Abstract
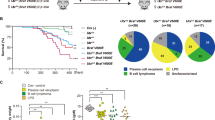
Mutations in the gene encoding the KMT2D (or MLL2) methyltransferase are highly recurrent and occur early during tumorigenesis in diffuse large B cell lymphoma (DLBCL) and follicular lymphoma (FL). However, the functional consequences of these mutations and their role in lymphomagenesis are unknown. Here we show that FL- and DLBCL-associated KMT2D mutations impair KMT2D enzymatic activity, leading to diminished global H3K4 methylation in germinal-center (GC) B cells and DLBCL cells. Conditional deletion of Kmt2d early during B cell development, but not after initiation of the GC reaction, results in an increase in GC B cells and enhances B cell proliferation in mice. Moreover, genetic ablation of Kmt2d in mice overexpressing Bcl2 increases the incidence of GC-derived lymphomas resembling human tumors. These findings suggest that KMT2D acts as a tumor suppressor gene whose early loss facilitates lymphomagenesis by remodeling the epigenetic landscape of the cancer precursor cells. Eradication of KMT2D-deficient cells may thus represent a rational therapeutic approach for targeting early tumorigenic events.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Referenced accessions
NCBI Reference Sequence
References
Shaffer III, A.L., Young, R.M. & Staudt, L.M. Pathogenesis of human B cell lymphomas. Annu. Rev. Immunol. 30, 565–610 (2012).
Basso, K. & Dalla-Favera, R. Germinal centers and B cell lymphomagenesis. Nat. Rev. Immunol. 15, 172–184 (2015).
Morin, R.D. et al. Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature 476, 298–303 (2011).
Pasqualucci, L. et al. Inactivating mutations of acetyltransferase genes in B cell lymphoma. Nature 471, 189–195 (2011).
Pasqualucci, L. et al. Analysis of the coding genome of diffuse large B cell lymphoma. Nat. Genet. 43, 830–837 (2011).
Lohr, J.G. et al. Discovery and prioritization of somatic mutations in diffuse large B cell lymphoma (DLBCL) by whole-exome sequencing. Proc. Natl. Acad. Sci. USA 109, 3879–3884 (2012).
Okosun, J. et al. Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat. Genet. 46, 176–181 (2014).
Pasqualucci, L. et al. Genetics of follicular lymphoma transformation. Cell Rep. 6, 130–140 (2014).
Alizadeh, A.A. et al. Distinct types of diffuse large B cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
Green, M.R. et al. Hierarchy in somatic mutations arising during genomic evolution and progression of follicular lymphoma. Blood 121, 1604–1611 (2013).
Green, M.R. et al. Mutations in early follicular lymphoma progenitors are associated with suppressed antigen presentation. Proc. Natl. Acad. Sci. USA 112, E1116–E1125 (2015).
Shilatifard, A. The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu. Rev. Biochem. 81, 65–95 (2012).
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007).
Li, B., Carey, M. & Workman, J.L. The role of chromatin during transcription. Cell 128, 707–719 (2007).
Briggs, S.D. et al. Histone H3 lysine 4 methylation is mediated by Set1 and required for cell growth and rDNA silencing in Saccharomyces cerevisiae. Genes Dev. 15, 3286–3295 (2001).
Roguev, A. et al. The Saccharomyces cerevisiae Set1 complex includes an Ash2 homologue and methylates histone 3 lysine 4. EMBO J. 20, 7137–7148 (2001).
Miller, T. et al. COMPASS: a complex of proteins associated with a trithorax-related SET domain protein. Proc. Natl. Acad. Sci. USA 98, 12902–12907 (2001).
Krogan, N.J. et al. COMPASS, a histone H3 (lysine 4) methyltransferase required for telomeric silencing of gene expression. J. Biol. Chem. 277, 10753–10755 (2002).
Herz, H.M. et al. Enhancer-associated H3K4 monomethylation by Trithorax-related, the Drosophila homolog of mammalian Mll3/Mll4. Genes Dev. 26, 2604–2620 (2012).
