013).
Yildirim, E. et al. Xist RNA is a potent suppressor of hematologic cancer in mice. Cell 152, 727–742 (2013).
Necsulea, A. et al. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature 505, 635–640 (2014).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012).
Ulitsky, I., Shkumatava, A., Jan, C.H., Sive, H. & Bartel, D.P. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell 147, 1537–1550 (2011).
Ulitsky, I. & Bartel, D.P. lincRNAs: genomics, evolution, and mechanisms. Cell 154, 26–46 (2013).
Bejerano, G. et al. Ultraconserved elements in the human genome. Science 304, 1321–1325 (2004).
Calin, G.A. et al. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 12, 215–229 (2007).
Schorderet, P. & Duboule, D. Structural and functional differences in the long non-coding RNA hotair in mouse and human. PLoS Genet. 7, e1002071 (2011).
Rinn, J.L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Li, L. et al. Targeted disruption of Hotair leads to homeotic transformation and gene derepression. Cell Rep. 5, 3–12 (2013).
Zhang, B. et al. The lncRNA Malat1 is dispensable for mouse development but its transcription plays a cis-regulatory role in the adult. Cell Rep. 2, 111–123 (2012).
Nakagawa, S., Naganuma, T., Shioi, G. & Hirose, T. Paraspeckles are subpopulation-specific nuclear bodies that are not essential in mice. J. Cell Biol. 193, 31–39 (2011).
Nakagawa, S. et al. The lncRNA Neat1 is required for corpus luteum formation and the establishment of pregnancy in a subpopulation of mice. Development 141, 4618–4627 (2014).
Trimarchi, T. et al. Genome-wide mapping and characterization of Notch-regulated long noncoding RNAs in acute leukemia. Cell 158, 593–606 (2014).
Léveillé, N. et al. Genome-wide profiling of p53-regulated enhancer RNAs uncovers a subset of enhancers controlled by a lncRNA. Nat. Commun. 6, 6520 (2015).
Zhou, Y. et al. Activation of p53 by MEG3 non-coding RNA. J. Biol. Chem. 282, 24731–24742 (2007).
Winkle, M. et al. Long noncoding RNAs as a novel component of the Myc transcriptional network. FASEB J. 29, 2338–2346 (2015).
Jia, L. et al. Functional enhancers at the gene-poor 8q24 cancer-linked locus. PLoS Genet. 5, e1000597 (2009).
Sur, I., Tuupanen, S., Whitington, T., Aaltonen, L.A. & Taipale, J. Lessons from functional analysis of genome-wide association studies. Cancer Res. 73, 4180–4184 (2013).
Ling, H. et al. CCAT2, a novel noncoding RNA mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res. 23, 1446–1461 (2013).
Kim, T. et al. Long-range interaction and correlation between MYC enhancer and oncogenic long noncoding RNA CARLo-5. Proc. Natl. Acad. Sci. USA 111, 4173–4178 (2014).
Tseng, Y.Y. et al. PVT1 dependence in cancer with MYC copy-number increase. Nature 512, 82–86 (2014).
Prensner, J.R. et al. Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat. Biotechnol. 29, 742–749 (2011).
Prensner, J.R. et al. PCAT-1, a long noncoding RNA, regulates BRCA2 and controls homologous recombination in cancer. Cancer Res. 74, 1651–1660 (2014).
Prensner, J.R. et al. The long non-coding RNA PCAT-1 promotes prostate cancer cell proliferation through cMyc. Neoplasia 16, 900–908 (2014).
Prensner, J.R. et al. The IncRNAs PCGEM1 and PRNCR1 are not implicated in castration resistant prostate cancer. Oncotarget 5, 1434–1438 (2014).
Dawson, M.A. & Kouzarides, T. Cancer epigenetics: from mechanism to therapy. Cell 150, 12–27 (2012).
Marchese, F.P. & Huarte, M. Long non-coding RNAs and chromatin modifiers: Their place in the epigenetic code. Epigenetics 9, 21–26 (2014).
Guttman, M. & Rinn, J.L. Modular regulatory principles of large non-coding RNAs. Nature 482, 339–346 (2012).
Khalil, A.M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl. Acad. Sci. USA 106, 11667–11672 (2009).
Zhang, J., Zhang, P., Wang, L., Piao, H.L. & Ma, L. Long non-coding RNA HOTAIR in carcinogenesis and metastasis. Acta Biochim. Biophys. Sin. (Shanghai) 46, 1–5 (2014).
Tsai, M.C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010).
Zhang, E.B. et al. P53-regulated long non-coding RNA TUG1 affects cell proliferation in human non-small cell lung cancer, partly through epigenetically regulating HOXB7 expression. Cell Death Dis. 5, e1243 (2014).
Davidovich, C., Zheng, L., Goodrich, K.J. & Cech, T.R. Promiscuous RNA binding by Polycomb repressive complex 2. Nat. Struct. Mol. Biol. 20, 1250–1257 (2013).
Prensner, J.R. et al. The long noncoding RNA SCHLAP1 promotes aggressive prostate cancer and antagonizes the SWI/SNF complex. Nat. Genet. 45, 1392–1398 (2013).
