Abstract
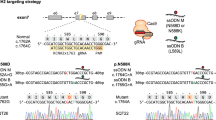
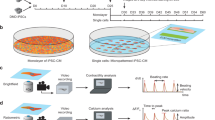
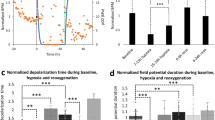
Study of monogenic mitochondrial cardiomyopathies may yield insights into mitochondrial roles in cardiac development and disease. Here, we combined patient-derived and genetically engineered induced pluripotent stem cells (iPSCs) with tissue engineering to elucidate the pathophysiology underlying the cardiomyopathy of Barth syndrome (BTHS), a mitochondrial disorder caused by mutation of the gene encoding tafazzin (TAZ). Using BTHS iPSC-derived cardiomyocytes (iPSC-CMs), we defined metabolic, structural and functional abnormalities associated with TAZ mutation. BTHS iPSC-CMs assembled sparse and irregular sarcomeres, and engineered BTHS 'heart-on-chip' tissues contracted weakly. Gene replacement and genome editing demonstrated that TAZ mutation is necessary and sufficient for these phenotypes. Sarcomere assembly and myocardial contraction abnormalities occurred in the context of normal whole-cell ATP levels. Excess levels of reactive oxygen species mechanistically linked TAZ mutation to impaired cardiomyocyte function. Our study provides new insights into the pathogenesis of Barth syndrome, suggests new treatment strategies and advances iPSC-based in vitro modeling of cardiomyopathy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Bione, S. et al. A novel X-linked gene, G4.5. is responsible for Barth syndrome. Nat. Genet. 12, 385–389 (1996).
Houtkooper, R.H. et al. The enigmatic role of tafazzin in cardiolipin metabolism. Biochim. Biophys. Acta 1788, 2003–2014 (2009).
Chicco, A.J. & Sparagna, G.C. Role of cardiolipin alterations in mitochondrial dysfunction and disease. Am. J. Physiol. Cell Physiol. 292, C33–C44 (2007).
Karikó, K., Buckstein, M., Ni, H. & Weissman, D. Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 23, 165–175 (2005).
Grosberg, A., Alford, P.W., McCain, M.L. & Parker, K.K. Ensembles of engineered cardiac tissues for physiological and pharmacological study: heart on a chip. Lab Chip 11, 4165–4173 (2011).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Warren, L. et al. Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7, 618–630 (2010).
Warren, L., Ni, Y., Wang, J. & Guo, X. Feeder-free derivation of human induced pluripotent stem cells with messenger RNA. Sci. Rep. 2, 657 (2012).
Shiba, Y. et al. Human ES-cell-derived cardiomyocytes electrically couple and suppress arrhythmias in injured hearts. Nature 489, 322–325 (2012).
Uosaki, H. et al. Efficient and scalable purification of cardiomyocytes from human embryonic and induced pluripotent stem cells by VCAM1 surface expression. PLoS ONE 6, e23657 (2011).
Elliott, D.A. et al. NKX2–5(eGFP/w) hESCs for isolation of human cardiac progenitors and cardiomyocytes. Nat. Methods 8, 1037–1040 (2011).
Schlame, M. et al. Phospholipid abnormalities in children with Barth syndrome. J. Am. Coll. Cardiol. 42, 1994–1999 (2003).
Kulik, W. et al. Bloodspot assay using HPLC-tandem mass spectrometry for detection of Barth syndrome. Clin. Chem. 54, 371–378 (2008).
Rana, P., Anson, B., Engle, S. & Will, Y. Characterization of human-induced pluripotent stem cell-derived cardiomyocytes: bioenergetics and utilization in safety screening. Toxicol. Sci. 130, 117–131 (2012).
Karikó, K. et al. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol. Ther. 16, 1833–1840 (2008).
Zangi, L. et al. Modified mRNA directs the fate of heart progenitor cells and induces vascular regeneration after myocardial infarction. Nat. Biotechnol. 31, 898–907 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Hom, J.R. et al. The permeability transition pore controls cardiac mitochondrial maturation and myocyte differentiation. Dev. Cell 21, 469–478 (2011).
