Abstract
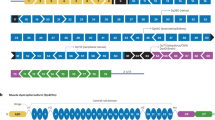
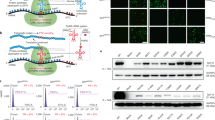
Most mutations that truncate the reading frame of the DMD gene cause loss of dystrophin expression and lead to Duchenne muscular dystrophy. However, amelioration of disease severity has been shown to result from alternative translation initiation beginning in DMD exon 6 that leads to expression of a highly functional N-truncated dystrophin. Here we demonstrate that this isoform results from usage of an internal ribosome entry site (IRES) within exon 5 that is glucocorticoid inducible. We confirmed IRES activity by both peptide sequencing and ribosome profiling in muscle from individuals with minimal symptoms despite the presence of truncating mutations. We generated a truncated reading frame upstream of the IRES by exon skipping, which led to synthesis of a functional N-truncated isoform in both human subject–derived cell lines and in a new DMD mouse model, where expression of the truncated isoform protected muscle from contraction-induced injury and corrected muscle force to the same level as that observed in control mice. These results support a potential therapeutic approach for patients with mutations within the 5′ exons of DMD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Referenced accessions
NCBI Reference Sequence
Change history
25 August 2014
In the version of this article initially published online, the nucleotide numbering of the AS (–3 to + 16), B (+17 to +40) and C (+49 to –10) targeted sequences in Figure 3b was incorrect, and thus the schematic of the sequences was incorrectly drawn. The correct numbering is as follows: AS, –3 to + 18; B, +17 to +44; and C, +49 to –8. The correction has no impact on the results of the study or its conclusions. The error has been corrected for all versions of this article.
13 March 2015
In the version of this article initially published, three participants of the study were not included as co-authors. Also, one of the individuals mentioned in the Acknowledgments section of the report was incorrectly included and thus has been removed at their request, and the name of another individual mentioned in the Acknowledgments was originally misspelled (“Fabbri” should have been "Fabri"). The errors have been corrected in the HTML and PDF versions of the article.
References
Monaco, A.P. Dystrophin, the protein product of the Duchenne/Becker muscular dystrophy gene. Trends Biochem. Sci. 14, 412–415 (1989).
Flanigan, K.M. et al. DMD Trp3X nonsense mutation associated with a founder effect in North American families with mild Becker muscular dystrophy. Neuromuscul. Disord. 19, 743–748 (2009).
Flanigan, K.M. et al. Mutational spectrum of DMD mutations in dystrophinopathy patients: application of modern diagnostic techniques to a large cohort. Hum. Mutat. 30, 1657–1666 (2009).
Gurvich, O.L. et al. DMD exon 1 truncating point mutations: amelioration of phenotype by alternative translation initiation in exon 6. Hum. Mutat. 30, 633–640 (2009).
Witting, N., Duno, M. & Vissing, J. Becker muscular dystrophy with widespread muscle hypertrophy and a non-sense mutation of exon 2. Neuromuscul. Disord. 23, 25–28 (2013).
Gimona, M., Djinovic-Carugo, K., Kranewitter, W.J. & Winder, S.J. Functional plasticity of CH domains. FEBS Lett. 513, 98–106 (2002).
Hertz, M.I., Landry, D.M., Willis, A.E., Luo, G. & Thompson, S.R. Ribosomal protein S25 dependency reveals a common mechanism for diverse internal ribosome entry sites and ribosome shunting. Mol. Cell. Biol. 33, 1016–1026 (2013).
Spriggs, K.A., Bushell, M. & Willis, A.E. Translational regulation of gene expression during conditions of cell stress. Mol. Cell 40, 228–237 (2010).
Jang, S.K. et al. A segment of the 5′ nontranslated region of encephalomyocarditis virus RNA directs internal entry of ribosomes during in vitro translation. J. Virol. 62, 2636–2643 (1988).
Miura, P., Thompson, J., Chakkalakal, J.V., Holcik, M. & Jasmin, B.J. The utrophin A 5′-untranslated region confers internal ribosome entry site-mediated translational control during regeneration of skeletal muscle fibers. J. Biol. Chem. 280, 32997–33005 (2005).
