Abstract
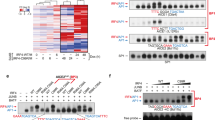
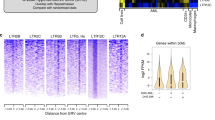
Mammalian genomes contain many repetitive elements, including long terminal repeats (LTRs), which have long been suspected to have a role in tumorigenesis. Here we present evidence that aberrant LTR activation contributes to lineage-inappropriate gene expression in transformed human cells and that such gene expression is central for tumor cell survival. We show that B cell–derived Hodgkin's lymphoma cells depend on the activity of the non-B, myeloid-specific proto-oncogene colony-stimulating factor 1 receptor (CSF1R). In these cells, CSF1R transcription initiates at an aberrantly activated endogenous LTR of the MaLR family (THE1B). Derepression of the THE1 subfamily of MaLR LTRs is widespread in the genome of Hodgkin's lymphoma cells and is associated with impaired epigenetic control due to loss of expression of the corepressor CBFA2T3. Furthermore, we detect LTR-driven CSF1R transcripts in anaplastic large cell lymphoma, in which CSF1R is known to be expressed aberrantly. We conclude that LTR derepression is involved in the pathogenesis of human lymphomas, a finding that might have diagnostic, prognostic and therapeutic implications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Jern, P. & Coffin, J.M. Effects of retroviruses on host genome function. Annu. Rev. Genet. 42, 709–732 (2008).
Smit, A.F. Identification of a new, abundant superfamily of mammalian LTR-transposons. Nucleic Acids Res. 21, 1863–1872 (1993).
Maksakova, I.A., Mager, D.L. & Reiss, D. Keeping active endogenous retroviral-like elements in check: the epigenetic perspective. Cell. Mol. Life Sci. 65, 3329–3347 (2008).
Faulkner, G.J. et al. The regulated retrotransposon transcriptome of mammalian cells. Nat. Genet. 41, 563–571 (2009).
Druker, R. & Whitelaw, E. Retrotransposon-derived elements in the mammalian genome: a potential source of disease. J. Inherit. Metab. Dis. 27, 319–330 (2004).
Ehrlich, M. DNA methylation in cancer: too much, but also too little. Oncogene 21, 5400–5413 (2002).
Fan, T. et al. DNA hypomethylation caused by Lsh deletion promotes erythroleukemia development. Epigenetics 3, 134–142 (2008).
Howard, G., Eiges, R., Gaudet, F., Jaenisch, R. & Eden, A. Activation and transposition of endogenous retroviral elements in hypomethylation induced tumors in mice. Oncogene 27, 404–408 (2008).
Eden, A., Gaudet, F., Waghmare, A. & Jaenisch, R. Chromosomal instability and tumors promoted by DNA hypomethylation. Science 300, 455 (2003).
Huang, J. et al. Lsh, an epigenetic guardian of repetitive elements. Nucleic Acids Res. 32, 5019–5028 (2004).
Borowitz, M.J., Béné, M.C., Harris, N.L., Porwit, A. & Matutes, E. Acute leukaemias of ambiguous lineage. in WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues (eds. Swerdlow, S.H., Campo, E., Harris, N.L., Jaffe, E.S., Pileri, S.A., Stein, H., Thiele, J. & Vardiman, J.W.) 149–155 (IARC, Lyon, 2008).
Feldman, A.L. et al. Clonally related follicular lymphomas and histiocytic/dendritic cell sarcomas: evidence for transdifferentiation of the follicular lymphoma clone. Blood 111, 5433–5439 (2008).
Küppers, R. The biology of Hodgkin′s lymphoma. Nat. Rev. Cancer 9, 15–27 (2009).
Janz, M., Dörken, B. & Mathas, S. Reprogramming of B lymphoid cells in human lymphoma pathogenesis. Cell Cycle 5, 1057–1061 (2006).
Küppers, R. et al. Identification of Hodgkin and Reed-Sternberg cell-specific genes by gene expression profiling. J. Clin. Invest. 111, 529–537 (2003).