Gerdes, A.M. & Capasso, J.M. Structural remodeling and mechanical dysfunction of cardiac myocytes in heart failure. J. Mol. Cell. Cardiol. 27, 849–856 (1995).
Agarwal, A., Goss, J.A., Cho, A., McCain, M.L. & Parker, K.K. Microfluidic heart on a chip for higher throughput pharmacological studies. Lab Chip 13, 3599–3608 (2013).
Feinberg, A.W. et al. Muscular thin films for building actuators and powering devices. Science 317, 1366–1370 (2007).
Domian, I.J. et al. Generation of functional ventricular heart muscle from mouse ventricular progenitor cells. Science 326, 426–429 (2009).
Parker, K.K., Tan, J., Chen, C.S. & Tung, L. Myofibrillar architecture in engineered cardiac myocytes. Circ. Res. 103, 340–342 (2008).
Alford, P.W., Feinberg, A.W., Sheehy, S.P. & Parker, K.K. Biohybrid thin films for measuring contractility in engineered cardiovascular muscle. Biomaterials 31, 3613–3621 (2010).
Malhotra, A. et al. Role of calcium-independent phospholipase A2 in the pathogenesis of Barth syndrome. Proc. Natl. Acad. Sci. USA 106, 2337–2341 (2009).
Valianpour, F. et al. Linoleic acid supplementation of Barth syndrome fibroblasts restores cardiolipin levels: implications for treatment. J. Lipid Res. 44, 560–566 (2003).
Rigaud, C. et al. Natural history of Barth syndrome: a national cohort study of 22 patients. Orphanet J. Rare Dis. 8, 70 (2013).
Steinberg, S.F. Oxidative stress and sarcomeric proteins. Circ. Res. 112, 393–405 (2013).
Sun, N. et al. Patient-specific induced pluripotent stem cells as a model for familial dilated cardiomyopathy. Sci. Transl. Med. 4, 130ra47 (2012).
Lan, F. et al. Abnormal calcium handling properties underlie familial hypertrophic cardiomyopathy pathology in patient-specific induced pluripotent stem cells. Cell Stem Cell 12, 101–113 (2013).
Park, I.H. et al. Disease-specific induced pluripotent stem cells. Cell 134, 877–886 (2008).
Chan, E.M. et al. Live cell imaging distinguishes bona fide human iPS cells from partially reprogrammed cells. Nat. Biotechnol. 27, 1033–1037 (2009).
Ichida, F. et al. Novel gene mutations in patients with left ventricular noncompaction or Barth syndrome. Circulation 103, 1256–1263 (2001).
Whitman, G.J. et al. Diagnosis and therapeutic evaluation of a pediatric case of cardiomyopathy using phosphorus-31 nuclear magnetic resonance spectroscopy. J. Am. Coll. Cardiol. 5, 745–749 (1985).
Chen, Y. & Dorn, G.W.n. PINK1-phosphorylated mitofusin 2 is a Parkin receptor for culling damaged mitochondria. Science 340, 471–475 (2013).
He, A., Kong, S.W., Ma, Q. & Pu, W.T. Co-occupancy by multiple cardiac transcription factors identifies transcriptional enhancers active in heart. Proc. Natl. Acad. Sci. USA 108, 5632–5637 (2011).
Houtkooper, R.H. et al. Cardiolipin and monolysocardiolipin analysis in fibroblasts, lymphocytes, and tissues using high-performance liquid chromatography-mass spectrometry as a diagnostic test for Barth syndrome. Anal. Biochem. 15, 230–237 (2009).
Bray, M.A., Sheehy, S.P. & Parker, K.K. Sarcomere alignment is regulated by myocyte shape. Cell Motil. Cytoskeleton 65, 641–651 (2008).
