Abstract
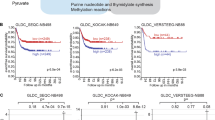
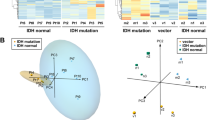
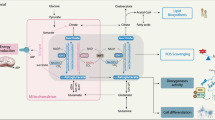
Here we show that glioblastoma express high levels of branched-chain amino acid transaminase 1 (BCAT1), the enzyme that initiates the catabolism of branched-chain amino acids (BCAAs). Expression of BCAT1 was exclusive to tumors carrying wild-type isocitrate dehydrogenase 1 (IDH1) and IDH2 genes and was highly correlated with methylation patterns in the BCAT1 promoter region. BCAT1 expression was dependent on the concentration of α-ketoglutarate substrate in glioma cell lines and could be suppressed by ectopic overexpression of mutant IDH1 in immortalized human astrocytes, providing a link between IDH1 function and BCAT1 expression. Suppression of BCAT1 in glioma cell lines blocked the excretion of glutamate and led to reduced proliferation and invasiveness in vitro, as well as significant decreases in tumor growth in a glioblastoma xenograft model. These findings suggest a central role for BCAT1 in glioma pathogenesis, making BCAT1 and BCAA metabolism attractive targets for the development of targeted therapeutic approaches to treat patients with glioblastoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Accessions
Ensembl
Gene Expression Omnibus
References
Lieth, E. et al. Nitrogen shuttling between neurons and glial cells during glutamate synthesis. J. Neurochem. 76, 1712–1723 (2001).
Hutson, S.M. The case for regulating indispensable amino acid metabolism: the branched-chain α-keto acid dehydrogenase kinase-knockout mouse. Biochem. J. 400, e1–e3 (2006).
Ichihara, A. & Koyama, E. Transaminase of branched chain amino acids. I. Branched chain amino acids–α-ketoglutarate transaminase. J. Biochem. 59, 160–169 (1966).
Taylor, R.T. & Jenkins, W.T. Leucine aminotransferase. II. Purification and characterization. J. Biol. Chem. 241, 4396–4405 (1966).
García-Espinosa, M.A., Wallin, R., Hutson, S.M. & Sweatt, A.J. Widespread neuronal expression of branched-chain aminotransferase in the CNS: implications for leucine/glutamate metabolism and for signaling by amino acids. J. Neurochem. 100, 1458–1468 (2007).
Hall, T.R., Wallin, R., Reinhart, G.D. & Hutson, S.M. Branched chain aminotransferase isoenzymes. Purification and characterization of the rat brain isoenzyme. J. Biol. Chem. 268, 3092–3098 (1993).
Sweatt, A.J. et al. Branched-chain amino acid catabolism: unique segregation of pathway enzymes in organ systems and peripheral nerves. Am. J. Physiol. Endocrinol. Metab. 286, E64–E76 (2004).
Hutson, S.M., Sweatt, A.J. & Lanoue, K.F. Branched-chain [corrected] amino acid metabolism: implications for establishing safe intakes. J. Nutr. 135 (suppl. 6), 1557S–1564S (2005).
Eagle, H. Nutrition needs of mammalian cells in tissue culture. Science 122, 501–514 (1955).
DeBerardinis, R.J. & Cheng, T. Q's next: the diverse functions of glutamine in metabolism, cell biology and cancer. Oncogene 29, 313–324 (2010).
DeBerardinis, R.J. et al. Beyond aerobic glycolysis: transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc. Natl. Acad. Sci. USA 104, 19345–19350 (2007).
Seltzer, M.J. et al. Inhibition of glutaminase preferentially slows growth of glioma cells with mutant IDH1. Cancer Res. 70, 8981–8987 (2010).
Wise, D.R. et al. Myc regulates a transcriptional program that stimulates mitochondrial glutaminolysis and leads to glutamine addiction. Proc. Natl. Acad. Sci. USA 105, 18782–18787 (2008).
Yang, C. et al. Glioblastoma cells require glutamate dehydrogenase to survive impairments of glucose metabolism or Akt signaling. Cancer Res. 69, 7986–7993 (2009).
Metallo, C.M. et al. Reductive glutamine metabolism by IDH1 mediates lipogenesis under hypoxia. Nature 481, 380–384 (2012).
