Abstract
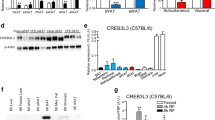
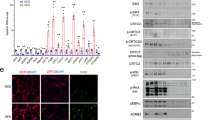
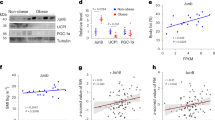
Obesity develops as a result of altered energy homeostasis favoring fat storage. Here we describe a new transcription co-regulator for adiposity and energy metabolism, SERTA domain containing 2 (TRIP-Br2, also called SERTAD2). TRIP-Br2–null mice are resistant to obesity and obesity-related insulin resistance. Adipocytes of these knockout mice showed greater stimulated lipolysis secondary to enhanced expression of hormone sensitive lipase (HSL) and β3-adrenergic (Adrb3) receptors. The knockout mice also have higher energy expenditure because of increased adipocyte thermogenesis and oxidative metabolism caused by upregulating key enzymes in their respective processes. Our data show that a cell-cycle transcriptional co-regulator, TRIP-Br2, modulates fat storage through simultaneous regulation of lipolysis, thermogenesis and oxidative metabolism. These data, together with the observation that TRIP-Br2 expression is selectively elevated in visceral fat in obese humans, suggests that this transcriptional co-regulator is a new therapeutic target for counteracting the development of obesity, insulin resistance and hyperlipidemia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kahn, B.B. & Flier, J.S. Obesity and insulin resistance. J. Clin. Invest. 106, 473–481 (2000).
Friedman, J.M. A war on obesity, not the obese. Science 299, 856–858 (2003).
Rosen, E.D. & Spiegelman, B.M. Adipocytes as regulators of energy balance and glucose homeostasis. Nature 444, 847–853 (2006).
Duncan, R.E., Ahmadian, M., Jaworski, K., Sarkadi-Nagy, E. & Sul, H.S. Regulation of lipolysis in adipocytes. Annu. Rev. Nutr. 27, 79–101 (2007).
Lafontan, M. & Langin, D. Lipolysis and lipid mobilization in human adipose tissue. Prog. Lipid Res. 48, 275–297 (2009).
Hamann, A., Flier, J.S. & Lowell, B.B. Decreased brown fat markedly enhances susceptibility to diet-induced obesity, diabetes, and hyperlipidemia. Endocrinology 137, 21–29 (1996).
Desvergne, B., Michalik, L. & Wahli, W. Transcriptional regulation of metabolism. Physiol. Rev. 86, 465–514 (2006).
Feige, J.N. & Auwerx, J. Transcriptional coregulators in the control of energy homeostasis. Trends Cell Biol. 17, 292–301 (2007).
Rosenfeld, M.G., Lunyak, V.V. & Glass, C.K. Sensors and signals: a coactivator/corepressor/epigenetic code for integrating signal-dependent programs of transcriptional response. Genes Dev. 20, 1405–1428 (2006).
Spiegelman, B.M. & Heinrich, R. Biological control through regulated transcriptional coactivators. Cell 119, 157–167 (2004).
Lehrke, M. & Lazar, M.A. The many faces of PPARγ. Cell 123, 993–999 (2005).
Hsu, S.I. et al. TRIP-Br: a novel family of PHD zinc finger- and bromodomain-interacting proteins that regulate the transcriptional activity of E2F–1/DP-1. EMBO J. 20, 2273–2285 (2001).
Cheong, J.K. et al. TRIP-Br2 promotes oncogenesis in nude mice and is frequently overexpressed in multiple human tumors. J. Transl. Med. 7, 8 (2009).
Darwish, H., Cho, J.M., Loignon, M. & Alaoui-Jamali, M.A. Overexpression of SERTAD3, a putative oncogene located within the 19q13 amplicon, induces E2F activity and promotes tumor growth. Oncogene 26, 4319–4328 (2007).
Hayashi, R., Goto, Y., Ikeda, R., Yokoyama, K.K. & Yoshida, K. CDCA4 is an E2F transcription factor family–induced nuclear factor that regulates E2F-dependent transcriptional activation and cell proliferation. J. Biol. Chem. 281, 35633–35648 (2006).