Hu, D. et al. The MLL3/MLL4 branches of the COMPASS family function as major histone H3K4 monomethylases at enhancers. Mol. Cell. Biol. 33, 4745–4754 (2013).
Lee, J.E. et al. H3K4 mono- and di-methyltransferase MLL4 is required for enhancer activation during cell differentiation. eLife 2, e01503 (2013).
Guo, C. et al. KMT2D maintains neoplastic cell proliferation and global histone H3 lysine 4 monomethylation. Oncotarget 4, 2144–2153 (2013).
Kaikkonen, M.U. et al. Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol. Cell 51, 310–325 (2013).
Ng, S.B. et al. Exome sequencing identifies MLL2 mutations as a cause of Kabuki syndrome. Nat. Genet. 42, 790–793 (2010).
Guo, C. et al. Global identification of MLL2-targeted loci reveals MLL2's role in diverse signaling pathways. Proc. Natl. Acad. Sci. USA 109, 17603–17608 (2012).
Santos, M.A. et al. DNA damage–induced differentiation of leukemic cells as an anti-cancer barrier. Nature 514, 107–111 (2014).
Dhar, S.S. et al. Trans-tail regulation of MLL4-catalyzed H3K4 methylation by H4R3 symmetric dimethylation is mediated by a tandem PHD of MLL4. Genes Dev. 26, 2749–2762 (2012).
Casola, S. et al. Tracking germinal center B cells expressing germline immunoglobulin γ1 transcripts by conditional gene targeting. Proc. Natl. Acad. Sci. USA 103, 7396–7401 (2006).
Rickert, R.C., Roes, J. & Rajewsky, K. B lymphocyte–specific, Cre-mediated mutagenesis in mice. Nucleic Acids Res. 25, 1317–1318 (1997).
Carsetti, R., Kohler, G. & Lamers, M.C. Transitional B cells are the target of negative selection in the B cell compartment. J. Exp. Med. 181, 2129–2140 (1995).
Allman, D. et al. Resolution of three nonproliferative immature splenic B cell subsets reveals multiple selection points during peripheral B cell maturation. J. Immunol. 167, 6834–6840 (2001).
Jacob, J. & Kelsoe, G. In situ studies of the primary immune response to (4-hydroxy-3-nitrophenyl)acetyl. II. A common clonal origin for periarteriolar lymphoid sheath–associated foci and germinal centers. J. Exp. Med. 176, 679–687 (1992).
Lim, W.K., Lyashenko, E. & Califano, A. Master regulators used as breast cancer metastasis classifier. Pacific Symposium on Biocomputing 504–515 (2009).
Pao, L.I. et al. B cell–specific deletion of protein-tyrosine phosphatase Shp1 promotes B-1a cell development and causes systemic autoimmunity. Immunity 27, 35–48 (2007).
Yokouchi, M., Suzuki, R., Masuhara, M., Komiya, S., Inoue, A. & Yoshimura, A. Cloning and characterization of APS, an adaptor molecule containing PH and SH2 domains that is tyrosine phosphorylated upon B cell receptor stimulation. Oncogene 15, 7–15 (1997).
Sauer, K. & Cooke, M.P. Regulation of immune cell development through soluble inositol-1,3,4,5-tetrakisphosphate. Nat. Rev. Immunol. 10, 257–271 (2010).
Liu, Y.J. et al. Mechanism of antigen-driven selection in germinal centers. Nature 342, 929–931 (1989).
Egle, A., Harris, A.W., Bath, M.L., O'Reilly, L. & Cory, S. VavP-Bcl2 transgenic mice develop follicular lymphoma preceded by germinal center hyperplasia. Blood 103, 2276–2283 (2004).
Béguelin, W. et al. EZH2 is required for germinal center formation and somatic EZH2 mutations promote lymphoid transformation. Cancer Cell 23, 677–692 (2013).