Mullen, A.R. et al. Reductive carboxylation supports growth in tumour cells with defective mitochondria. Nature 481, 385–388 (2012).
Wise, D.R. et al. Hypoxia promotes isocitrate dehydrogenase–dependent carboxylation of α-ketoglutarate to citrate to support cell growth and viability. Proc. Natl. Acad. Sci. USA 108, 19611–19616 (2011).
Balss, J. et al. Analysis of the IDH1 codon 132 mutation in brain tumors. Acta Neuropathol. 116, 597–602 (2008).
Ichimura, K. et al. IDH1 mutations are present in the majority of common adult gliomas but rare in primary glioblastomas. Neuro-oncol. 11, 341–347 (2009).
Parsons, D.W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–1812 (2008).
Yan, H. et al. IDH1 and IDH2 mutations in gliomas. N. Engl. J. Med. 360, 765–773 (2009).
Brennan, C. et al. Glioblastoma subclasses can be defined by activity among signal transduction pathways and associated genomic alterations. PLoS ONE 4, e7752 (2009).
Colman, H. et al. A multigene predictor of outcome in glioblastoma. Neuro-oncol. 12, 49–57 (2010).
Noushmehr, H. et al. Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17, 510–522 (2010).
Sturm, D. et al. Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22, 425–437 (2012).
Toedt, G. et al. Molecular signatures classify astrocytic gliomas by IDH1 mutation status. Int. J. Cancer 128, 1095–1103 (2011).
Verhaak, R.G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010).
Hartmann, C. et al. Patients with IDH1 wild type anaplastic astrocytomas exhibit worse prognosis than IDH1-mutated glioblastomas, and IDH1 mutation status accounts for the unfavorable prognostic effect of higher age: implications for classification of gliomas. Acta Neuropathol. 120, 707–718 (2010).
Sasaki, M. et al. D-2-hydroxyglutarate produced by mutant IDH1 perturbs collagen maturation and basement membrane function. Genes Dev. 26, 2038–2049 (2012).
Sasaki, M. et al. IDH1(R132H) mutation increases murine haematopoietic progenitors and alters epigenetics. Nature 488, 656–659 (2012).
Dang, L., Jin, S. & Su, S.M. IDH mutations in glioma and acute myeloid leukemia. Trends Mol. Med. 16, 387–397 (2010).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009).
Figueroa, M.E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18, 553–567 (2010).
Lu, C. et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483, 474–478 (2012).
Xu, W. et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate–dependent dioxygenases. Cancer Cell 19, 17–30 (2011).
Schramm, G. et al. PathWave: discovering patterns of differentially regulated enzymes in metabolic pathways. Bioinformatics 26, 1225–1231 (2010).
Gravendeel, L.A. et al. Intrinsic gene expression profiles of gliomas are a better predictor of survival than histology. Cancer Res. 69, 9065–9072 (2009).
Turcan, S. et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483, 479–483 (2012).
Reitman, Z.J. & Yan, H. Isocitrate dehydrogenase 1 and 2 mutations in cancer: alterations at a crossroads of cellular metabolism. J. Natl. Cancer Inst. 102, 932–941 (2010).
She, P. et al. Disruption of BCATm in mice leads to increased energy expenditure associated with the activation of a futile protein turnover cycle. Cell Metab. 6, 181–194 (2007).
Goto, M. et al. Structural determinants for branched-chain aminotransferase isozyme-specific inhibition by the anticonvulsant drug gabapentin. J. Biol. Chem. 280, 37246–37256 (2005).
Sontheimer, H. A role for glutamate in growth and invasion of primary brain tumors. J. Neurochem. 105, 287–295 (2008).
Pasquali, M., Monsen, G., Richardson, L., Alston, M. & Longo, N. Biochemical findings in common inborn errors of metabolism. Am. J. Med. Genet. C. Semin. Med. Genet. 142C, 64–76 (2006).
Koppenol, W.H., Bounds, P.L. & Dang, C.V. Otto Warburg's contributions to current concepts of cancer metabolism. Nat. Rev. Cancer 11, 325–337 (2011).
de Bont, J.M. et al. Differential expression and prognostic significance of SOX genes in pediatric medulloblastoma and ependymoma identified by microarray analysis. Neuro-oncol. 10, 648–660 (2008).