Lai, I.L., Wang, S.Y., Yao, Y.L. & Yang, W.M. Transcriptional and subcellular regulation of the TRIP-Br family. Gene 388, 102–109 (2007).
Hirose, T. et al. Regulation of CREB-mediated transcription by association of CDK4 binding protein p34SEI-1 with CBP. Int. J. Mol. Med. 2003 11, 705–712 (2003).
Kim, S.S., Chen, Y.M., O'Leary, E., Witzgall, R., Vidal, M. & Bonventre, J.V. A novel member of the RING finger family, KRIP-1, associates with the KRAB-A transcriptional repressor domain of zinc finger proteins. Proc. Natl. Acad. Sci. USA. 93, 15299–15304 (1996).
Fajas, L. et al. E2Fs regulate adipocyte differentiation. Dev. Cell 3, 39–49 (2002).
Tseng, Y.H. et al. Prediction of preadipocyte differentiation by gene expression reveals role of insulin receptor substrates and necdin. Nat. Cell Biol. 7, 601–611 (2005).
Sim, K.G., Cheong, J.K. & Hsu, S.I. The TRIP-Br family of transcriptional regulators is essential for the execution of cyclin E–mediated cell cycle progression. Cell Cycle 5, 1111–1115 (2006).
Almind, K. & Kahn, C.R. Genetic determinants of energy expenditure and insulin resistance in diet-induced obesity in mice. Diabetes 53, 3274–3285 (2004).
Jandacek, R.J., Heubi, J.E. & Tso, P. A novel, noninvasive method for the measurement of intestinal fat absorption. Gastroenterology 127, 139–144 (2004).
Hotta, K. et al. Plasma concentrations of a novel, adipose-specific protein, adiponectin, in type 2 diabetic patients. Arterioscler. Thromb. Vasc. Biol. 20, 1595–1599 (2000).
Rosenbaum, M. & Leibel, R.L. The role of leptin in human physiology. N. Engl. J. Med. 341, 913–915 (1999).
Postic, C. & Girard, J. Contribution of de novo fatty acid synthesis to hepatic steatosis and insulin resistance: lessons from genetically engineered mice. J. Clin. Invest. 118, 829–838 (2008).
Hotamisligil, G.S. Inflammation and metabolic disorders. Nature 444, 860–867 (2006).
Schenk, S., Saberi, M. & Olefsky, J.M. Insulin sensitivity: modulation by nutrients and inflammation. J. Clin. Invest. 118, 2992–3002 (2008).
Gregoire, F.M., Smas, C.M. & Sul, H.S. Understanding adipocyte differentiation. Physiol. Rev. 78, 783–809 (1998).
Rosen, E.D. & MacDougald, O.A. Adipocyte differentiation from the inside out. Nat. Rev. Mol. Cell Biol. 7, 885–896 (2006).
Kersten, S. Mechanisms of nutritional and hormonal regulation of lipogenesis. EMBO Rep. 2, 282–286 (2001).
Uysal, K.T., Scheja, L., Wiesbrock, S.M., Bonner-Weir, S. & Hotamisligil, G.S. Improved glucose and lipid metabolism in genetically obese mice lacking aP2. Endocrinology 141, 3388–3396 (2000).
Turner, S.M. et al. Measurement of TG synthesis and turnover in vivo by 2H2O incorporation into the glycerol moiety and application of MIDA. Am. J. Physiol. Endocrinol. Metab. 285, E790–E803 (2003).
Ahmadian, M. et al. Adipose overexpression of desnutrin promotes fatty acid use and attenuates diet-induced obesity. Diabetes 58, 855–866 (2009).
Zimmermann, R. et al. Fat mobilization in adipose tissue is promoted by adipose triglyceride lipase. Science 306, 1383–1386 (2004).
Holm, C., Osterlund, T., Laurell, H. & Contreras, J.A. Molecular mechanisms regulating hormone-sensitive lipase and lipolysis. Annu. Rev. Nutr. 20, 365–393 (2000).
