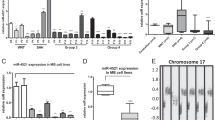
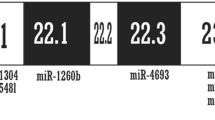
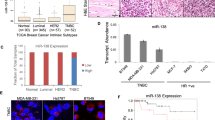
Abstract
Inactivation of the p53 tumor suppressor pathway allows cell survival in times of stress and occurs in many human cancers; however, normal embryonic stem cells and some cancers such as neuroblastoma maintain wild-type human TP53 and mouse Trp53 (referred to collectively as p53 herein). Here we describe a miRNA, miR-380-5p, that represses p53 expression via a conserved sequence in the p53 3′ untranslated region (UTR). miR-380-5p is highly expressed in mouse embryonic stem cells and neuroblastomas, and high expression correlates with poor outcome in neuroblastomas with neuroblastoma derived v-myc myelocytomatosis viral-related oncogene (MYCN) amplification. miR-380 overexpression cooperates with activated HRAS oncoprotein to transform primary cells, block oncogene-induced senescence and form tumors in mice. Conversely, inhibition of endogenous miR-380-5p in embryonic stem or neuroblastoma cells results in induction of p53, and extensive apoptotic cell death. In vivo delivery of a miR-380-5p antagonist decreases tumor size in an orthotopic mouse model of neuroblastoma. We demonstrate a new mechanism of p53 regulation in cancer and stem cells and uncover a potential therapeutic target for neuroblastoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Horn, H.F. & Vousden, K.H. Coping with stress: multiple ways to activate p53. Oncogene 26, 1306–1316 (2007).
Levine, A.J., Hu, W. & Feng, Z. The P53 pathway: what questions remain to be explored? Cell Death Differ. 13, 1027–1036 (2006).
Vogelstein, B. & Kinzler, K.W. Cancer genes and the pathways they control. Nat. Med. 10, 789–799 (2004).
Petitjean, A., Achatz, M.I., Borresen-Dale, A.L., Hainaut, P. & Olivier, M. TP53 mutations in human cancers: functional selection and impact on cancer prognosis and outcomes. Oncogene 26, 2157–2165 (2007).
Vazquez, A., Bond, E.E., Levine, A.J. & Bond, G.L. The genetics of the p53 pathway, apoptosis and cancer therapy. Nat. Rev. Drug Discov. 7, 979–987 (2008).
Martins, C.P., Brown-Swigart, L. & Evan, G.I. Modeling the therapeutic efficacy of p53 restoration in tumors. Cell 127, 1323–1334 (2006).
Ventura, A. et al. Restoration of p53 function leads to tumour regression in vivo. Nature 445, 661–665 (2007).
Xue, W. et al. Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature 445, 656–660 (2007).
Tweddle, D.A. et al. The p53 pathway and its inactivation in neuroblastoma. Cancer Lett. 197, 93–98 (2003).
Yoon, H. et al. Gene expression profiling of isogenic cells with different TP53 gene dosage reveals numerous genes that are affected by TP53 dosage and identifies CSPG2 as a direct target of p53. Proc. Natl. Acad. Sci. USA 99, 15632–15637 (2002).
Zhao, R. et al. Analysis of p53-regulated gene expression patterns using oligonucleotide arrays. Genes Dev. 14, 981–993 (2000).
Clarke, A.R., Gledhill, S., Hooper, M.L., Bird, C.C. & Wyllie, A.H. p53 dependence of early apoptotic and proliferative responses within the mouse intestinal epithelium following γ-irradiation. Oncogene 9, 1767–1773 (1994).
Clarke, A.R. et al. Thymocyte apoptosis induced by p53-dependent and independent pathways. Nature 362, 849–852 (1993).
Lowe, S.W., Schmitt, E.M., Smith, S.W., Osborne, B.A. & Jacks, T. p53 is required for radiation-induced apoptosis in mouse thymocytes. Nature 362, 847–849 (1993).
Gottlieb, E. et al. Transgenic mouse model for studying the transcriptional activity of the p53 protein: age- and tissue-dependent changes in radiation-induced activation during embryogenesis. EMBO J. 16, 1381–1390 (1997).
Venkatachalam, S. et al. Retention of wild-type p53 in tumors from p53 heterozygous mice: reduction of p53 dosage can promote cancer formation. EMBO J. 17, 4657–4667 (1998).
Halaby, M.J. & Yang, D.Q. p53 translational control: a new facet of p53 regulation and its implication for tumorigenesis and cancer therapeutics. Gene 395, 1–7 (2007).