Mathas, S. et al. Intrinsic inhibition of transcription factor E2A by HLH proteins ABF-1 and Id2 mediates reprogramming of neoplastic B cells in Hodgkin lymphoma. Nat. Immunol. 7, 207–215 (2006).
Pixley, F.J. & Stanley, E.R. CSF-1 regulation of the wandering macrophage: complexity in action. Trends Cell Biol. 14, 628–638 (2004).
Moreau, A., Praloran, V., Berrada, L., Coupey, L. & Gaillard, F. Immunohistochemical detection of cells positive for colony-stimulating factor 1 in lymph nodes from reactive lymphadenitis, and Hodgkin′s disease. Leukemia 6, 126–130 (1992).
Irvine, K.M. et al. A CSF-1 receptor kinase inhibitor targets effector functions and inhibits pro-inflammatory cytokine production from murine macrophage populations. FASEB J. 20, 1921–1923 (2006).
Burns, C.J. et al. Discovery of 2-(α-methylbenzylamino) pyrazines as potent Type II inhibitors of FMS. Bioorg. Med. Chem. Lett. 19, 1206–1209 (2009).
Bonifer, C. & Hume, D.A. The transcriptional regulation of the colony-stimulating factor 1 receptor (CSF1R) gene during hematopoiesis. Front. Biosci. 13, 549–560 (2008).
Jundt, F. et al. Loss of PU.1 expression is associated with defective immunoglobulin transcription in Hodgkin and Reed-Sternberg cells of classical Hodgkin disease. Blood 99, 3060–3062 (2002).
Follows, G.A., Tagoh, H., Lefevre, P., Morgan, G.J. & Bonifer, C. Differential transcription factor occupancy but evolutionarily conserved chromatin features at the human and mouse M-CSF (CSF-1) receptor loci. Nucleic Acids Res. 31, 5805–5816 (2003).
Visvader, J. & Verma, I.M. Differential transcription of exon 1 of the human c-fms gene in placental trophoblasts and monocytes. Mol. Cell. Biol. 9, 1336–1341 (1989).
Hug, B.A. & Lazar, M.A. ETO interacting proteins. Oncogene 23, 4270–4274 (2004).
Joos, S. et al. Classical Hodgkin lymphoma is characterized by recurrent copy number gains of the short arm of chromosome 2. Blood 99, 1381–1387 (2002).
Ohshima, K. et al. Chromosome 16q deletion and loss of E-cadherin expression in Hodgkin and Reed-Sternberg cells. Int. J. Cancer 92, 678–682 (2001).
Delhase, M., Hayakawa, M., Chen, Y. & Karin, M. Positive and negative regulation of IκB kinase activity through IKKβ subunit phosphorylation. Science 284, 309–313 (1999).
Mathas, S. et al. Gene deregulation and spatial genome reorganization near breakpoints prior to formation of translocations in anaplastic large cell lymphoma. Proc. Natl. Acad. Sci. USA 106, 5831–5836 (2009).
Kacinski, B.M. CSF-1 and its receptor in ovarian, endometrial and breast cancer. Ann. Med. 27, 79–85 (1995).
Chambers, S.K., Kacinski, B.M., Ivins, C.M. & Carcangiu, M.L. Overexpression of epithelial macrophage colony-stimulating factor (CSF-1) and CSF-1 receptor: a poor prognostic factor in epithelial ovarian cancer, contrasted with a protective effect of stromal CSF-1. Clin. Cancer Res. 3, 999–1007 (1997).
Kluger, H.M. et al. Macrophage colony-stimulating factor-1 receptor expression is associated with poor outcome in breast cancer by large cohort tissue microarray analysis. Clin. Cancer Res. 10, 173–177 (2004).
Gamou, T. et al. The partner gene of AML1 in t(16;21) myeloid malignancies is a novel member of the MTG8(ETO) family. Blood 91, 4028–4037 (1998).
