Abstract
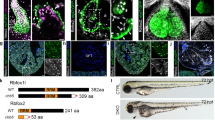
Left ventricular noncompaction (LVNC) causes prominent ventricular trabeculations and reduces cardiac systolic function. The clinical presentation of LVNC ranges from asymptomatic to heart failure. We show that germline mutations in human MIB1 (mindbomb homolog 1), which encodes an E3 ubiquitin ligase that promotes endocytosis of the NOTCH ligands DELTA and JAGGED, cause LVNC in autosomal-dominant pedigrees, with affected individuals showing reduced NOTCH1 activity and reduced expression of target genes. Functional studies in cells and zebrafish embryos and in silico modeling indicate that MIB1 functions as a dimer, which is disrupted by the human mutations. Targeted inactivation of Mib1 in mouse myocardium causes LVNC, a phenotype mimicked by inactivation of myocardial Jagged1 or endocardial Notch1. Myocardial Mib1 mutants show reduced ventricular Notch1 activity, expansion of compact myocardium to proliferative, immature trabeculae and abnormal expression of cardiac development and disease genes. These results implicate NOTCH signaling in LVNC and indicate that MIB1 mutations arrest chamber myocardium development, preventing trabecular maturation and compaction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ritter, M. et al. Isolated noncompaction of the myocardium in adults. Mayo Clin. Proc. 72, 26–31 (1997).
Sedmera, D., Pexieder, T., Vuillemin, M., Thompson, R.P. & Anderson, R.H. Developmental patterning of the myocardium. Anat. Rec. 258, 319–337 (2000).
Towbin, J.A. Left ventricular noncompaction: a new form of heart failure. Heart Fail. Clin. 6, 453–469, viii (2010).
Captur, G. & Nihoyannopoulos, P. Left ventricular non-compaction: genetic heterogeneity, diagnosis and clinical course. Int. J. Cardiol. 140, 145–153 (2010).
Oechslin, E.N., Attenhofer Jost, C.H., Rojas, J.R., Kaufmann, P.A. & Jenni, R. Long-term follow-up of 34 adults with isolated left ventricular noncompaction: a distinct cardiomyopathy with poor prognosis. J. Am. Coll. Cardiol. 36, 493–500 (2000).
Stöllberger, C. & Finsterer, J. Left ventricular hypertrabeculation/noncompaction. J. Am. Soc. Echocardiogr. 17, 91–100 (2004).
Maron, B.J. et al. Contemporary definitions and classification of the cardiomyopathies: an American Heart Association Scientific Statement from the Council on Clinical Cardiology, Heart Failure and Transplantation Committee; Quality of Care and Outcomes Research and Functional Genomics and Translational Biology Interdisciplinary Working Groups; and Council on Epidemiology and Prevention. Circulation 113, 1807–1816 (2006).
Klaassen, S. et al. Mutations in sarcomere protein genes in left ventricular noncompaction. Circulation 117, 2893–2901 (2008).
Ichida, F. et al. Novel gene mutations in patients with left ventricular noncompaction or Barth syndrome. Circulation 103, 1256–1263 (2001).
Hermida-Prieto, M. et al. Familial dilated cardiomyopathy and isolated left ventricular noncompaction associated with lamin A/C gene mutations. Am. J. Cardiol. 94, 50–54 (2004).
Shou, W. et al. Cardiac defects and altered ryanodine receptor function in mice lacking FKBP12. Nature 391, 489–492 (1998).
Finsterer, J. Cardiogenetics, neurogenetics, and pathogenetics of left ventricular hypertrabeculation/noncompaction. Pediatr. Cardiol. 30, 659–681 (2009).
Bione, S. et al. A novel X-linked gene, G4.5. is responsible for Barth syndrome. Nat. Genet. 12, 385–389 (1996).
Artavanis-Tsakonas, S., Rand, M.D. & Lake, R.J. Notch signaling: cell fate control and signal integration in development. Science 284, 770–776 (1999).
High, F.A. & Epstein, J.A. The multifaceted role of Notch in cardiac development and disease. Nat. Rev. Genet. 9, 49–61 (2008).
de la Pompa, J.L. & Epstein, J.A. Coordinating tissue interactions: notch signaling in cardiac development and disease. Dev. Cell 22, 244–254 (2012).
