Abstract
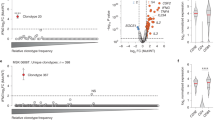
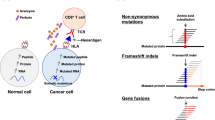
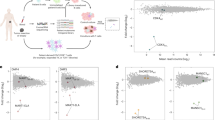
Substantial regressions of metastatic lesions have been observed in up to 70% of patients with melanoma who received adoptively transferred autologous tumor-infiltrating lymphocytes (TILs) in phase 2 clinical trials1,2. In addition, 40% of patients treated in a recent trial experienced complete regressions of all measurable lesions for at least 5 years following TIL treatment3. To evaluate the potential association between the ability of TILs to mediate durable regressions and their ability to recognize potent antigens that presumably include mutated gene products, we developed a new screening approach involving mining whole-exome sequence data to identify mutated proteins expressed in patient tumors. We then synthesized and evaluated candidate mutated T cell epitopes that were identified using a major histocompatibility complex–binding algorithm4 for recognition by TILs. Using this approach, we identified mutated antigens expressed on autologous tumor cells that were recognized by three bulk TIL lines from three individuals with melanoma that were associated with objective tumor regressions following adoptive transfer. This simplified approach for identifying mutated antigens recognized by T cells avoids the need to generate and laboriously screen cDNA libraries from tumors and may represent a generally applicable method for identifying mutated antigens expressed in a variety of tumor types.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
14 May 2013
In the version of this article initially published online, the second sentence of the abstract stated, “In addition, 40% of patients treated in a recent trial experienced complete regressions of all measurable lesions lasting between 5 and 9 years after treatment3.” The correct statement should read, “In addition, 40% of patients treated in a recent trial experienced complete regressions of all measurable lesions for at least 5 years following TIL treatment3.” The error has been corrected for the print, PDF and HTML versions of this article.
References
Dudley, M.E. et al. Cancer regression and autoimmunity in patients after clonal repopulation with antitumor lymphocytes. Science 298, 850–854 (2002).
Dudley, M.E. et al. Adoptive cell therapy for patients with metastatic melanoma: evaluation of intensive myeloablative chemoradiation preparative regimens. J. Clin. Oncol. 26, 5233–5239 (2008).
Rosenberg, S.A. et al. Durable complete responses in heavily pretreated patients with metastatic melanoma using T-cell transfer immunotherapy. Clin. Cancer Res. 17, 4550–4557 (2011).
Nielsen, M. et al. NetMHCpan, a method for quantitative predictions of peptide binding to any HLA-A and -B locus protein of known sequence. PLoS ONE 2, e796 (2007).
Marincola, F.M. et al. Locus-specific analysis of human leukocyte antigen class I expression in melanoma cell lines. J. Immunother. Emphasis Tumor Immunol. 16, 13–23 (1994).
Salter, R.D. & Cresswell, P. Impaired assembly and transport of HLA-A and -B antigens in a mutant TxB cell hybrid. EMBO J. 5, 943–949 (1986).
Amit, S. et al. Axin-mediated CKI phosphorylation of β-catenin at Ser45: a molecular switch for the Wnt pathway. Genes Dev. 16, 1066–1076 (2002).
Zhou, J., Dudley, M.E., Rosenberg, S.A. & Robbins, P.F. Persistence of multiple tumor-specific T-cell clones is associated with complete tumor regression in a melanoma patient receiving adoptive cell transfer therapy. J. Immunother. 28, 53–62 (2005).
Goshima, G., Mayer, M., Zhang, N., Stuurman, N. & Vale, R.D. Augmin: a protein complex required for centrosome-independent microtubule generation within the spindle. J. Cell Biol. 181, 421–429 (2008).
Rammensee, H., Bachmann, J., Emmerich, N.P., Bachor, O.A. & Stevanovic, S. SYFPEITHI: database for MHC ligands and peptide motifs. Immunogenetics 50, 213–219 (1999).
Morel, S. et al. Processing of some antigens by the standard proteasome but not by the immunoproteasome results in poor presentation by dendritic cells. Immunity 12, 107–117 (2000).
Chapiro, J. et al. Destructive cleavage of antigenic peptides either by the immunoproteasome or by the standard proteasome results in differential antigen presentation. J. Immunol. 176, 1053–1061 (2006).
Guillaume, B. et al. Two abundant proteasome subtypes that uniquely process some antigens presented by HLA class I molecules. Proc. Natl. Acad. Sci. USA 107, 18599–18604 (2010).
Kahle, J.J. et al. Comparison of an expanded ataxia interactome with patient medical records reveals a relationship between macular degeneration and ataxia. Hum. Mol. Genet. 20, 510–527 (2011).