Goto, M., Shinno, H. & Ichihara, A. Isozyme patterns of branched-chain amino acid transaminase in human tissues and tumors. Gann 68, 663–667 (1977).
Weggen, S. et al. Identification of amplified genes from SV40 large T antigen–induced rat PNET cell lines by subtractive cDNA analysis and radiation hybrid mapping. Oncogene 20, 2023–2031 (2001).
Yoshikawa, R. et al. ECA39 is a novel distant metastasis-related biomarker in colorectal cancer. World J. Gastroenterol. 12, 5884–5889 (2006).
Zhou, W. et al. Functional evidence for a nasopharyngeal carcinoma-related gene BCAT1 located at 12p12. Oncol. Res. 16, 405–413 (2007).
Reitman, Z.J. et al. Profiling the effects of isocitrate dehydrogenase 1 and 2 mutations on the cellular metabolome. Proc. Natl. Acad. Sci. USA 108, 3270–3275 (2011).
Zhao, S. et al. Glioma-derived mutations in IDH1 dominantly inhibit IDH1 catalytic activity and induce HIF-1α. Science 324, 261–265 (2009).
Davoodi, J. et al. Overexpression and characterization of the human mitochondrial and cytosolic branched-chain aminotransferases. J. Biol. Chem. 273, 4982–4989 (1998).
Locasale, J.W. et al. Phosphoglycerate dehydrogenase diverts glycolytic flux and contributes to oncogenesis. Nat. Genet. 43, 869–874 (2011).
Possemato, R. et al. Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature 476, 346–350 (2011).
Jain, M. et al. Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation. Science 336, 1040–1044 (2012).
Hartmann, C. et al. Type and frequency of IDH1 and IDH2 mutations are related to astrocytic and oligodendroglial differentiation and age: a study of 1,010 diffuse gliomas. Acta Neuropathol. 118, 469–474 (2009).
Capper, D., Zentgraf, H., Balss, J., Hartmann, C. & von Deimling, A. Monoclonal antibody specific for IDH1 R132H mutation. Acta Neuropathol. 118, 599–601 (2009).
Buckingham, S.C. et al. Glutamate release by primary brain tumors induces epileptic activity. Nat. Med. 17, 1269–1274 (2011).
Grzendowski, M. et al. Simultaneous extraction of nucleic acids and proteins from tissue specimens by ultracentrifugation: a protocol using the high-salt protein fraction for quantitative proteome analysis. Proteomics 9, 4985–4990 (2009).
Ehrich, M. et al. Quantitative high-throughput analysis of DNA methylation patterns by base-specific cleavage and mass spectrometry. Proc. Natl. Acad. Sci. USA 102, 15785–15790 (2005).
Campos, B. et al. Differentiation therapy exerts antitumor effects on stem-like glioma cells. Clin. Cancer Res. 16, 2715–2728 (2010).
Turcan, S. et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483, 479–483 (2012).
Wiederschain, D. et al. Single-vector inducible lentiviral RNAi system for oncology target validation. Cell Cycle 8, 498–504 (2009).
Bai, A.H. et al. MicroRNA-182 promotes leptomeningeal spread of non-sonic hedgehog-medulloblastoma. Acta Neuropathol. 123, 529–538 (2012).
Rolli, C.G., Seufferlein, T., Kemkemer, R. & Spatz, J.P. Impact of tumor cell cytoskeleton organization on invasiveness and migration: a microchannel-based approach. PLoS ONE 5, e8726 (2010).
Sze, D.Y. & Jardetzky, O. Determination of metabolite and nucleotide concentrations in proliferating lymphocytes by 1H-NMR of acid extracts. Biochim. Biophys. Acta 1054, 181–197 (1990).
Lowry, O.H., Rosebrough, N.J., Farr, A.L. & Randall, R.J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193, 265–275 (1951).
Helenius, A. & Simons, K. The binding of detergents to lipophilic and hydrophilic proteins. J. Biol. Chem. 247, 3656–3661 (1972).
Sauer, S.W. et al. Intracerebral accumulation of glutaric and 3-hydroxyglutaric acids secondary to limited flux across the blood-brain barrier constitute a biochemical risk factor for neurodegeneration in glutaryl-CoA dehydrogenase deficiency. J. Neurochem. 97, 899–910 (2006).