Mourtada-Maarabouni, M., Pickard, M.R., Hedge, V.L., Farzaneh, F. & Williams, G.T. GAS5, a non-protein-coding RNA, controls apoptosis and is downregulated in breast cancer. Oncogene 28, 195–208 (2009).
Kino, T., Hurt, D.E., Ichijo, T., Nader, N. & Chrousos, G.P. Noncoding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci. Signal. 3, ra8 (2010).
Lee, J.T. & Bartolomei, M.S. X-inactivation, imprinting, and long noncoding RNAs in health and disease. Cell 152, 1308–1323 (2013).
Yoon, J.H. et al. LincRNA-p21 suppresses target mRNA translation. Mol. Cell 47, 648–655 (2012).
Godinho, M., Meijer, D., Setyono-Han, B., Dorssers, L.C. & van Agthoven, T. Characterization of BCAR4, a novel oncogene causing endocrine resistance in human breast cancer cells. J. Cell. Physiol. 226, 1741–1749 (2011).
Xing, Z. et al. lncRNA directs cooperative epigenetic regulation downstream of chemokine signals. Cell 159, 1110–1125 (2014).
Beltran, M. et al. A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Genes Dev. 22, 756–769 (2008).
Salmena, L., Poliseno, L., Tay, Y., Kats, L. & Pandolfi, P.P. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146, 353–358 (2011).
Cesana, M. et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 147, 358–369 (2011).
Poliseno, L. et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 465, 1033–1038 (2010).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013).
Hansen, T.B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388 (2013).
Poliseno, L. et al. Deletion of PTENP1 pseudogene in human melanoma. J. Invest. Dermatol. 131, 2497–2500 (2011).
Cabili, M.N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 25, 1915–1927 (2011).
Panzitt, K. et al. Characterization of HULC, a novel gene with striking up-regulation in hepatocellular carcinoma, as noncoding RNA. Gastroenterology 132, 330–342 (2007).
Record, M., Subra, C., Silvente-Poirot, S. & Poirot, M. Exosomes as intercellular signalosomes and pharmacological effectors. Biochem. Pharmacol. 81, 1171–1182 (2011).
Donner, A. A platform for RNA. SciBX 6 doi:10.1038/scibx.2013.1151 (24 October 2013).
Meng, L. et al. Towards a therapy for Angelman syndrome by targeting a long non-coding RNA. Nature 518, 409–412 (2015).
Ling, H., Fabbri, M. & Calin, G.A. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat. Rev. Drug Discov. 12, 847–865 (2013).
Mizrahi, A. et al. Development of targeted therapy for ovarian cancer mediated by a plasmid expressing diphtheria toxin under the control of H19 regulatory sequences. J. Transl. Med. 7, 69 (2009).
Jiang, J. et al. Translating dosage compensation to trisomy 21. Nature 500, 296–300 (2013).
Chu, C., Qu, K., Zhong, F.L., Artandi, S.E. & Chang, H.Y. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol. Cell 44, 667–678 (2011).
Simon, M.D. et al. The genomic binding sites of a noncoding RNA. Proc. Natl. Acad. Sci. USA 108, 20497–20502 (2011).
Engreitz, J.M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013).
McHugh, C.A., Russell, P. & Guttman, M. Methods for comprehensive experimental identification of RNA-protein interactions. Genome Biol. 15, 203 (2014).
Hezroni, H. et al. Principles of long noncoding RNA evolution derived from direct comparison of transcriptomes in 17 species. Cell Rep. 11, 1110–1122 (2015).
Zhou, Y., Zhang, X. & Klibanski, A. MEG3 noncoding RNA: a tumor suppressor. J. Mol. Endocrinol. 48, R45–R53 (2012).
Wilusz, J.E., Freier, S.M. & Spector, D.L. 3′ end processing of a long nuclear-retained noncoding RNA yields a tRNA-like cytoplasmic RNA. Cell 135, 919–932 (2008).
Acknowledgements
I thank the members of my laboratory and D. O'Brien for critical reading of the manuscript, and I apologize to colleagues whose work was not discussed or cited owing to space constraints. I am supported by European Research Council grant ERC-2011-StG 281877, and by Spanish Ministry of Science Grant RYC-2011-08347.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1 (PDF 602 kb)
Rights and permissions
About this article
Cite this article
Huarte, M. The emerging role of lncRNAs in cancer. Nat Med 21, 1253–1261 (2015). https://doi.org/10.1038/nm.3981
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3981
This article is cited by
-
METTL3-mediated m6A modification of lncRNA TSPAN12 promotes metastasis of hepatocellular carcinoma through SENP1-depentent deSUMOylation of EIF3I
Oncogene (2024)
-
LncRNA TDRKH-AS1 promotes breast cancer progression via the miR-134-5p/CREB1 axis
Journal of Translational Medicine (2023)
-
LINC02159 promotes non-small cell lung cancer progression via ALYREF/YAP1 signaling
Molecular Cancer (2023)
-
The role of LncRNAs in tumor immunotherapy
Cancer Cell International (2023)
-
Identification of cuproptosis-related lncRNAs to predict prognosis and immune infiltration characteristics in alimentary tract malignancies
BMC Bioinformatics (2023)