Miura, P., Andrews, M., Holcik, M. & Jasmin, B.J. IRES-mediated translation of utrophin A is enhanced by glucocorticoid treatment in skeletal muscle cells. PLoS ONE 3, e2309 (2008).
Cornelis, S. et al. Identification and characterization of a novel cell cycle–regulated internal ribosome entry site. Mol. Cell 5, 597–605 (2000).
Komar, A.A. et al. Internal initiation drives the synthesis of Ure2 protein lacking the prion domain and affects [URE3] propagation in yeast cells. EMBO J. 22, 1199–1209 (2003).
Lauring, A.S. & Overbaugh, J. Evidence that an IRES within the Notch2 coding region can direct expression of a nuclear form of the protein. Mol. Cell 6, 939–945 (2000).
Maier, D., Nagel, A.C. & Preiss, A. Two isoforms of the Notch antagonist Hairless are produced by differential translation initiation. Proc. Natl. Acad. Sci. USA 99, 15480–15485 (2002).
Baird, S.D., Turcotte, M., Korneluk, R.G. & Holcik, M. Searching for IRES. RNA 12, 1755–1785 (2006).
Heppner Goss, K., Trzepacz, C., Tuohy, T.M. & Groden, J. Attenuated APC alleles produce functional protein from internal translation initiation. Proc. Natl. Acad. Sci. USA 99, 8161–8166 (2002).
White, S.J. et al. Duplications in the DMD gene. Hum. Mutat. 27, 938–945 (2006).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J.V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Ameur, A. et al. Total RNA sequencing reveals nascent transcription and widespread co-transcriptional splicing in the human brain. Nat. Struct. Mol. Biol. 18, 1435–1440 (2011).
Johannes, G., Carter, M.S., Eisen, M.B., Brown, P.O. & Sarnow, P. Identification of eukaryotic mRNAs that are translated at reduced cap binding complex eIF4F concentrations using a cDNA microarray. Proc. Natl. Acad. Sci. USA 96, 13118–13123 (1999).
Murray, E.L. & Schoenberg, D.R. A+U-rich instability elements differentially activate 5′-3′ and 3′-5′ mRNA decay. Mol. Cell. Biol. 27, 2791–2799 (2007).
Stoneley, M. et al. Analysis of the c-myc IRES; a potential role for cell-type specific trans-acting factors and the nuclear compartment. Nucleic Acids Res. 28, 687–694 (2000).
Thompson, S.R. So you want to know if your message has an IRES? Wiley interdisciplinary reviews. RNA 3, 697–705 (2012).
Kozak, M. A second look at cellular mRNA sequences said to function as internal ribosome entry sites. Nucleic Acids Res. 33, 6593–6602 (2005).
Wood, M.J., Gait, M.J. & Yin, H. RNA-targeted splice-correction therapy for neuromuscular disease. Brain 133, 957–972 (2010).
Goyenvalle, A. et al. Rescue of dystrophic muscle through U7 snRNA–mediated exon skipping. Science 306, 1796–1799 (2004).
Vulin, A. et al. Muscle function recovery in golden retriever muscular dystrophy after AAV1–U7 exon skipping. Mol. Ther. 20, 2120–2133 (2012).
Chaouch, S. et al. Immortalized skin fibroblasts expressing conditional MyoD as a renewable and reliable source of converted human muscle cells to assess therapeutic strategies for muscular dystrophies: validation of an exon-skipping approach to restore dystrophin in Duchenne muscular dystrophy cells. Hum. Gene Ther. 20, 784–790 (2009).
Aartsma-Rus, A. et al. Functional analysis of 114 exon-internal AONs for targeted DMD exon skipping: indication for steric hindrance of SR protein binding sites. Oligonucleotides 15, 284–297 (2005).
Matsumura, K., Ervasti, J.M., Ohlendieck, K., Kahl, S.D. & Campbell, K.P. Association of dystrophin-related protein with dystrophin-associated proteins in mdx mouse muscle. Nature 360, 588–591 (1992).