She, P. et al. Disruption of BCATm in mice leads to increased energy expenditure associated with the activation of a futile protein turnover cycle. Cell Metab. 6, 181–194 (2007).
Suryawan, A. et al. A molecular model of human branched-chain amino acid metabolism. Am. J. Clin. Nutr. 68, 72–81 (1998).
Hutson, S.M. et al. Role of branched-chain aminotransferase isoenzymes and gabapentin in neurotransmitter metabolism. J. Neurochem. 71, 863–874 (1998).
Acknowledgements
We thank G. Tödt for bioinformatic expertise, M. Kirchgäßner, K. Pfleger, E. Wieland, A.-C. Klein, S. Emmerich and the DKFZ Light Microscopy Facility for excellent technical support, C. Schmidt (DKFZ) for providing the HEY1-pDest-vectors and A. von Deimling (University of Heidelberg) for providing the IDH1-specific antibodies. This work was supported by the German Federal Ministry of Education and Research (BMBF) within the National Genome Research Network NGFNplus (01GS0883 and 01GS0884) to B.R., P.L. and G.R., the German Cancer Aid (Deutsche Krebshilfe, grant number 108456) to B.R. and P.L. and the intramural funding program of the National Tumor Center Heidelberg. Y.J.P. was supported by the Roman Herzog research fellowship from the Hertie Foundation, Germany, and the Korean Health Technology R&D Project, Ministry of Health and Welfare, Republic of Korea (A120071-1211-0000200).
Author information
Authors and Affiliations
Contributions
M.T. and S.B. developed the concept, designed and performed experiments, analyzed data and wrote the manuscript. Y.J.P. and A.M.L. conducted the promoter methylation analysis, chromatin immunoprecipitation analysis and HEY1 knockdown. W.W. conducted dimethyl–α-KG and hypoxia experiments. M.S. assisted with experiments. S.V.P. conducted lentiviral knockdown in NCH421k cells. A.H.C.B. assisted with microchannel experiments. D.K. conducted western blot analysis of patient samples. R.M.P. and R. König mapped metabolic pathways. J.F. performed IDH1 mutation studies and immunohistochemical analyses. D.L. performed animal experiments. I.W., V.H., S.T., T.A.C., D.S., H.W. and S.M.P. assisted with the methylation analysis. A.A. and S.M.H. performed and interpreted BCAT1 inhibition experiments. C.G.R. and R. Kemkemer synthesized the microchannel chips. B.C. and C.H.-M. analyzed tissue microarray data. K.S. conducted the MS/MS and gas chromatography–mass spectrometry. W.-E.H. performed the 1H-NMR spectroscopy. M.J. performed magnetic resonance imaging. C.P. was involved in the methylation analysis. J.G.O. analyzed the mass spectrometry data. G.R. performed histological and immunohistochemical analyses and contributed to writing the manuscript. P.L. oversaw all research phases and contributed to writing the manuscript. B.R. developed the concept, supervised the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Tables 1–5 and Supplementary Methods (PDF 1918 kb)
Supplementary Video 1
Migration of U-87MG cells transduced with nontarget shRNA (AVI 13511 kb)
Supplementary Video 2
Migration of U-87MG cells transduced with BCAT1 shRNAI (AVI 14761 kb)
Rights and permissions
About this article
Cite this article
Tönjes, M., Barbus, S., Park, Y. et al. BCAT1 promotes cell proliferation through amino acid catabolism in gliomas carrying wild-type IDH1. Nat Med 19, 901–908 (2013). https://doi.org/10.1038/nm.3217
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3217
This article is cited by
-
HuR promotes castration-resistant prostate cancer progression by altering ERK5 activation via posttranscriptional regulation of BCAT1
Journal of Translational Medicine (2024)
-
Amino acid metabolism in tumor biology and therapy
Cell Death & Disease (2024)
-
High expression of BCAT1 sensitizes AML cells to PARP inhibitor by suppressing DNA damage response
Journal of Molecular Medicine (2024)
-
BCAT1 controls embryonic neural stem cells proliferation and differentiation in the upper layer neurons
Molecular Brain (2023)
-
Inhibition of branched-chain alpha-keto acid dehydrogenase kinase augments the sensitivity of ovarian and breast cancer cells to paclitaxel
British Journal of Cancer (2023)