Itoh, M. et al. Mind bomb is a ubiquitin ligase that is essential for efficient activation of Notch signaling by Delta. Dev. Cell 4, 67–82 (2003).
Barsi, J.C., Rajendra, R., Wu, J.I. & Artzt, K. Mind bomb1 is a ubiquitin ligase essential for mouse embryonic development and Notch signaling. Mech. Dev. 122, 1106–1117 (2005).
Hansson, E.M. et al. Control of Notch-ligand endocytosis by ligand-receptor interaction. J. Cell Sci. 123, 2931–2942 (2010).
Kopan, R. & Ilagan, M.X. The canonical Notch signaling pathway: unfolding the activation mechanism. Cell 137, 216–233 (2009).
Koo, B.K. et al. An obligatory role of mind bomb-1 in notch signaling of mammalian development. PLoS ONE 2, e1221 (2007).
Stanley, E.G. et al. Efficient Cre-mediated deletion in cardiac progenitor cells conferred by a 3′UTR-ires-Cre allele of the homeobox gene Nkx2–5. Int. J. Dev. Biol. 46, 431–439 (2002).
Jiao, K. et al. An essential role of Bmp4 in the atrioventricular septation of the mouse heart. Genes Dev. 17, 2362–2367 (2003).
Jenni, R., Oechslin, E., Schneider, J., Attenhofer Jost, C. & Kaufmann, P.A. Echocardiographic and pathoanatomical characteristics of isolated left ventricular non-compaction: a step towards classification as a distinct cardiomyopathy. Heart 86, 666–671 (2001).
Stöllberger, C. & Finsterer, J. Trabeculation and left ventricular hypertrabeculation/noncompaction. J. Am. Soc. Echocardiogr. 17, 1120–1121 (2004).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Davydov, E.V. et al. Identifying a high fraction of the human genome to be under selective constraint using GERP. PLoS Comput. Biol. 6, e1001025 (2010).
Ichida, F. Left ventricular noncompaction. Circ. J. 73, 19–26 (2009).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
MacArthur, D.G. et al. A systematic survey of loss-of-function variants in human protein-coding genes. Science 335, 823–828 (2012).
Frischmeyer, P.A. et al. An mRNA surveillance mechanism that eliminates transcripts lacking termination codons. Science 295, 2258–2261 (2002).
Macías-Vidal, J., Gort, L., Lluch, M., Pineda, M. & Coll, M.J. Nonsense-mediated mRNA decay process in nine alleles of Niemann-Pick type C patients from Spain. Mol. Genet. Metab. 97, 60–64 (2009).
Van de Walle, I. et al. An early decrease in Notch activation is required for human TCR-αβ lineage differentiation at the expense of TCR-γδ T cells. Blood 113, 2988–2998 (2009).
Zhang, C., Li, Q., Lim, C.H., Qiu, X. & Jiang, Y.J. The characterization of zebrafish antimorphic mib alleles reveals that Mib and Mind bomb-2 (Mib2) function redundantly. Dev. Biol. 305, 14–27 (2007).
Sedgwick, S.G. & Smerdon, S.J. The ankyrin repeat: a diversity of interactions on a common structural framework. Trends Biochem. Sci. 24, 311–316 (1999).
Rodighiero, S. et al. Fixation, mounting and sealing with nail polish of cell specimens lead to incorrect FRET measurements using acceptor photobleaching. Cell. Physiol. Biochem. 21, 489–498 (2008).
Chapman, G., Liu, L., Sahlgren, C., Dahlqvist, C. & Lendahl, U. High levels of Notch signaling down-regulate Numb and Numblike. J. Cell Biol. 175, 535–540 (2006).
Del Monte, G., Grego-Bessa, J., Gonzalez-Rajal, A., Bolos, V. & De La Pompa, J.L. Monitoring Notch1 activity in development: evidence for a feedback regulatory loop. Dev. Dyn. 236, 2594–2614 (2007).
Parsons, M.J. et al. Notch-responsive cells initiate the secondary transition in larval zebrafish pancreas. Mech. Dev. 126, 898–912 (2009).
