Abstract
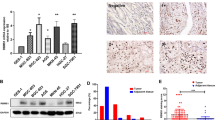
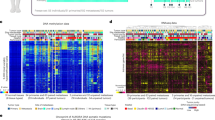
There is a pressing need to identify prognostic markers of metastatic disease and targets for treatment. Combining high-throughput RNA sequencing, functional characterization, mechanistic studies and clinical validation, we identify leukemia inhibitory factor receptor (LIFR) as a breast cancer metastasis suppressor downstream of the microRNA miR-9 and upstream of Hippo signaling. Restoring LIFR expression in highly malignant tumor cells suppresses metastasis by triggering a Hippo kinase cascade that leads to phosphorylation, cytoplasmic retention and functional inactivation of the transcriptional coactivator YES-associated protein (YAP). Conversely, loss of LIFR in nonmetastatic breast cancer cells induces migration, invasion and metastatic colonization through activation of YAP. LIFR is downregulated in human breast carcinomas and inversely correlates with metastasis. Notably, in approximately 1,000 nonmetastatic breast tumors, LIFR expression status correlated with metastasis-free, recurrence-free and overall survival outcomes in the patients. These findings identify LIFR as a metastasis suppressor that functions through the Hippo-YAP pathway and has significant prognostic power.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Lee, Y.T. Breast carcinoma: pattern of metastasis at autopsy. J. Surg. Oncol. 23, 175–180 (1983).
Weigelt, B., Peterse, J.L. & van 't Veer, L.J. Breast cancer metastasis: markers and models. Nat. Rev. Cancer 5, 591–602 (2005).
Steeg, P.S. Tumor metastasis: mechanistic insights and clinical challenges. Nat. Med. 12, 895–904 (2006).
Steeg, P.S. & Theodorescu, D. Metastasis: a therapeutic target for cancer. Nat. Clin. Pract. Oncol. 5, 206–219 (2008).
Vaidya, K.S. & Welch, D.R. Metastasis suppressors and their roles in breast carcinoma. J. Mammary Gland Biol. Neoplasia 12, 175–190 (2007).
Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
He, L. & Hannon, G.J. MicroRNAs: small RNAs with a big role in gene regulation. Nat. Rev. Genet. 5, 522–531 (2004).
Bartel, D.P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Nicoloso, M.S., Spizzo, R., Shimizu, M., Rossi, S. & Calin, G.A. MicroRNAs—the micro steering wheel of tumour metastases. Nat. Rev. Cancer 9, 293–302 (2009).
Ma, L., Teruya-Feldstein, J. & Weinberg, R.A. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 449, 682–688 (2007).
Tavazoie, S.F. et al. Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451, 147–152 (2008).
Huang, Q. et al. The microRNAs miR-373 and miR-520c promote tumour invasion and metastasis. Nat. Cell Biol. 10, 202–210 (2008).
Valastyan, S. et al. A pleiotropically acting microRNA, miR-31, inhibits breast cancer metastasis. Cell 137, 1032–1046 (2009).
Ma, L. et al. miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin and cancer metastasis. Nat. Cell Biol. 12, 247–256 (2010).
Ma, L. et al. Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat. Biotechnol. 28, 341–347 (2010).
Huntsman, D.G. et al. Early gastric cancer in young, asymptomatic carriers of germ-line E-cadherin mutations. N. Engl. J. Med. 344, 1904–1909 (2001).
Chan, J.K. & Wong, C.S. Loss of E-cadherin is the fundamental defect in diffuse-type gastric carcinoma and infiltrating lobular carcinoma of the breast. Adv. Anat. Pathol. 8, 165–172 (2001).
Lewis, B.P., Burge, C.B. & Bartel, D.P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005).
Gearing, D.P. The leukemia inhibitory factor and its receptor. Adv. Immunol. 53, 31–58 (1993).
Kishimoto, T., Akira, S., Narazaki, M. & Taga, T. Interleukin-6 family of cytokines and gp130. Blood 86, 1243–1254 (1995).
Tamm, C., Bower, N. & Anneren, C. Regulation of mouse embryonic stem cell self-renewal by a Yes-YAP-TEAD2 signaling pathway downstream of LIF. J. Cell Sci. 124, 1136–1144 (2011).
Zhao, B. et al. Angiomotin is a novel Hippo pathway component that inhibits YAP oncoprotein. Genes Dev. 25, 51–63 (2011).
Zhao, B. et al. Inactivation of YAP oncoprotein by the Hippo pathway is involved in cell contact inhibition and tissue growth control. Genes Dev. 21, 2747–2761 (2007).
Zhao, B. et al. TEAD mediates YAP-dependent gene induction and growth control. Genes Dev. 22, 1962–1971 (2008).
Zhao, B., Li, L., Lei, Q. & Guan, K.L. The Hippo-YAP pathway in organ size control and tumorigenesis: an updated version. Genes Dev. 24, 862–874 (2010).
Pan, D. The hippo signaling pathway in development and cancer. Dev. Cell 19, 491–505 (2010).
Kang, Y. et al. A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 3, 537–549 (2003).
Dong, J. et al. Elucidation of a universal size-control mechanism in Drosophila and mammals. Cell 130, 1120–1133 (2007).