Mottillo, E.P., Shen, X.J. & Granneman, J.G. Role of hormone-sensitive lipase in β-adrenergic remodeling of white adipose tissue. Am. J. Physiol. Endocrinol. Metab. 293, E1188–E1197 (2007).
Langin, D. et al. Adipocyte lipases and defect of lipolysis in human obesity. Diabetes 54, 3190–3197 (2005).
Lowell, B.B. & Bachman, E.S. β-adrenergic receptors, diet-induced thermogenesis, and obesity. J. Biol. Chem. 278, 29385–29388 (2003).
Cermak, T. et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 39, e82 (2011).
Lonard, D.M., Lanz, R.B. & O'Malley, B.W. Nuclear receptor coregulators and human disease. Endocr. Rev. 28, 575–587 (2007).
Jaworski, K. et al. AdPLA ablation increases lipolysis and prevents obesity induced by high-fat feeding or leptin deficiency. Nat. Med. 15, 159–168 (2009).
Lucas, S., Tavernier, G., Tiraby, C., Mairal, A. & Langin, D. Expression of human hormone-sensitive lipase in white adipose tissue of transgenic mice increases lipase activity but does not enhance in vitro lipolysis. J. Lipid Res. 44, 154–163 (2003).
Kopecky, J. et al. Reduction of dietary obesity in aP2-Ucp transgenic mice: mechanism and adipose tissue morphology. Am. J. Physiol. 270, E776–E786 (1996).
Wang, Y.X. et al. Peroxisome-proliferator–activated receptor δ activates fat metabolism to prevent obesity. Cell 113, 159–170 (2003).
Puigserver, P. et al. A cold-inducible coactivator of nuclear receptors linked to adaptive thermogenesis. Cell 92, 829–839 (1998).
Leone, T.C. et al. PGC-1α deficiency causes multi-system energy metabolic derangements: muscle dysfunction, abnormal weight control and hepatic steatosis. PLoS Biol. 3, e101 (2005).
Kosteli, A. et al. Weight loss and lipolysis promote a dynamic immune response in murine adipose tissue. J. Clin. Invest. 120, 3466–3479 (2010).
Blanchet, E., Annicotte, J.S. & Fajas, L. Cell cycle regulators in the control of metabolism. Cell Cycle 8, 4029–4031 (2009).
Berndt, J. et al. Plasma visfatin concentrations and fat depot-specific mRNA expression in humans. Diabetes 54, 2911–2916 (2005).
Blüher, M., Unger, R., Rassoul, F., Richter, V. & Paschke, R. Relation between glycaemic control, hyperinsulinaemia and plasma concentrations of soluble adhesion molecules in patients with impaired glucose tolerance or Type II diabetes. Diabetologia 45, 210–216 (2002).
Carpenter, A.E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Chiang, S.H. et al. The protein kinase IKKɛ regulates energy balance in obese mice. Cell 138, 961–975 (2009).
Ahmadian, M. et al. Desnutrin/ATGL is regulated by AMPK and is required for a brown adipose phenotype. Cell Metab. 13, 739–748 (2011).
Yehuda-Shnaidman, E., Buehrer, B., Pi, J., Kumar, N. & Collins, S. Acute stimulation of white adipocyte respiration by PKA-induced lipolysis. Diabetes 59, 2474–2483 (2010).
Wu, M. et al. Multiparameter metabolic analysis reveals a close link between attenuated mitochondrial bioenergetic function and enhanced glycolysis dependency in human tumor cells. Am. J. Physiol. Cell Physiol. 292, C125–C136 (2007).
Boucher, J. et al. Human α2A-adrenergic receptor gene expressed in transgenic mouse adipose tissue under the control of its regulatory elements. J. Mol. Endocrinol. 29, 251–264 (2002).
Macotela, Y., Boucher, J., Tran, T.T. & Kahn, C.R. Sex and depot differences in adipocyte insulin sensitivity and glucose metabolism. Diabetes 58, 803–812 (2009).
Folch, J., Lees, M. & Sloane Stanley, G.H. A simple method for the isolation and purification of total lipides from animal tissues. J. Biol. Chem. 226, 497–509 (1957).