McKenzie, G.J. et al. Nuclear Ca2+ and CaM kinase IV specify hormonal- and Notch-responsiveness. J. Neurochem. 93, 171–185 (2005).
Koibuchi, N. & Chin, M.T. CHF1/Hey2 plays a pivotal role in left ventricular maturation through suppression of ectopic atrial gene expression. Circ. Res. 100, 850–855 (2007).
Singh, M.K. et al. Tbx20 is essential for cardiac chamber differentiation and repression of Tbx2. Development 132, 2697–2707 (2005).
Moens, C.B., Stanton, B.R., Parada, L.F. & Rossant, J. Defects in heart and lung development in compound heterozygotes for two different targeted mutations at the N-myc locus. Development 119, 485–499 (1993).
Zeller, R., Bloch, K.D., Williams, B.S., Arceci, R.J. & Seidman, C.E. Localized expression of the atrial natriuretic factor gene during cardiac embryogenesis. Genes Dev. 1, 693–698 (1987).
Chen, H. et al. BMP10 is essential for maintaining cardiac growth during murine cardiogenesis. Development 131, 2219–2231 (2004).
Van Kempen, M.J. et al. Developmental changes of connexin40 and connexin43 mRNA distribution patterns in the rat heart. Cardiovasc. Res. 32, 886–900 (1996).
Zhang, C., Li, Q. & Jiang, Y.J. Zebrafish Mib and Mib2 are mutual E3 ubiquitin ligases with common and specific δ substrates. J. Mol. Biol. 366, 1115–1128 (2007).
Grego-Bessa, J. et al. Notch signaling is essential for ventricular chamber development. Dev. Cell 12, 415–429 (2007).
MacGrogan, D., Nus, M. & de la Pompa, J.L. Notch signaling in cardiac development and disease. Curr. Top. Dev. Biol. 92, 333–365 (2010).
Gridley, T. Notch signaling in the vasculature. Curr. Top. Dev. Biol. 92, 277–309 (2010).
Sarma, R.J., Chana, A. & Elkayam, U. Left ventricular noncompaction. Prog. Cardiovasc. Dis. 52, 264–273 (2010).
Wang, Y. et al. Ephrin-B2 controls VEGF-induced angiogenesis and lymphangiogenesis. Nature 465, 483–486 (2010).
Mancini, S.J. et al. Jagged1-dependent Notch signaling is dispensable for hematopoietic stem cell self-renewal and differentiation. Blood 105, 2340–2342 (2005).
Radtke, F. et al. Deficient T cell fate specification in mice with an induced inactivation of Notch1. Immunity 10, 547–558 (1999).
Soriano, P. Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat. Genet. 21, 70–71 (1999).
Kanzler, B., Kuschert, S.J., Liu, Y.H. & Mallo, M. Hoxa-2 restricts the chondrogenic domain and inhibits bone formation during development of the branchial area. Development 125, 2587–2597 (1998).
del Monte, G. et al. Differential Notch signaling in the epicardium is required for cardiac inflow development and coronary vessel morphogenesis. Circ. Res. 108, 824–836 (2011).
Li, B. & Dewey, C.N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Jin, Y., Blue, E.K., Dixon, S., Shao, Z. & Gallagher, P.J. A death-associated protein kinase (DAPK)-interacting protein, DIP-1, is an E3 ubiquitin ligase that promotes tumor necrosis factor-induced apoptosis and regulates the cellular levels of DAPK. J. Biol. Chem. 277, 46980–46986 (2002).
Cruz-Adalia, A. et al. CD69 limits the severity of cardiomyopathy after autoimmune myocarditis. Circulation 122, 1396–1404 (2010).
Jacquier, A. et al. Measurement of trabeculated left ventricular mass using cardiac magnetic resonance imaging in the diagnosis of left ventricular non-compaction. Eur. Heart J. 31, 1098–1104 (2010).
Petersen, S.E. et al. Left ventricular non-compaction: insights from cardiovascular magnetic resonance imaging. J. Am. Coll. Cardiol. 46, 101–105 (2005).