Zhang, J. et al. Genetic heterogeneity of diffuse large B cell lymphoma. Proc. Natl. Acad. Sci. USA 110, 1398–1403 (2013).
Creyghton, M.P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl. Acad. Sci. USA 107, 21931–21936 (2010).
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011).
Whyte, W.A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Chapuy, B. et al. Discovery and characterization of super-enhancer–associated dependencies in diffuse large B cell lymphoma. Cancer Cell 24, 777–790 (2013).
Ardehali, M.B. et al. Drosophila Set1 is the major histone H3 lysine 4 trimethyltransferase with role in transcription. EMBO J. 30, 2817–2828 (2011).
Calo, E. & Wysocka, J. Modification of enhancer chromatin: what, how and why? Mol. Cell 49, 825–837 (2013).
Petruk, S. et al. TrxG and PcG proteins but not methylated histones remain associated with DNA through replication. Cell 150, 922–933 (2012).
Pasqualucci, L. The genetic basis of diffuse large B cell lymphoma. Curr. Opin. Hematol. 20, 336–344 (2013).
Pasqualucci, L. et al. AID is required for germinal center–derived lymphomagenesis. Nat. Genet. 40, 108–112 (2008).
Pasqualucci, L. et al. Hypermutation of multiple proto-oncogenes in B cell diffuse large-cell lymphomas. Nature 412, 341–346 (2001).
Hatzi, K. & Melnick, A. Breaking bad in the germinal center: how deregulation of BCL6 contributes to lymphomagenesis. Trends Mol. Med. 20, 343–352 (2014).
Cattoretti, G. et al. Deregulated BCL6 expression recapitulates the pathogenesis of human diffuse large B cell lymphomas in mice. Cancer Cell 7, 445–455 (2005).
Mandelbaum, J. et al. BLIMP1 is a tumor suppressor gene frequently disrupted in activated B cell–like diffuse large B cell lymphoma. Cancer Cell 18, 568–579 (2010).
Souers, A.J. et al. ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets. Nat. Med. 19, 202–208 (2013).
Knutson, S.K. et al. A selective inhibitor of EZH2 blocks H3K27 methylation and kills mutant lymphoma cells. Nat. Chem. Biol. 8, 890–896 (2012).
McCabe, M.T. et al. EZH2 inhibition as a therapeutic strategy for lymphoma with EZH2-activating mutations. Nature 492, 108–112 (2012).
Klein, U. et al. Transcriptional analysis of the B cell germinal center reaction. Proc. Natl. Acad. Sci. USA 100, 2639–2644 (2003).
Compagno, M. et al. Mutations of multiple genes cause deregulation of NF-κB in diffuse large B cell lymphoma. Nature 459, 717–721 (2009).
Dominguez-Sola, D. et al. The proto-oncogene is required for selection in the germinal center and cyclic reentry. Nat. Immunol. 13, 1083–1091 (2012).
Thomas, M. et al. Full deacylation of polyethylenimine dramatically boosts its gene delivery efficiency and specificity to mouse lung. Proc. Natl. Acad. Sci. USA 102, 5679–5684 (2005).
Bereshchenko, O.R., Gu, W. & Dalla-Favera, R. Acetylation inactivates the transcriptional repressor BCL6. Nat. Genet. 32, 606–613 (2002).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Green, M.R. & Sambrook, J. Molecular Cloning: A Laboratory Manual 4th edn. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2012).
Jolly, C.J., Klix, N. & Neuberger, M.S. Rapid methods for the analysis of immunoglobulin gene hypermutation: application to transgenic and gene-targeted mice. Nucleic Acids Res. 25, 1913–1919 (1997).
Klein, U. et al. Transcription factor IRF4 controls plasma cell differentiation and class-switch recombination. Nat. Immunol. 7, 773–782 (2006).
Reich, M. et al. GenePattern 2.0. Nat. Genet. 38, 500–501 (2006).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate–a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Subramanian, A., Kuehn, H., Gould, J., Tamayo, P. & Mesirov, J.P. GSEA-P: a desktop application for gene set enrichment analysis. Bioinformatics 23, 3251–3253 (2007).