Flynt, A.S. & Lai, E.C. Biological principles of microRNA-mediated regulation: shared themes amid diversity. Nat. Rev. Genet. 9, 831–842 (2008).
Ventura, A. & Jacks, T. MicroRNAs and cancer: short RNAs go a long way. Cell 136, 586–591 (2009).
Esquela-Kerscher, A. & Slack, F.J. Oncomirs—microRNAs with a role in cancer. Nat. Rev. Cancer 6, 259–269 (2006).
Enright, A.J. et al. MicroRNA targets in Drosophila. Genome Biol. 5, R1 (2003).
Saetrom, P. et al. Distance constraints between microRNA target sites dictate efficacy and cooperativity. Nucleic Acids Res. 35, 2333–2342 (2007).
Zhao, Y., Samal, E. & Srivastava, D. Serum response factor regulates a muscle-specific microRNA that targets Hand2 during cardiogenesis. Nature 436, 214–220 (2005).
Kuhn, D.E. et al. Experimental validation of miRNA targets. Methods 44, 47–54 (2008).
Seitz, H. et al. A large imprinted microRNA gene cluster at the mouse Dlk1-Gtl2 domain. Genome Res. 14, 1741–1748 (2004).
Tedeschi, A. & Di Giovanni, S. The non-apoptotic role of p53 in neuronal biology: enlightening the dark side of the moon. EMBO Rep. 10, 576–583 (2009).
Le, M.T. et al. MicroRNA-125b is a novel negative regulator of p53. Genes Dev. 23, 862–876 (2009).
Fu, L., Ma, W. & Benchimol, S. A translation repressor element resides in the 3′ untranslated region of human p53 mRNA. Oncogene 18, 6419–6424 (1999).
Mazan-Mamczarz, K. et al. RNA-binding protein HuR enhances p53 translation in response to ultraviolet light irradiation. Proc. Natl. Acad. Sci. USA 100, 8354–8359 (2003).
Sarkisian, C.J. et al. Dose-dependent oncogene-induced senescence in vivo and its evasion during mammary tumorigenesis. Nat. Cell Biol. 9, 493–505 (2007).
Swarbrick, A., Roy, E., Allen, T. & Bishop, J.M. Id1 cooperates with oncogenic Ras to induce metastatic mammary carcinoma by subversion of the cellular senescence response. Proc. Natl. Acad. Sci. USA 105, 5402–5407 (2008).
Welm, A.L., Kim, S., Welm, B.E. & Bishop, J.M. MET and MYC cooperate in mammary tumorigenesis. Proc. Natl. Acad. Sci. USA 102, 4324–4329 (2005).
Hemann, M.T. et al. An epi-allelic series of p53 hypomorphs created by stable RNAi produces distinct tumor phenotypes in vivo. Nat. Genet. 33, 396–400 (2003).
Scott, G.K. et al. Coordinate suppression of ERBB2 and ERBB3 by enforced expression of micro-RNA miR-125a or miR-125b. J. Biol. Chem. 282, 1479–1486 (2007).
Tweddle, D.A., Malcolm, A.J., Bown, N., Pearson, A.D. & Lunec, J. Evidence for the development of p53 mutations after cytotoxic therapy in a neuroblastoma cell line. Cancer Res. 61, 8–13 (2001).
Weiss, W.A., Aldape, K., Mohapatra, G., Feuerstein, B.G. & Bishop, J.M. Targeted expression of MYCN causes neuroblastoma in transgenic mice. EMBO J. 16, 2985–2995 (1997).
Chesler, L. et al. Chemotherapy-induced apoptosis in a transgenic model of neuroblastoma proceeds through p53 induction. Neoplasia 10, 1268–1274 (2008).
Liu, T. et al. Activation of tissue transglutaminase transcription by histone deacetylase inhibition as a therapeutic approach for Myc oncogenesis. Proc. Natl. Acad. Sci. USA 104, 18682–18687 (2007).
Chesler, L. et al. Inhibition of phosphatidylinositol 3-kinase destabilizes Mycn protein and blocks malignant progression in neuroblastoma. Cancer Res. 66, 8139–8146 (2006).
Hogarty, M.D. et al. ODC1 is a critical determinant of MYCN oncogenesis and a therapeutic target in neuroblastoma. Cancer Res. 68, 9735–9745 (2008).
Rounbehler, R.J. et al. Targeting ornithine decarboxylase impairs development of MYCN-amplified neuroblastoma. Cancer Res. 69, 547–553 (2009).