Stöllberger, C., Finsterer, J. & Blazek, G. Left ventricular hypertrabeculation/noncompaction and association with additional cardiac abnormalities and neuromuscular disorders. Am. J. Cardiol. 90, 899–902 (2002).
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat. Protoc. 5, 725–738 (2010).
Roy, A., Xu, D., Poisson, J. & Zhang, Y. A protocol for computer-based protein structure and function prediction. J. Vis. Exp. e3259 (2011).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40 (2008).
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201 (2006).
Kiefer, F., Arnold, K., Kunzli, M., Bordoli, L. & Schwede, T. The SWISS-MODEL Repository and associated resources. Nucleic Acids Res. 37, D387–D392 (2009).
Peitsch, M.C. Protein modeling by e-mail. Nat. Biotechnol. 13, 658–660 (1995).
Jones, D.T., Taylor, W.R. & Thornton, J.M. A model recognition approach to the prediction of all-helical membrane protein structure and topology. Biochemistry 33, 3038–3049 (1994).
Ritchie, D.W. & Venkatraman, V. Ultra-fast FFT protein docking on graphics processors. Bioinformatics 26, 2398–2405 (2010).
Macindoe, G., Mavridis, L., Venkatraman, V., Devignes, M.D. & Ritchie, D.W. HexServer: an FFT-based protein docking server powered by graphics processors. Nucleic Acids Res. 38, W445–9 (2010).
Ritchie, D.W., Kozakov, D. & Vajda, S. Accelerating and focusing protein-protein docking correlations using multi-dimensional rotational FFT generating functions. Bioinformatics 24, 1865–1873 (2008).
Ritchie, D.W. High-order analytic translation matrix elements for real-space six-dimensional polar Fourier correlations. J. Appl. Cryst. 38, 808–818 (2005).
Mustard, D. & Ritchie, D.W. Docking essential dynamics eigenstructures. Proteins 60, 269–274 (2005).
Ritchie, D.W. Evaluation of protein docking predictions using Hex 3.1 in CAPRI rounds 1 and 2. Proteins 52, 98–106 (2003).
Ritchie, D.W. & Kemp, G.J.L. Protein docking using spherical polar Fourier correlations. Proteins 39, 178–194 (2000).
Ritchie, D.W. & Kemp, G.J.L. Fast computation, rotation, and comparison of low resolution spherical harmonic molecular surfaces. J. Comput. Chem. 20, 383–395 (1999).
Ritchie, D.W. Parametric Protein Shape Recognition. PhD thesis, Univ. Aberdeen (1998).
Kozakov, D. et al. Achieving reliability and high accuracy in automated protein docking: ClusPro, PIPER, SDU, and stability analysis in CAPRI rounds 13–19. Proteins 78, 3124–3130 (2010).
Kozakov, D., Brenke, R., Comeau, S.R. & Vajda, S. PIPER: an FFT-based protein docking program with pairwise potentials. Proteins 65, 392–406 (2006).
Comeau, S.R., Gatchell, D.W., Vajda, S. & Camacho, C.J. ClusPro: an automated docking and discrimination method for the prediction of protein complexes. Bioinformatics 20, 45–50 (2004).
Copeland, J.W. & Treisman, R. The diaphanous-related formin mDia1 controls serum response factor activity through its effects on actin polymerization. Mol. Biol. Cell 13, 4088–4099 (2002).
Acknowledgements
We thank P.J. Gallagher (Indiana University) for the Mib1-specific antibody, U. Lendahl (Karolinska Institute) for the Jag1- and Notch1-HEK293 cells, A. Martín-Pendas (Cancer Center, CSIC, Salamanca) for the pCELF plamid, D. Saura and J. González for carrying out the CMRI of the patients, M. Manzanares, A. Martín-Pendas, J.M. Pérez-Pomares, J.M. Redondo, J.J. Schott and M. Torres for critical reading of the manuscript and S. Bartlett (CNIC) for English editing. This study was funded by grants SAF2010-17555, RD06/0014/0038 (RECAVA) and RD06/0010/1013 (TERCEL) from the Spanish Ministry of Economy and Competition (MINECO) to J.L.d.l.P. G.L. has a PhD fellowship from the MINECO (FPI Program, BES-2008-002904). The CNIC is supported by the MINECO and the Pro-CNIC Foundation.
