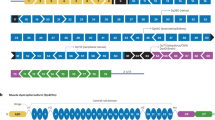
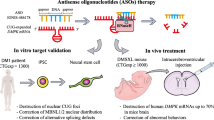
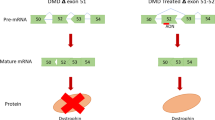
Abstract
Antisense oligonucleotides (AONs) hold promise for therapeutic correction of many genetic diseases via exon skipping, and the first AON-based drugs have entered clinical trials for neuromuscular disorders1,2. However, despite advances in AON chemistry and design, systemic use of AONs is limited because of poor tissue uptake, and recent clinical reports confirm that sufficient therapeutic efficacy has not yet been achieved. Here we present a new class of AONs made of tricyclo-DNA (tcDNA), which displays unique pharmacological properties and unprecedented uptake by many tissues after systemic administration. We demonstrate these properties in two mouse models of Duchenne muscular dystrophy (DMD), a neurogenetic disease typically caused by frame-shifting deletions or nonsense mutations in the gene encoding dystrophin3,4 and characterized by progressive muscle weakness, cardiomyopathy, respiratory failure5 and neurocognitive impairment6. Although current naked AONs do not enter the heart or cross the blood-brain barrier to any substantial extent, we show that systemic delivery of tcDNA-AONs promotes a high degree of rescue of dystrophin expression in skeletal muscles, the heart and, to a lesser extent, the brain. Our results demonstrate for the first time a physiological improvement of cardio-respiratory functions and a correction of behavioral features in DMD model mice. This makes tcDNA-AON chemistry particularly attractive as a potential future therapy for patients with DMD and other neuromuscular disorders or with other diseases that are eligible for exon-skipping approaches requiring whole-body treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Goemans, N.M. et al. Systemic administration of PRO051 in Duchenne's muscular dystrophy. N. Engl. J. Med. 364, 1513–1522 (2011).
Cirak, S. et al. Exon skipping and dystrophin restoration in patients with Duchenne muscular dystrophy after systemic phosphorodiamidate morpholino oligomer treatment: an open-label, phase 2, dose-escalation study. Lancet 378, 595–605 (2011).
Koenig, M. et al. Complete cloning of the Duchenne muscular dystrophy (DMD) cDNA and preliminary genomic organization of the DMD gene in normal and affected individuals. Cell 50, 509–517 (1987).
Muntoni, F., Torelli, S. & Ferlini, A. Dystrophin and mutations: one gene, several proteins, multiple phenotypes. Lancet Neurol. 2, 731–740 (2003).
Emery, A.E. The muscular dystrophies. Lancet 359, 687–695 (2002).
Bresolin, N. et al. Cognitive impairment in Duchenne muscular dystrophy. Neuromuscul. Disord. 4, 359–369 (1994).
Mendell, J.R. et al. Eteplirsen for the treatment of Duchenne muscular dystrophy. Ann. Neurol. 74, 637–647 (2013).
Lu, Q.L., Cirak, S. & Partridge, T. What can we learn from clinical trials of exon skipping for DMD? Mol. Ther. Nucleic Acids 3, e152 (2014).
Townsend, D., Yasuda, S.L., Chamberlain, J.S. & Metzger, J.M. Emergent dilated cardiomyopathy caused by targeted repair of dystrophic skeletal muscle. Mol. Ther. 16, 832–835 (2008).
Renneberg, D. & Leumann, C.J. Watson-Crick base-pairing properties of tricyclo-DNA. J. Am. Chem. Soc. 124, 5993–6002 (2002).
Ittig, D., Gerber, A.B. & Leumann, C.J. Position-dependent effects on stability in tricyclo-DNA modified oligonucleotide duplexes. Nucleic Acids Res. 39, 373–380 (2011).
Renneberg, D., Bouliong, E., Reber, U., Schumperli, D. & Leumann, C.J. Antisense properties of tricyclo-DNA. Nucleic Acids Res. 30, 2751–2757 (2002).
Ivanova, G. et al. Tricyclo-DNA containing oligonucleotides as steric block inhibitors of human immunodeficiency virus type 1 tat-dependent trans-activation and HIV-1 infectivity. Oligonucleotides 17, 54–65 (2007).
Murray, S. et al. TricycloDNA-modified oligo-2′-deoxyribonucleotides reduce scavenger receptor B1 mRNA in hepatic and extra-hepatic tissues—a comparative study of oligonucleotide length, design and chemistry. Nucleic Acids Res. 40, 6135–6143 (2012).
Nel, A.E. et al. Understanding biophysicochemical interactions at the nano-bio interface. Nat. Mater. 8, 543–557 (2009).
Sicinski, P. et al. The molecular basis of muscular dystrophy in the mdx mouse: a point mutation. Science 244, 1578–1580 (1989).
Mann, C.J., Honeyman, K., McClorey, G., Fletcher, S. & Wilton, S.D. Improved antisense oligonucleotide induced exon skipping in the mdx mouse model of muscular dystrophy. J. Gene Med. 4, 644–654 (2002).
Alter, J. et al. Systemic delivery of morpholino oligonucleotide restores dystrophin expression bodywide and improves dystrophic pathology. Nat. Med. 12, 175–177 (2006).
Heemskerk, H.A. et al. In vivo comparison of 2′-O-methyl phosphorothioate and morpholino antisense oligonucleotides for Duchenne muscular dystrophy exon skipping. J. Gene Med. 11, 257–266 (2009).