Davis, S. et al. Potent inhibition of microRNA in vivo without degradation. Nucleic Acids Res. 37, 70–77 (2009).
Stadtfeld, M. et al. Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. Nature 465, 175–181 (2010).
Kawamura, T. et al. Linking the p53 tumour suppressor pathway to somatic cell reprogramming. Nature 460, 1140–1144 (2009).
Kota, J. et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 137, 1005–1017 (2009).
Medina, P.P., Nolde, M. & Slack, F.J. OncomiR addiction in an in vivo model of microRNA-21–induced pre-B-cell lymphoma. Nature advance online publication, doi:10.1038/nature09284 (8 August 2010).
Ma, L. et al. Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat. Biotechnol. 28, 341–347 (2010).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Haber, M. et al. Association of high-level MRP1 expression with poor clinical outcome in a large prospective study of primary neuroblastoma. J. Clin. Oncol. 24, 1546–1553 (2006).
Grundhoff, A., Sullivan, C.S. & Ganem, D. A combined computational and microarray-based approach identifies novel microRNAs encoded by human gamma-herpesviruses. RNA 12, 733–750 (2006).
Acknowledgements
We gratefully thank J.M. Bishop, N.K. Hayward and R.L. Sutherland for their support of this project, the Children's Oncology Group and M. Grimmer (University of California–San Francisco) for providing tumor samples, D. Lynch and J. Brugge (Harvard Medical School) for MCF10A cells expressing the ecotropic receptor, R. Jaenisch (Whitehead Institute) for Trp53−/− ES cells and S. Lowe (Cold Spring Harbor) for the p53 shRNA retrovirus. TH-MYCN transgenic mice were from W. Weiss (University of California–San Francisco). This work was supported by grants from the US National Institutes of Health: P50-CA58207, K08-CA104032 and 1R01CA136717 (to A.G.), 5R01DC005991 (to N.L.), R01CA102321, R01NS055750 and P01CA081403 (to W.W.) and K08NS48118 (to R.B.); the Susan G. Komen Foundation; the University of California–San Francisco Program for Breakthrough Biomedical Research (to A.G.); the G.W. Hooper Foundation; the Australian National Health and Medical Research Council (to T.P., M.H., M.D.N. and A. Swarbrick) and the Cancer Institute New South Wales (M.H. and M.D.N.). A. Swarbrick is a recipient of a Cancer Institute New South Wales early career development fellowship, and R.L.J. received a US National Science Foundation fellowship. A.G. is a V-Foundation Scholar, A. Shaw is a Cancer Institute New South Wales Research Scholar and Y.P. is supported by an Australian Postgraduate Award from the Australian National Health and Medical Research Council.
Author information
Authors and Affiliations
Contributions
S.L.W., A. Swarbrick and A.G. conceived and designed the experiments, discussed the results and wrote the manuscript. A. Swarbrick, S.L.W., A. Shaw, Y.P., A.N., A.G., R.L.J., C.S.S., C.S.H., P.L., A.B., N.H., Y.C. and L.L. performed experiments. L.J.A. and M.D.N. performed statistical analysis of the human neuroblastoma data set. A.C. provided human samples and clinical data, and E.G.M. provided anti-miRs for in vivo studies. M.H., T.P., W.W., N.L. and C.S.S. supervised experiments or experimental design.
Corresponding authors
Ethics declarations
Competing interests
E.G.M. is an employee and shareholder of Regulus Therapeutics.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Table 1 and Supplementary Methods (PDF 1169 kb)
Rights and permissions
About this article
Cite this article
Swarbrick, A., Woods, S., Shaw, A. et al. miR-380-5p represses p53 to control cellular survival and is associated with poor outcome in MYCN-amplified neuroblastoma. Nat Med 16, 1134–1140 (2010). https://doi.org/10.1038/nm.2227
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2227
This article is cited by
-
Identification of PARN nuclease activity inhibitors by computational-based docking and high-throughput screening
Scientific Reports (2023)
-
miR-380-5p-mediated repression of TEP1 and TSPYL5 interferes with telomerase activity and favours the emergence of an “ALT-like” phenotype in diffuse malignant peritoneal mesothelioma cells
Journal of Hematology & Oncology (2017)
-
Neuroblastoma treatment in the post-genomic era
Journal of Biomedical Science (2017)
-
Species-specific mutual regulation of p53 and miR-138 between human, rat and mouse
Scientific Reports (2016)
-
MicroRNA-25 promotes gastric cancer migration, invasion and proliferation by directly targeting transducer of ERBB2, 1 and correlates with poor survival
Oncogene (2015)