Kozak, M. & Shatkin, A.J. Migration of 40 S ribosomal subunits on messenger RNA in the presence of edeine. J. Biol. Chem. 253, 6568–6577 (1978).
Calvo, S.E., Pagliarini, D.J. & Mootha, V.K. Upstream open reading frames cause widespread reduction of protein expression and are polymorphic among humans. Proc. Natl. Acad. Sci. USA 106, 7507–7512 (2009).
Kozak, M. Context effects and inefficient initiation at non-AUG codons in eucaryotic cell-free translation systems. Mol. Cell. Biol. 9, 5073–5080 (1989).
Hellen, C.U. & Sarnow, P. Internal ribosome entry sites in eukaryotic mRNA molecules. Genes Dev. 15, 1593–1612 (2001).
López-Lastra, M., Rivas, A. & Barria, M.I. Protein synthesis in eukaryotes: the growing biological relevance of cap-independent translation initiation. Biol. Res. 38, 121–146 (2005).
Yaman, I. et al. The zipper model of translational control: a small upstream ORF is the switch that controls structural remodeling of an mRNA leader. Cell 113, 519–531 (2003).
Cencig, S. et al. Mapping and characterization of the minimal internal ribosome entry segment in the human c-myc mRNA 5′ untranslated region. Oncogene 23, 267–277 (2004).
Gerbasi, V.R. & Link, A.J. The myotonic dystrophy type 2 protein ZNF9 is part of an ITAF complex that promotes cap-independent translation. Mol. Cell. Proteomics 6, 1049–1058 (2007).
Pickering, B.M., Mitchell, S.A., Spriggs, K.A., Stoneley, M. & Willis, A.E. Bag-1 internal ribosome entry segment activity is promoted by structural changes mediated by poly(rC) binding protein 1 and recruitment of polypyrimidine tract binding protein 1. Mol. Cell. Biol. 24, 5595–5605 (2004).
Campbell, K.P. Three muscular dystrophies: loss of cytoskeleton-extracellular matrix linkage. Cell 80, 675–679 (1995).
Bloch, R.J. & Gonzalez-Serratos, H. Lateral force transmission across costameres in skeletal muscle. Exerc. Sport Sci. Rev. 31, 73–78 (2003).
Stradal, T., Kranewitter, W., Winder, S.J. & Gimona, M. CH domains revisited. FEBS Lett. 431, 134–137 (1998).
Gimona, M. & Winder, S.J. Single calponin homology domains are not actin-binding domains. Curr. Biol. 8, R674–R675 (1998).
Norwood, F.L., Sutherland-Smith, A.J., Keep, N.H. & Kendrick-Jones, J. The structure of the N-terminal actin-binding domain of human dystrophin and how mutations in this domain may cause Duchenne or Becker muscular dystrophy. Structure 8, 481–491 (2000).
Sutherland-Smith, A.J. et al. An atomic model for actin binding by the CH domains and spectrin-repeat modules of utrophin and dystrophin. J. Mol. Biol. 329, 15–33 (2003).
Rybakova, I.N., Humston, J.L., Sonnemann, K.J. & Ervasti, J.M. Dystrophin and utrophin bind actin through distinct modes of contact. J. Biol. Chem. 281, 9996–10001 (2006).
Lin, A.Y. et al. Impacts of dystrophin and utrophin domains on actin structural dynamics: implications for therapeutic design. J. Mol. Biol. 420, 87–98 (2012).
Henderson, D.M., Lin, A.Y., Thomas, D.D. & Ervasti, J.M. The carboxy-terminal third of dystrophin enhances actin binding activity. J. Mol. Biol. 416, 414–424 (2012).
Corrado, K. et al. Transgenic mdx mice expressing dystrophin with a deletion in the actin-binding domain display a “mild Becker” phenotype. J. Cell Biol. 134, 873–884 (1996).
Le Rumeur, E., Winder, S.J. & Hubert, J.F. Dystrophin: more than just the sum of its parts. Biochim. Biophys. Acta 1804, 1713–1722 (2010).