Kohler, M. et al. Quality of life, physical disability, and respiratory impairment in Duchenne muscular dystrophy. Am. J. Respir. Crit. Care Med. 172, 1032–1036 (2005).
Quinlan, J.G. et al. Evolution of the mdx mouse cardiomyopathy: physiological and morphological findings. Neuromuscul. Disord. 14, 491–496 (2004).
Spurney, C.F. et al. Dystrophin-deficient cardiomyopathy in mouse: expression of Nox4 and Lox are associated with fibrosis and altered functional parameters in the heart. Neuromuscul. Disord. 18, 371–381 (2008).
Perronnet, C. & Vaillend, C. Dystrophins, utrophins, and associated scaffolding complexes: role in mammalian brain and implications for therapeutic strategies. J. Biomed. Biotechnol. 2010, 849426 (2010).
Sekiguchi, M. et al. A deficit of brain dystrophin impairs specific amygdala GABAergic transmission and enhances defensive behaviour in mice. Brain 132, 124–135 (2009).
Yamamoto, K. et al. Reduction of abnormal behavioral response to brief restraint by information from other mice in dystrophin-deficient mdx mice. Neuromuscul. Disord. 20, 505–511 (2010).
Deconinck, A.E. et al. Utrophin-dystrophin-deficient mice as a model for Duchenne muscular dystrophy. Cell 90, 717–727 (1997).
Grady, R.M. et al. Skeletal and cardiac myopathies in mice lacking utrophin and dystrophin: a model for Duchenne muscular dystrophy. Cell 90, 729–738 (1997).
Wu, C.C. et al. Selection of oligonucleotide aptamers with enhanced uptake and activation of human leukemia B cells. Hum. Gene Ther. 14, 849–860 (2003).
Malerba, A., Thorogood, F.C., Dickson, G. & Graham, I.R. Dosing regimen has a significant impact on the efficiency of morpholino oligomer-induced exon skipping in mdx mice. Hum. Gene Ther. 955–965 (2009).
Malerba, A. et al. Chronic systemic therapy with low-dose morpholino oligomers ameliorates the pathology and normalizes locomotor behavior in mdx mice. Mol. Ther. 19, 345–354 (2011).
Verhaart, I.E. et al. Dose-dependent pharmacokinetic profiles of 2′-O-methyl phosphorothioate antisense oligonucleotides in mdx mice. Nucleic Acid Ther 23, 228–237 (2013).
Goyenvalle, A. et al. Rescue of severely affected dystrophin/utrophin-deficient mice through scAAV-U7snRNA-mediated exon skipping. Hum. Mol. Genet. 2559–2571 (2012).
Goyenvalle, A. et al. Prevention of dystrophic pathology in severely affected dystrophin/utrophin-deficient mice by morpholino-oligomer-mediated exon-skipping. Mol. Ther. 18, 198–205 (2010).
Acknowledgements
We thank P.O. Buclez for technical assistance as well as D. Staunton and the Biophysical Instrumentation Facility at the University of Oxford for help with the SEC-MALS experiment. We also thank S. Vinit, A. Jacquet and N. Mougenot for assistance with the plethysmography and echocardiography experiments, and D. Mornet (INSERM, Montpellier) for the dystrophin-specific primary antibody 5G5. This work was supported by the Agence Nationale de la Recherche (Chair of Excellence HandiMedEx), the Association Monegasque Contre les Myopathies, the Duchenne Parent Project France, the aktion Benni & Co. and the UK Medical Research Council.
Author information
Authors and Affiliations
Contributions
A.G., S.E.A., K.E., T.V., S.S., M.J.A.W., K.E.D., C.V., C.L and L.G. participated in the planning, design and interpretation of experiments. A.G., G.G., A.B., S.E.A., K.E., A.A., R.C., B.D., A.F., C.B., H.A. and C.V. carried out experiments. A.G. and L.G. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Branislav Dugovic is employed by Synthena, which produces tricyclo-DNA oligomers. Christian Leumann and Luis Garcia are co-funders of Synthena.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1 and Supplementary Figures 1–8 (PDF 673 kb)
Control dKO mouse
Representative decreased ambulation of a 8-week old control dKO mouse. The reduced musculature considerably affects the mobility of the mice, which display a very severe dystrophic phenotype, including contracted and stiff limbs, and very pronounced kyphosis as a result of the degenerative process. (AVI 4243 kb)
TcDNA treated dKO mouse.
The dystrophic pathology of the treated mouse appears greatly improved as the animal demonstrates only a minimal kyphosis and is very mobile compared to the untreated control. (AVI 5886 kb)
Source data
Rights and permissions
About this article
Cite this article
Goyenvalle, A., Griffith, G., Babbs, A. et al. Functional correction in mouse models of muscular dystrophy using exon-skipping tricyclo-DNA oligomers. Nat Med 21, 270–275 (2015). https://doi.org/10.1038/nm.3765
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3765
This article is cited by
-
Splice-Modulating Antisense Oligonucleotides as Therapeutics for Inherited Metabolic Diseases
BioDrugs (2024)
-
The unconditioned fear response in dystrophin-deficient mice is associated with adrenal and vascular function
Scientific Reports (2023)
-
In vivo restoration of dystrophin expression in mdx mice using intra-muscular and intra-arterial injections of hydrogel microsphere carriers of exon skipping antisense oligonucleotides
Cell Death & Disease (2022)
-
Epileptic disorders in Becker and Duchenne muscular dystrophies: a systematic review and meta-analysis
Journal of Neurology (2022)
-
Advances in oligonucleotide drug delivery
Nature Reviews Drug Discovery (2020)