Abstract
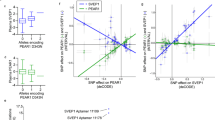
Racial differences in the pathophysiology of atherothrombosis are poorly understood. We explored the function and transcriptome of platelets in healthy black (n = 70) and white (n = 84) subjects. Platelet aggregation and calcium mobilization induced by the PAR4 thrombin receptor were significantly greater in black subjects. Numerous differentially expressed RNAs were associated with both race and PAR4 reactivity, including PCTP (encoding phosphatidylcholine transfer protein), and platelets from black subjects expressed higher levels of PC-TP protein. PC-TP inhibition or depletion blocked PAR4- but not PAR1-mediated activation of platelets and megakaryocytic cell lines. miR-376c levels were differentially expressed by race and PAR4 reactivity and were inversely correlated with PCTP mRNA levels, PC-TP protein levels and PAR4 reactivity. miR-376c regulated the expression of PC-TP in human megakaryocytes. A disproportionately high number of microRNAs that were differentially expressed by race and PAR4 reactivity, including miR-376c, are encoded in the DLK1-DIO3 locus and were expressed at lower levels in platelets from black subjects. These results suggest that PC-TP contributes to the racial difference in PAR4-mediated platelet activation, indicate a genomic contribution to platelet function that differs by race and emphasize a need to consider the effects of race when developing anti-thrombotic drugs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Libby, P. Mechanisms of acute coronary syndromes and their implications for therapy. N. Engl. J. Med. 368, 2004–2013 (2013).
Leger, A.J., Covic, L. & Kuliopulos, A. Protease-activated receptors in cardiovascular diseases. Circulation 114, 1070–1077 (2006).
Abrams, C.S. & Brass, L.F. Platelet signal transduction. in Hemostasis and Thrombosis: Basic Principles and Clinical Practice (eds. Colman, R.W., Hirsh, J., Marder, V.J., Clowes, A.W. & George, J.N.) 617–629 (Lippincott Williams & Wilkins, Philadelphia, PA, 2006).
Macfarlane, S.R., Seatter, M.J., Kanke, T., Hunter, G.D. & Plevin, R. Proteinase-activated receptors. Pharmacol. Rev. 53, 245–282 (2001).
Lova, P. et al. Contribution of protease-activated receptors 1 and 4 and glycoprotein Ib-IX-V in the G(i)-independent activation of platelet Rap1B by thrombin. J. Biol. Chem. 279, 25299–25306 (2004).
Henriksen, R.A. & Hanks, V.K. PAR-4 agonist AYPGKF stimulates thromboxane production by human platelets. Arterioscler. Thromb. Vasc. Biol. 22, 861–866 (2002).
Holinstat, M. et al. PAR4, but not PAR1, signals human platelet aggregation via Ca2+ mobilization and synergistic P2Y12 receptor activation. J. Biol. Chem. 281, 26665–26674 (2006).
O'Donnell, C.J. et al. Genetic and environmental contributions to platelet aggregation: the Framingham Heart Study. Circulation 103, 3051–3056 (2001).
Bray, P.F. et al. Heritability of platelet function in families with premature coronary artery disease. J. Thromb. Haemost. 5, 1617–1623 (2007).
Thomas, K.L., Honeycutt, E., Shaw, L.K. & Peterson, E.D. Racial differences in long-term survival among patients with coronary artery disease. Am. Heart J. 160, 744–751 (2010).
Berry, J.D. et al. Lifetime risks of cardiovascular disease. N. Engl. J. Med. 366, 321–329 (2012).
Quinton, T.M., Kim, S., Derian, C.K., Jin, J. & Kunapuli, S.P. Plasmin-mediated activation of platelets occurs by cleavage of protease-activated receptor 4. J. Biol. Chem. 279, 18434–18439 (2004).
Nagalla, S. et al. Platelet microRNA-mRNA coexpression profiles correlate with platelet reactivity. Blood 117, 5189–5197 (2011).
Benjamini, Y. & Hochberg, Y. Controlling for the false discovery rate: a practical and powerful approach to multiple testing. J.R. Stat. Soc. 57, 289–300 (1995).
Zhang, W. et al. Evaluation of genetic variation contributing to differences in gene expression between populations. Am. J. Hum. Genet. 82, 631–640 (2008).
Kang, H.W., Wei, J. & Cohen, D.E. PC-TP/StARD2: of membranes and metabolism. Trends Endocrinol. Metab. 21, 449–456 (2010).
Xu, Q. et al. Investigation of variation in gene expression profiling of human blood by extended principle component analysis. PLoS One 6, e26905 (2011).
van Helvoort, A. et al. Mice without phosphatidylcholine transfer protein have no defects in the secretion of phosphatidylcholine into bile or into lung airspaces. Proc. Natl. Acad. Sci. USA 96, 11501–11506 (1999).