Author information
Authors and Affiliations
Contributions
G.L., J.C.C., B.M.-P., D.M., A.G.-R., B.P., G.D. and D.D. performed experiments. C.T. analyzed the sequencing data, F.M. performed in silico modeling, J.L.I.-G. performed CMRI in mice, and Y.-Y.K. provided the Mib1 conditional mutant. G.P., M.S.-M., B.I., C.M., P.G.-P., J.R.G. and L.M. provided samples, and diagnosed and evaluated patients with LVNC. L.F.-F. and L.J.J.-B. performed and evaluated echocardiography in mice and performed CMRI on the parents of the probands. J.L.d.l.P. designed experiments, reviewed the data and wrote the manuscript. All authors reviewed the manuscript during its preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Tables 1 and 2 (PDF 1730 kb)
Supplementary Video 1
6-month-old wild-type (WT) echocardiography analysis. Ep, epicardium; en, endocardium. (AVI 740 kb)
Supplementary Video 2
6-month-old Mib1flox;cTnT-Cre echocardiography analysis. Ep, epicardium. Asterisks mark where prominent trabeculae can be observed. (AVI 755 kb)
Supplementary Video 3
6-month-old WT CMRI. 4 chambers view. (AVI 199 kb)
Supplementary Video 4
6-month-old WT CMRI. short axis view. (AVI 176 kb)
Supplementary Video 5
6-month-old Mib1flox;cTnT-Cre CMRI 4 chambers view. (AVI 220 kb)
Supplementary Video 6
6-month-old Mib1flox;cTnT-Cre CMRI short axis view. (AVI 66 kb)
Supplementary Video 7
Healthy control CMRI 4 chamber view (AVI 630 kb)
Supplementary Video 8
Patient 1.II.3 CMRI 4 chamber view. (AVI 542 kb)
Supplementary Video 9
Patient 2.I.2 CMRI 4 chamber view. (AVI 735 kb)
Supplementary Video 10
Healthy control CMRI 2 chamber view. (AVI 555 kb)
Supplementary Video 11
Patient 1.II.3 CMRI 2 chamber view. (AVI 611 kb)
Supplementary Video 12
Patient 2.I.2 CMRI 2 chamber view. (AVI 388 kb)
Supplementary Video 13
Healthy control CMRI short axis view. (AVI 370 kb)
Supplementary Video 14
Patient 1.II.3 CMRI short axis view. (AVI 448 kb)
Supplementary Video 15
Patient 2.I.2 CMRI short axis view. (AVI 499 kb)
Supplementary Table 3
Ventricular alterations and non-compaction index in LVNC affected family members and control individuals based on echocardiographic and CMRI data. (XLS 39 kb)
Supplementary Data Set 1
E14.5 WT vs. Mib1flox;cTnT-Cre. RNA-Seq data, all genes (XLSX 140 kb)
Supplementary Data Set 2
E14.5 WT vs. Mib1flox;cTnT-Cre. RNA-Seq data, only up-regulated and down-regulated genes (XLSX 2123 kb)
Rights and permissions
About this article
Cite this article
Luxán, G., Casanova, J., Martínez-Poveda, B. et al. Mutations in the NOTCH pathway regulator MIB1 cause left ventricular noncompaction cardiomyopathy. Nat Med 19, 193–201 (2013). https://doi.org/10.1038/nm.3046
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3046
This article is cited by
-
Left ventricular hypertrophy and metabolic resetting in the Notch3-deficient adult mouse heart
Scientific Reports (2023)
-
Endothelial deletion of PTBP1 disrupts ventricular chamber development
Nature Communications (2023)
-
Notch signaling pathway: architecture, disease, and therapeutics
Signal Transduction and Targeted Therapy (2022)
-
Overlap phenotypes of the left ventricular noncompaction and hypertrophic cardiomyopathy with complex arrhythmias and heart failure induced by the novel truncated DSC2 mutation
Orphanet Journal of Rare Diseases (2021)
-
Oestrogen Receptor β Activation Protects Against Myocardial Infarction via Notch1 Signalling
Cardiovascular Drugs and Therapy (2020)