Cordenonsi, M. et al. The Hippo transducer TAZ confers cancer stem cell–related traits on breast cancer cells. Cell 147, 759–772 (2011).
Rhodes, D.R. et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 6, 1–6 (2004).
Richardson, A.L. et al. X chromosomal abnormalities in basal-like human breast cancer. Cancer Cell 9, 121–132 (2006).
Zhao, H. et al. Different gene expression patterns in invasive lobular and ductal carcinomas of the breast. Mol. Biol. Cell 15, 2523–2536 (2004).
Finak, G. et al. Stromal gene expression predicts clinical outcome in breast cancer. Nat. Med. 14, 518–527 (2008).
Sørlie, T. et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl. Acad. Sci. USA 98, 10869–10874 (2001).
Radvanyi, L. et al. The gene associated with trichorhinophalangeal syndrome in humans is overexpressed in breast cancer. Proc. Natl. Acad. Sci. USA 102, 11005–11010 (2005).
Sorlie, T. et al. Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc. Natl. Acad. Sci. USA 100, 8418–8423 (2003).
Turashvili, G. et al. Novel markers for differentiation of lobular and ductal invasive breast carcinomas by laser microdissection and microarray analysis. BMC Cancer 7, 55 (2007).
Perou, C.M. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000).
Karnoub, A.E. et al. Mesenchymal stem cells within tumour stroma promote breast cancer metastasis. Nature 449, 557–563 (2007).
Smith, S.C. & Theodorescu, D. Learning therapeutic lessons from metastasis suppressor proteins. Nat. Rev. Cancer 9, 253–264 (2009).
Stewart, S.A. et al. Lentivirus-delivered stable gene silencing by RNAi in primary cells. RNA 9, 493–501 (2003).
Rehmsmeier, M., Steffen, P., Hochsmann, M. & Giegerich, R. Fast and effective prediction of microRNA/target duplexes. RNA 10, 1507–1517 (2004).
Acknowledgements
We are grateful to R.A. Weinberg for his advice and reagents. We thank S. Ethier for providing cell lines; F. Reinhardt for advice on mouse surgery; the Genome Technology Core at the Whitehead Institute, the ShRNA and ORFeome Core at MD Anderson Cancer Center and the Histology Core Laboratories at MD Anderson Cancer Center and Memorial Sloan-Kettering Cancer Center for technical assistance; and members of the Ma Lab for discussion. We thank J. Chen, K. Muller, W. Pagel, K. Keyomarsi, L. Li and R. Cleveland for critical reading of the manuscript. This work is supported by the US National Institutes of Health grants R00CA138572 (to L.M.), R01CA166051 (to L.M.), R01CA109311 (to M.-C.H.) and P01CA099031 (to M.-C.H.), a Cancer Prevention and Research Institute of Texas Scholar Award R1004 (to L.M.), a University of Texas STARS Award (to L.M.), a Faculty Development Award (to L.M.) from the MD Anderson Cancer Center Support grant CA016672 from the US National Institutes of Health, Center for Biological Pathways (to Y.S. and M.-C.H.), a Susan G. Komen for the Cure grant SAC110016 (to M.-C.H.), the National Breast Cancer Foundation, Inc. and the Sister Institution Fund of China Medical University and Hospital and MD Anderson Cancer Center (to M.-C.H.).
Author information
Authors and Affiliations
Contributions
L.M. conceived of and supervised the project. D.C. and L.M. designed, performed and analyzed most of the experiments. Y.S. maintained shRNA and open reading frame (ORF) libraries, constructed RNA-Seq libraries and analyzed RNA-Seq data. Y.W. and M.-C.H. performed studies on tissue microarrays of human patient samples. P.Z. performed some biochemical work. A.H.R. and H.-K.L. performed tail vein injection experiments. J.T.-F. performed histopathological analysis. S.G. processed the raw data from the Solexa sequencer. H.L. performed TCGA data analysis. L.M. wrote the manuscript with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Discussion and Supplementary Figures 1–15 (PDF 2281 kb)
Supplementary Table
Supplementary Tables 1–4 (XLS 153 kb)
Rights and permissions
About this article
Cite this article
Chen, D., Sun, Y., Wei, Y. et al. LIFR is a breast cancer metastasis suppressor upstream of the Hippo-YAP pathway and a prognostic marker. Nat Med 18, 1511–1517 (2012). https://doi.org/10.1038/nm.2940
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2940
This article is cited by
-
PTHrP intracrine actions divergently influence breast cancer growth through p27 and LIFR
Breast Cancer Research (2024)
-
Targeting metastasis-initiating cancer stem cells in gastric cancer with leukaemia inhibitory factor
Cell Death Discovery (2024)
-
The Role of Breast Cancer Cells in Bone Metastasis: Suitable Seeds for Nourishing Soil
Current Osteoporosis Reports (2024)
-
Evaluation of the expression of the long non-coding RNAs, LOWEG and MINCR, and their clinical significance in human gastric cancer
Egyptian Journal of Medical Human Genetics (2023)
-
Predictive and prognostic biomarkers of bone metastasis in breast cancer: current status and future directions
Cell & Bioscience (2023)