Bhosle, R.C., Michele, D.E., Campbell, K.P., Li, Z. & Robson, R.M. Interactions of intermediate filament protein synemin with dystrophin and utrophin. Biochem. Biophys. Res. Commun. 346, 768–777 (2006).
Rezniczek, G.A. et al. Plectin 1f scaffolding at the sarcolemma of dystrophic (mdx) muscle fibers through multiple interactions with β-dystroglycan. J. Cell Biol. 176, 965–977 (2007).
Prins, K.W. et al. Dystrophin is a microtubule-associated protein. J. Cell Biol. 186, 363–369 (2009).
Beggs, A.H. et al. Exploring the molecular basis for variability among patients with Becker muscular dystrophy: dystrophin gene and protein studies. Am. J. Hum. Genet. 49, 54–67 (1991).
Winnard, A.V. et al. Characterization of translational frame exception patients in Duchenne/Becker muscular dystrophy. Hum. Mol. Genet. 2, 737–744 (1993).
Winnard, A.V., Mendell, J.R., Prior, T.W., Florence, J. & Burghes, A.H. Frameshift deletions of exons 3–7 and revertant fibers in Duchenne muscular dystrophy: mechanisms of dystrophin production. Am. J. Hum. Genet. 56, 158–166 (1995).
Heald, A., Anderson, L.V., Bushby, K.M. & Shaw, P.J. Becker muscular dystrophy with onset after 60 years. Neurology 44, 2388–2390 (1994).
Johannes, G., Carter, M.S., Eisen, M.B., Brown, P.O. & Sarnow, P. Identification of eukaryotic mRNAs that are translated at reduced cap binding complex eIF4F concentrations using a cDNA microarray. Proc. Natl. Acad. Sci. USA 96, 13118–13123 (1999).
Goyenvalle, A. et al. Rescue of dystrophic muscle through U7 snRNA-mediated exon skipping. Science 306, 1796–1799 (2004).
Wein, N. et al. Efficient bypass of mutations in dysferlin deficient patient cells by antisense-induced exon skipping. Hum. Mutat. 31, 136–142 (2010).
Clark, K.R., Liu, X., McGrath, J.P. & Johnson, P.R. Highly purified recombinant adeno-associated virus vectors are biologically active and free of detectable helper and wild-type viruses. Hum. Gene Ther. 10, 1031–1039 (1999).
Bovolenta, M. et al. A novel custom high density-comparative genomic hybridization array detects common rearrangements as well as deep intronic mutations in dystrophinopathies. BMC Genomics 9, 572 (2008).
Morris, G.E. et al. An epitope structure for the C-terminal domain of dystrophin and utrophin. Biochemistry 37, 11117–11127 (1998).
Spurney, C.F. et al. Preclinical drug trials in the mdx mouse: assessment of reliable and sensitive outcome measures. Muscle Nerve 39, 591–602 (2009).
Hakim, C.H., Grange, R.W. & Duan, D. The passive mechanical properties of the extensor digitorum longus muscle are compromised in 2- to 20-mo-old mdx mice. J. Appl. Physiol. 110, 1656–1663 (2011).
Julesz, B. Spatial nonlinearities in the instantaneous perception of textures with identical power spectra. Phil. Trans. R. Soc. Lond. B 290, 83–94 (1980).
Porikli, F. Integral histogram: A fast way to extract histograms in cartesian spaces. Proc. IEEE Comput. Soc. Conf. Comput. Vis. Pattern Recognit. 1, 829–836 (2005).
Schmid, C. Constructing models for content-based image retrieval. Proc. IEEE Comput. Soc. Conf. Comput. Vis. Pattern Recognit. 2, II-39–II-45 (2001).
Di Zenzo, S. A note on the gradient of a multi-image. Comput. Vis. Graph. Image Process. 33, 116–125 (1986).
Gevers, T. Adaptive image segmentation by combining photometric invariant region and edge information. IEEE Trans. Pattern Anal. Mach. Intrll. 24, 848–852 (2002).