Morgenstern, J.P. & Land, H. Advanced mammalian gene transfer: high titre retroviral vectors with multiple drug selection markers and a complementary helper-free packaging cell line. Nucleic Acids Res. 18, 3587–3596 (1990).
Kulkarni, R.N. et al. Tissue-specific knockout of the insulin receptor in pancreatic beta cells creates an insulin secretory defect similar to that in type 2 diabetes. Cell 96, 329–339 (1999).
Lu, Y.C. et al. Cyclophilin a protects Peg3 from hypermethylation and inactive histone modification. J. Biol. Chem. 281, 39081–39087 (2006).
Acknowledgements
The authors thank C.R. Kahn (Joslin Diabetes Center) for providing reagents and discussions, E. Rosen for discussions, E. Morgan and K. Parlee for excellent assistance in the preparation of this manuscript, H. Li for assistance with hormone assays, O.P. McGuinness for mouse metabolic phenotyping, L. Fajas (INSERM) for providing reagents, M. Mori (Joslin Diabetes Center) for providing samples and R. Zechner for providing the protocol for triglyceride hydrolase activities. Funds to generate some reagents used in this research were supported by US National Institutes of Health (NIH) grant RO1 DK 67536 (R.N.K.) and the Joslin Graetz Bridge Funds (R.N.K.), NIH grants R01 HL073168 (A.D.), K99 DK090210 (C.W.L.), DK51586 and DK58825 (S.R.F.) and the Joslin Diabetes and Endocrinology Research Center Specialized Assay and Advanced Microscopy Cores (NIH P30 DK36836). The human studies were supported by a grant of the Deutsche Forschungsgemeinschaft Clinical Research group 'Atherobesity' (KFO152; BL 833/1-1). C.W.L. was supported by a US National Institutes of Health Interdisciplinary training grant (1RL9EB008539-01) (SysCODE), K99 DK090210 and R00 DK090210. D.K. is the recipient of a research fellowship (Manpei Suzuki Diabetes Foundation, Japan) and a Juvenile Diabetes Research Foundation postdoctoral Fellowship. S.I.-H.H. and J.K.C. were supported by the R. Glenn Davis (Dialysis Center, Inc.) Endowed Professorship in Clinical and Translational Medicine and by the University of Florida, Division of Nephrology Gatorade Fund. S.I.-H.H. was supported as a Scholar of the Clinical Translational Science Institute at the University of Florida.
Author information
Authors and Affiliations
Contributions
C.W.L., J.K.C., S.I.-H.H. and R.N.K. conceived the project. C.W.L. and R.N.K. designed the experiments. A.D. analyzed human data. C.W.L., J.B., J.K.C., C.V., D.K., J.H., C.M., M.K.H., K.T. and H.-J.K. performed experiments and analyzed data. M.B. contributed human samples and supervised human expression analysis. S.R.F. supervised experiments. D.L., S.I.-H.H., Y.-H.T. and L.G. contributed reagents. C.W.L. and R.N.K. wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–26, Supplementary Tables 1 and 2 and Supplementary Methods (PDF 644 kb)
Rights and permissions
About this article
Cite this article
Liew, C., Boucher, J., Cheong, J. et al. Ablation of TRIP-Br2, a regulator of fat lipolysis, thermogenesis and oxidative metabolism, prevents diet-induced obesity and insulin resistance. Nat Med 19, 217–226 (2013). https://doi.org/10.1038/nm.3056
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3056
This article is cited by
-
Adipose expression of CREB3L3 modulates body weight during obesity
Scientific Reports (2021)
-
Transcriptome analyses of liver in newly-hatched chicks during the metabolic perturbation of fasting and re-feeding reveals THRSPA as the key lipogenic transcription factor
BMC Genomics (2020)
-
Transcriptional profiling of liver during the critical embryo-to-hatchling transition period in the chicken (Gallus gallus)
BMC Genomics (2018)
-
miR-128-3p regulates 3T3-L1 adipogenesis and lipolysis by targeting Pparg and Sertad2
Journal of Physiology and Biochemistry (2018)
-
Transcription regulator TRIP-Br2 mediates ER stress-induced brown adipocytes dysfunction
Scientific Reports (2017)