Chen, C.S., Mrksich, M., Huang, S., Whitesides, G.M. & Ingber, D.E. Geometric control of cell life and death. Science 276, 1425–1428 (1997).
Sato, Y. et al. Three-dimensional multi-scale line filter for segmentation and visualization of curvilinear structures in medical images. Med. Image Anal. 2, 143–168 (1998).
Melkman, A.A. On-line construction of the convex hull of a simple polyline. Inf. Process. Lett. 25, 11–12 (1987).
Wei, S. et al. T-tubule remodeling during transition from hypertrophy to heart failure. Circ. Res. 107, 520–531 (2010).
Timoshenko, S. & Woinowsky-Krieger, S. in Engineering Societies Monographs 5 (McGraw-Hill, 1959).
Stoney, G.G. The tension of metallic films deposited by electrolysis. Proc. R. Soc. Lond. 82, 172–175 (1909).
Acknowledgements
This work supported by the Barth Syndrome Foundation, the Boston Children's Hospital Translational Investigator Service, the US National Institutes of Health (NIH) NHLBI Progenitor Cell Biology Consortium (NIH U01 HL100401 and U01 HL100408), NIH RC1 HL099618, NIH UH2 TR000522 and charitable donations from E. Marram, K. Carpenter and G.F. Smith.
Author information
Authors and Affiliations
Contributions
G.W. designed and performed experiments and analyzed data. M.L.M. designed and performed experiments on MTFs and sarcomere organization and analyzed data. L.Y. and G.M.C. provided expert assistance and reagents for genome editing. F.S.P. designed the sarcomere organization analysis method. H.Y. developed the MTF analysis method. A.A. assisted with MTF substrate fabrication and experiments. D.J. provided advice on mitochondrial assays. D.Z. imaged iPSC-CMs to assess their mitochondrial organization and potential. L.Z. and K.R.C. provided expert assistance with modRNA, and J.C., J.D. and D.-Z.W. helped construct modRNAs. K.L. contributed to genome editing. R.J.A.W., W.K. and F.M.V. analyzed phospholipids. M.A.L. and C.E.M. provided expert assistance in iPSC differentiation to cardiomyocytes. A.H. developed TAZ shRNA viruses and provided technical assistance. J.G. and A.E.R. obtained patient samples. Q.M. assisted in teratoma analysis. J.W. contributed control iPSC lines. R.I.K. provided expert input, patient samples and 31P nuclear magnetic resonance data. K.K.P. and W.T.P. supervised the study. W.T.P. wrote the manuscript, and it was revised by K.K.P., G.W. and M.L.M.
Corresponding authors
Ethics declarations
Competing interests
J.W. is an employee of Allele Biotechnology & Pharmaceuticals.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–4 and Supplementary Figures 1–14 (PDF 8956 kb)
Supplementary Data Set
Supplementary Data Sets (XLSX 82 kb)
Supplementary Movies
Supplementary Movies 1–6 (PDF 12265 kb)
Rights and permissions
About this article
Cite this article
Wang, G., McCain, M., Yang, L. et al. Modeling the mitochondrial cardiomyopathy of Barth syndrome with induced pluripotent stem cell and heart-on-chip technologies. Nat Med 20, 616–623 (2014). https://doi.org/10.1038/nm.3545
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3545
This article is cited by
-
Preclinical Models of Cardiac Disease: A Comprehensive Overview for Clinical Scientists
Cardiovascular Engineering and Technology (2024)
-
Fibre-infused gel scaffolds guide cardiomyocyte alignment in 3D-printed ventricles
Nature Materials (2023)
-
Functional analysis of a common BAG3 allele associated with protection from heart failure
Nature Cardiovascular Research (2023)
-
Recent advances in pluripotent stem cell-derived cardiac organoids and heart-on-chip applications for studying anti-cancer drug-induced cardiotoxicity
Cell Biology and Toxicology (2023)
-
Application of Human Induced Pluripotent Stem Cells for Tissue Engineered Cardiomyocyte Modelling
Regenerative Engineering and Translational Medicine (2023)