Huang, D.W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Blecher-Gonen, R. et al. High-throughput chromatin immunoprecipitation for genome-wide mapping of in vivo protein-DNA interactions and epigenomic states. Nat. Protoc. 8, 539–554 (2013).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment–map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Giannopoulou, E.G. & Elemento, O. An integrated ChIP-seq analysis platform with customizable workflows. BMC Bioinformatics 12, 277 (2011).
Lin, C.Y. et al. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 151, 56–67 (2012).
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Acknowledgements
We would like to thank U. Klein and S. Zha for discussions, S. Nataraj, L. Belver, C. Scuoppo and N. De Silva for help and suggestions on various experimental procedures, E. McIntush (Bethyl Laboratories, Inc.) for help in generating several antibodies to KMT2D, the Flow Cytometry Shared Resource of the Herbert Irving Comprehensive Cancer Center at Columbia University for assistance with cell-sorting procedures, the Molecular Pathology lab for the preparation of mouse FFPE tissue samples and the Genomic Technologies Shared Resource for sequencing the ChIP-DNA libraries. We also thank S. Cory (Walter and Eliza Hall Institute of Medical Research, Parkville, Australia) for the VavP-Bcl2 mice and K. Rajewsky (Max Delbrück Center for Molecular Medicine, Berlin, Germany) for the CD19-Cre and Cγ1-Cre mice. The indicated DLBCL cell lines in Supplementary Table 5 were a kind gift of L. Staudt, M.A. Shipp and T. Gilmore. This work was supported by US National Institutes of Health grants RO1-CA172492 (L.P.) and RO1-CA37295 (R.D.-F.), and J.Z. is supported by a Lymphoma Research Foundation postdoctoral fellowship. L.P. is on leave from the University of Perugia.
Author information
Authors and Affiliations
Contributions
L.P. designed and directed the study, analyzed data and wrote the manuscript, with contributions from R.D.-F. and J.Z. J.Z. performed experiments, analyzed data and prepared figures. D.D.-S. and S.H. were responsible for histopathological analysis. J.-E.L. and K.G. generated the Kmt2d conditional knockout mouse model. K.B. performed the ChIP-Seq experiment and, together with A.B.H, contributed to the implementation of tools for the analysis of ChIP-Seq data. S.V. performed mutation analysis of the rearranged immunoglobulin genes from mouse tumors and provided technical support in various experiments. M.B. conducted the analysis of gene expression data. T.M. was responsible for animal husbandry. H.T. performed mouse autopsies and processed tissues for FACS and histologic analysis. R.D.-F. contributed to the study design and data analysis. All authors reviewed the manuscript and provided final approval for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Notes, Supplementary Figures 1–11 & Supplementary Tables 1–7 (PDF 5973 kb)
Rights and permissions
About this article
Cite this article
Zhang, J., Dominguez-Sola, D., Hussein, S. et al. Disruption of KMT2D perturbs germinal center B cell development and promotes lymphomagenesis. Nat Med 21, 1190–1198 (2015). https://doi.org/10.1038/nm.3940
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3940
This article is cited by
-
Histone methyltransferase KMT2D inhibits ENKTL carcinogenesis by epigenetically activating SGK1 and SOCS1
Genes & Genomics (2024)
-
Altered pathways and targeted therapy in double hit lymphoma
Journal of Hematology & Oncology (2022)
-
Identification of clinical implications and potential prognostic models of chromatin regulator mutations in multiple myeloma
Clinical Epigenetics (2022)
-
The follicular lymphoma epigenome regulates its microenvironment
Journal of Experimental & Clinical Cancer Research (2022)
-
Aberrant SPOP-CHAF1A ubiquitination axis triggers tumor autophagy that endows a therapeutical vulnerability in diffuse large B cell lymphoma
Journal of Translational Medicine (2022)