Rowley, J.W. et al. Genome-wide RNA-seq analysis of human and mouse platelet transcriptomes. Blood 118, e101–e111 (2011).
Wagle, N. et al. Small-molecule inhibitors of phosphatidylcholine transfer protein/StarD2 identified by high-throughput screening. Anal. Biochem. 383, 85–92 (2008).
Shishova, E.Y. et al. Genetic ablation or chemical inhibition of phosphatidylcholine transfer protein attenuates diet-induced hepatic glucose production. Hepatology 54, 664–674 (2011).
Ozaki, Y. et al. Thrombin-induced calcium oscillation in human platelets and MEG-01, a megakaryoblastic leukemia cell line. Biochem. Biophys. Res. Commun. 183, 864–871 (1992).
Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Guo, H., Ingolia, N.T., Weissman, J.S. & Bartel, D.P. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 466, 835–840 (2010).
Kondkar, A.A. et al. VAMP8/endobrevin is overexpressed in hyperreactive human platelets: suggested role for platelet microRNA. J. Thromb. Haemost. 8, 369–378 (2010).
Goodall, A.H. et al. Transcription profiling in human platelets reveals LRRFIP1 as a novel protein regulating platelet function. Blood 116, 4646–4656 (2010).
Cho, J.H. et al. Increased calcium stores in platelets from African Americans. Hypertension 25, 377–383 (1995).
Tang, H. et al. Genetic structure, self-identified race/ethnicity, and confounding in case-control association studies. Am. J. Hum. Genet. 76, 268–275 (2005).
Rosenberg, N.A. et al. Genetic structure of human populations. Science 298, 2381–2385 (2002).
Mountain, J.L. & Cavalli-Sforza, L.L. Multilocus genotypes, a tree of individuals, and human evolutionary history. Am. J. Hum. Genet. 61, 705–718 (1997).
Risch, N., Burchard, E., Ziv, E. & Tang, H. Categorization of humans in biomedical research: genes, race and disease. Genome Biol. 3, comment2007 (2002).
Tishkoff, S.A. et al. The genetic structure and history of Africans and African Americans. Science 324, 1035–1044 (2009).
Chahrour, M. et al. MeCP2, a key contributor to neurological disease, activates and represses transcription. Science 320, 1224–1229 (2008).
Schrick, K., Nguyen, D., Karlowski, W.M. & Mayer, K.F. START lipid/sterol-binding domains are amplified in plants and are predominantly associated with homeodomain transcription factors. Genome Biol. 5, R41 (2004).
Geijtenbeek, T.B., Smith, A.J., Borst, P. & Wirtz, K.W. cDNA cloning and tissue-specific expression of the phosphatidylcholine transfer protein gene. Biochem. J. 316, 49–55 (1996).
Plé, H. et al. Alteration of the platelet transcriptome in chronic kidney disease. Thromb. Haemost. 108, 605–615 (2012).
Baez, J.M., Tabas, I. & Cohen, D.E. Decreased lipid efflux and increased susceptibility to cholesterol-induced apoptosis in macrophages lacking phosphatidylcholine transfer protein. Biochem. J. 388, 57–63 (2005).
Lev, S. Non-vesicular lipid transport by lipid-transfer proteins and beyond. Nat. Rev. Mol. Cell Biol. 11, 739–750 (2010).
Mahadevappa, V.G. & Holub, B.J. Relative degradation of different molecular species of phosphatidylcholine in thrombin-stimulated human platelets. J. Biol. Chem. 259, 9369–9373 (1984).
Exton, J.H. Signaling through phosphatidylcholine breakdown. J. Biol. Chem. 265, 1–4 (1990).
O'Brien, K.A., Stojanovic-Terpo, A., Hay, N. & Du, X. An important role for Akt3 in platelet activation and thrombosis. Blood 118, 4215–4223 (2011).
Benetatos, L. et al. The microRNAs within the DLK1–DIO3 genomic region: involvement in disease pathogenesis. Cell. Mol. Life Sci. 70, 795–814 (2013).
Liu, L. et al. Activation of the imprinted Dlk1-Dio3 region correlates with pluripotency levels of mouse stem cells. J. Biol. Chem. 285, 19483–19490 (2010).
Wallace, C. et al. The imprinted DLK1–MEG3 gene region on chromosome 14q32.2 alters susceptibility to type 1 diabetes. Nat. Genet. 42, 68–71 (2010).
Fiore, R. et al. Mef2-mediated transcription of the miR379–410 cluster regulates activity-dependent dendritogenesis by fine-tuning Pumilio2 protein levels. EMBO J. 28, 697–710 (2009).
Song, G. & Wang, L. Transcriptional mechanism for the paired miR-433 and miR-127 genes by nuclear receptors SHP and ERRγ. Nucleic Acids Res. 36, 5727–5735 (2008).