Sapiro, G. & Ringach, D.L. Anisotropic diffusion of multivalued images with applications to color filtering. IEEE Trans. Image Process. 5, 1582–1586 (1996).
Acknowledgements
This work has been supported by the US National Institutes of Health National Institute of Neurological Disorders and Stroke (R01 NS043264 (K.M.F., M.T.H. and R.B.W.)), the National Institute of General Medical Sciences (R01 GM038277 and R01 GM084177 (D.R.S.)), and CureDuchenne (K.M.F.). N.W. has received fellowship support from the Ohio State University/Nationwide Children's Hospital Muscle Group and the Philippe Foundation. We wish to acknowledge the technical assistance of A. Rutherford, Y. Kaminoh and L. Taylor and are grateful for the reagents provided by P. Sarnow (pRDEF; Stanford University) and G. Morris (hybridoma cell lines for antibodies MANEX1A, MANCHO3 and MANDAG2; MDA Monoclonal Antibody Resource at the Wolfson Centre for Inherited Neuromuscular Disease). The EU BIO-NMD project (no. 241665) provided support to A. Ferlini and C.A.-K.S., and the Duchenne Parent Project Italy (DMD Diagnostics Project) provided support to A. Ferlini. Thanks are also due to C. Rodolico and M. Fabris for their technical support. The Clinical Proteomics Mass Spectrometry core facility at Karolinska University Hospital and Science for Life Laboratory provided assistance in mass spectrometry and data analysis.
Author information
Authors and Affiliations
Contributions
N.W. performed the immunostaining staining on human and mouse muscle, generated reagents for and performed all luciferase experiments, immortalized and transdifferentiated human cell lines, performed immunoblot and RT-PCR experiments on C2C12 and FibroMyoD cells, and performed RT-PCR on mouse tissues. A.V. generated the Dup2 mouse model, injected all mice and performed all immublotting on mouse tissue and on muscle biopsies from the subjects with Dup2 and frameshift mutations. B.M. generated the master exon 1–5 construct and performed the initial dual luciferase assays. K.N.H. and L.R.R.-K. performed or analyzed the specific force and eccentric contraction–induced damage tests. L.Y. developed and performed the H&E and EBD quantification. A. Ferlini, C.A.-K.S. and M.U. designed and performed analysis of the exon 2–deleted muscle. S.M., G.V. and C.P. clinically characterized the subject with exon 2 deletion, and S.B., M.B., and M.N. performed the genetic characterization of the patient with exon 2 deletion. M.S.F. performed cell and AON studies on a subject with exon 2 deletion. S.D.W. provided critical reagents (AONs) for cell-based experiments. B.B. and D.R.S. designed and performed the formaldehyde RNA electrophoresis and northern blotting. D.M.D. and R.B.W. designed and performed the ribosome profiling experiments. A. Findlay generated all the U7 constructs. N.W., M.T.H. and K.M.F. designed the experiments and analyzed and interpreted the data. N.W., R.B.W. and K.M.F. wrote the manuscript and the compiled the figures, with contributions from A.V., F.G., R.B.W., A. Ferlini and D.R.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Table 1 (PDF 18173 kb)
Rights and permissions
About this article
Cite this article
Wein, N., Vulin, A., Falzarano, M. et al. Translation from a DMD exon 5 IRES results in a functional dystrophin isoform that attenuates dystrophinopathy in humans and mice. Nat Med 20, 992–1000 (2014). https://doi.org/10.1038/nm.3628
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3628
This article is cited by
-
Duchenne muscular dystrophy
Nature Reviews Disease Primers (2021)
-
Small mutations in Duchenne/Becker muscular dystrophy in 164 unrelated Polish patients
Journal of Applied Genetics (2021)
-
Identification of therapeutics that target eEF1A2 and upregulate utrophin A translation in dystrophic muscles
Nature Communications (2020)
-
EMQN best practice guidelines for genetic testing in dystrophinopathies
European Journal of Human Genetics (2020)
-
Becker muscular dystrophy caused by exon 2-truncating mutation of DMD
Human Genome Variation (2019)