Blazek, D. et al. The cyclin K/Cdk12 complex maintains genomic stability via regulation of expression of DNA damage response genes. Genes Dev. 25, 2158–2172 (2011).
Kawakami, Y. et al. Cloning of the gene coding for a shared human melanoma antigen recognized by autologous T cells infiltrating into tumor. Proc. Natl. Acad. Sci. USA 91, 3515–3519 (1994).
van der Bruggen, P. et al. A gene encoding an antigen recognized by cytolytic T lymphocytes on a human melanoma. Science 254, 1643–1647 (1991).
Boël, P. et al. BAGE: a new gene encoding an antigen recognized on human melanomas by cytolytic T lymphocytes. Immunity 2, 167–175 (1995).
Robbins, P.F. et al. A mutated β-catenin gene encodes a melanoma-specific antigen recognized by tumor infiltrating lymphocytes. J. Exp. Med. 183, 1185–1192 (1996).
Cox, A.L. et al. Identification of a peptide recognized by five melanoma-specific human cytotoxic T cell lines. Science 264, 716–719 (1994).
Pieper, R. et al. Biochemical identification of a mutated human melanoma antigen recognized by CD4+ T cells. J. Exp. Med. 189, 757–766 (1999).
van der Bruggen, P., Stroobant, V., Vigneron, N. & Van den Eynde, B. Peptide database: T cell–defined tumor antigens. Cancer Immun. 〈http://www.cancerimmunity.org/peptide/〉 (2013).
Matsushita, H. et al. Cancer exome analysis reveals a T-cell–dependent mechanism of cancer immunoediting. Nature 482, 400–404 (2012).
Castle, J.C. et al. Exploiting the mutanome for tumor vaccination. Cancer Res. 72, 1081–1091 (2012).
Kvistborg, P. et al. TIL therapy broadens the tumor-reactive CD8+ T cell compartment in melanoma patients. OncoImmunology 1, 409–418 (2012).
Jones, S. et al. Frequent mutations of chromatin remodeling gene ARID1A in ovarian clear cell carcinoma. Science 330, 228–231 (2010).
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007).
Berger, M.F. et al. Melanoma genome sequencing reveals frequent PREX2 mutations. Nature 485, 502–506 (2012).
Topalian, S.L., Muul, L.M., Solomon, D. & Rosenberg, S.A. Expansion of human tumor infiltrating lymphocytes for use in immunotherapy trials. J. Immunol. Methods 102, 127–141 (1987).
Dudley, M.E. et al. Adoptive cell transfer therapy following non-myeloablative but lymphodepleting chemotherapy for the treatment of patients with refractory metastatic melanoma. J. Clin. Oncol. 23, 2346–2357 (2005).
Arnold, D. et al. Proteasome subunits encoded in the MHC are not generally required for the processing of peptides bound by MHC class I molecules. Nature 360, 171–174 (1992).
Acknowledgements
We thank B. Van den Eynde (Ludwig Institute) for providing HEK293 cells transfected with immunoproteasomal subunits and S. Schwarz and R. Fisch for assisting with experiments.
Author information
Authors and Affiliations
Contributions
P.F.R. designed and developed the experimental screening system, analyzed data and drafted the manuscript. Y.-C.L. and M.E.-G. performed experiments evaluating TIL responses against candidate mutated peptides and analyzed results. Y.F.L. cloned and sequenced gene products encoding candidate epitopes identified by exomic sequencing and analyzed results. J.K.T., C.G., E.T., J.C.L. and P.C. carried out bioinformatic analyses. J.G. provided advice on exomic sequencing, prepared samples for sequencing and carried out validation studies using Sanger sequencing. Y.S. provided advice on sequencing of DNA isolated from tumor and normal cells and assisted with data analysis. S.A.R. helped design the studies and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–7 (PDF 213 kb)
Rights and permissions
About this article
Cite this article
Robbins, P., Lu, YC., El-Gamil, M. et al. Mining exomic sequencing data to identify mutated antigens recognized by adoptively transferred tumor-reactive T cells. Nat Med 19, 747–752 (2013). https://doi.org/10.1038/nm.3161
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3161
This article is cited by
-
MHC II immunogenicity shapes the neoepitope landscape in human tumors
Nature Genetics (2023)
-
Identification of antigenic epitopes recognized by tumor infiltrating lymphocytes in high grade serous ovarian cancer by multi-omics profiling of the auto-antigen repertoire
Cancer Immunology, Immunotherapy (2023)
-
Peptide-Based Therapeutics in Cancer Therapy
Molecular Biotechnology (2023)
-
Using patient-derived tumor organoids from common epithelial cancers to analyze personalized T-cell responses to neoantigens
Cancer Immunology, Immunotherapy (2023)
-
Neoantigen-targeted CD8+ T cell responses with PD-1 blockade therapy
Nature (2023)