Edelstein, L.C. & Bray, P.F. MicroRNAs in platelet production and activation. Blood 117, 5289–5296 (2011).
Phimister, E.G. Medicine and the racial divide. N. Engl. J. Med. 348, 1081–1082 (2003).
Morrow, D.A. et al. Vorapaxar in the secondary prevention of atherothrombotic events. N. Engl. J. Med. 366, 1404–1413 (2012).
Bonaca, M.P. et al. Vorapaxar in patients with peripheral artery disease: results from TRA2°P-TIMI 50. Circulation 127, 1522–1529 (2013).
Scirica, B.M. et al. Vorapaxar for secondary prevention of thrombotic events for patients with previous myocardial infarction: a prespecified subgroup analysis of the TRA 2 degrees P-TIMI 50 trial. Lancet 380, 1317–1324 (2012).
Vergnolle, N. Protease-activated receptors as drug targets in inflammation and pain. Pharmacol. Ther. 123, 292–309 (2009).
Yee, D.L., Sun, C.W., Bergeron, A.L., Dong, J.F. & Bray, P.F. Aggregometry detects platelet hyperreactivity in healthy individuals. Blood 106, 2723–2729 (2005).
Yeung, J. et al. Protein kinase C regulation of 12-lipoxygenase–mediated human platelet activation. Mol. Pharmacol. 81, 420–430 (2012).
Fryer, J.D. et al. Exercise and genetic rescue of SCA1 via the transcriptional repressor Capicua. Science 334, 690–693 (2011).
Geiss, G.K. et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat. Biotechnol. 26, 317–325 (2008).
Tili, E. et al. The down-regulation of miR-125b in chronic lymphocytic leukemias leads to metabolic adaptation of cells to a transformed state. Blood 120, 2631–2638 (2012).
Gantner, B.N. et al. The Akt1 isoform is required for optimal IFN-β transcription through direct phosphorylation of β-catenin. J. Immunol. 189, 3104–3111 (2012).
Patel, S.R., Hartwig, J.H. & Italiano, J.E. Jr. The biogenesis of platelets from megakaryocyte proplatelets. J. Clin. Invest. 115, 3348–3354 (2005).
Gentleman, R.I.R.R. A language for data analysis and graphics. J. Comput. Stat. Graph. 5, 299–314 (1996).
Hochberg, Y. & Benjamini, Y. More powerful procedures for multiple significance testing. Stat. Med. 9, 811–818 (1990).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
1000 Genomes Project Consortium et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010); erratum 473, 544 (2011).
Acknowledgements
We thank S. McKenzie for helpful discussions, S. Kunapuli (Temple University) for the PAR1 inhibitor, J. Italiano (Harvard Medical School) for the antibody to tubulin, L. Ma for technical support, R. Baserga (Thomas Jefferson University) for the HCT116-Dicer knockout 2 cells and P. Yu for normalizing Affymetrix data. This work was supported by US National Institutes of Health (NIH) grant HL102482 (to P.F.B.) and the Cardeza Foundation for Hematologic Research. Compound A1 was developed with the support of NIH grants DK48873 and DK56626 (to D.E.C.).
Author information
Authors and Affiliations
Contributions
P.F.B. conceived the PRAX1 study. P.F.B. and C.S. designed the PRAX1 study. P.F.B., L.C.E., J.D., C.S. and S.N. supervised the project. L.C.E., L.M.S., E.S.C., R.T.M., M.H., N.M., D.E.C., J.D., C.S. and P.F.B. designed experiments. L.C.E., L.M.S., E.S.C., A.B., X.K., R.T.M. and M.H. collected data. L.C.E., L.M.S. and E.S.C. R.T.M., M.H., J.D., C.S. and P.F.B. analyzed data. L.C.E., E.S.C., D.E.C., C.S. and P.F.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1-5 and Supplementary Tables 1-5 (PDF 1641 kb)
Rights and permissions
About this article
Cite this article
Edelstein, L., Simon, L., Montoya, R. et al. Racial differences in human platelet PAR4 reactivity reflect expression of PCTP and miR-376c. Nat Med 19, 1609–1616 (2013). https://doi.org/10.1038/nm.3385
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3385
This article is cited by
-
Tumour-educated platelets for breast cancer detection: biological and technical insights
British Journal of Cancer (2023)
-
Toward platelet transcriptomics in cancer diagnosis, prognosis and therapy
British Journal of Cancer (2022)
-
Transcriptomic landscape of blood platelets in healthy donors
Scientific Reports (2021)
-
Platelet reactivity in response to aspirin and ticagrelor in African-Americans and European-Americans
Journal of Thrombosis and Thrombolysis (2021)
-
Platelet Function in Cardiovascular Disease: Activation of Molecules and Activation by Molecules
Cardiovascular Toxicology (2020)