Ehlers, A. et al. Histone acetylation and DNA demethylation of B cells result in a Hodgkin-like phenotype. Leukemia 22, 835–841 (2008).
Chyla, B.J. et al. Deletion of Mtg16, a target of t(16;21), alters hematopoietic progenitor cell proliferation and lineage allocation. Mol. Cell. Biol. 28, 6234–6247 (2008).
Mathas, S. et al. Aberrantly expressed c-Jun and JunB are a hallmark of Hodgkin lymphoma cells, stimulate proliferation and synergize with NF-κB. EMBO J. 21, 4104–4113 (2002).
Goardon, N. et al. ETO2 coordinates cellular proliferation and differentiation during erythropoiesis. EMBO J. 25, 357–366 (2006).
Walter, K., Bonifer, C. & Tagoh, H. Stem cell-specific epigenetic priming and B cell-specific transcriptional activation at the mouse Cd19 locus. Blood 112, 1673–1682 (2008).
Kumar, R. et al. CBFA2T3–ZNF652 corepressor complex regulates transcription of the E-box gene HEB. J. Biol. Chem. 283, 19026–19038 (2008).
Acknowledgements
We thank F. Hummel, S. Kressmann, S. Meier, C. Cieluch, B. Wollert-Wulf and R. Zühlke-Jenisch for outstanding technical assistance, H. Tagoh for help with experiments, P. Rahn for cell sorting and K. Rajewsky for helpful discussion and critical reading of the manuscript. M.G. is funded by the Federation of European Biochemical Societies. This work was supported in part by grants from the Deutsche Forschungsgemeinschaft (SFB/TRR54), the Wilhelm Sander-Stiftung, the Deutsche Krebshilfe, the KinderKrebsInitiative Buchholz/Holm-Seppensen, Susan G. Komen for the Cure, Leukaemia and Lymphoma Research, Cancer Research UK and Yorkshire Cancer Research. We also thank V. Diehl (University Cologne) for cell lines L1236 and L591, A. Engert (University Cologne) for L540Cy cells, P. Krammer (German Cancer Research Center Heidelberg) for BJAB cells, L. Staudt (US National Cancer Institute) for OCI-Ly3 and OCI-Ly10 cells, M. Falk (Helmholtz Zentrum München) for BL-60 cells and S. Rosen (Northwestern University) for MM1.S cells.
Author information
Authors and Affiliations
Contributions
B.L., K.W. and S.K. designed and performed experiments, interpreted data and contributed to writing of the manuscript; D.F.C., R.K., D.L. and M.F.H. gave technical support and contributed material; M.H., K.J., H.S. and I.A. performed and interpreted IHC and ISH analyses; J.R., M.G. and R. Siebert designed, performed and interpreted bisulfite pyrosequencing and FICTION analyses; M.A.B., K.D.W., R.B., E.S. and R. Stadhouders performed experiments; P.N.C. designed TD-PCR experiments and interpreted data; K.K. analyzed microarray data and performed real-time PCR analyses; M.J. and B.D. interpreted data and contributed to the writing of the manuscript; C.B. and S.M. designed research, interpreted data, wrote the manuscript and supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Tables 1–5 and Supplementary Methods (PDF 4469 kb)
Rights and permissions
About this article
Cite this article
Lamprecht, B., Walter, K., Kreher, S. et al. Derepression of an endogenous long terminal repeat activates the CSF1R proto-oncogene in human lymphoma. Nat Med 16, 571–579 (2010). https://doi.org/10.1038/nm.2129
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2129
This article is cited by
-
Activation of human endogenous retroviruses and its physiological consequences
Nature Reviews Molecular Cell Biology (2024)
-
Transposable elements as essential elements in the control of gene expression
Mobile DNA (2023)
-
Pan-cancer analysis identifies tumor-specific antigens derived from transposable elements
Nature Genetics (2023)
-
Transposable elements as tissue-specific enhancers in cancers of endodermal lineage
Nature Communications (2023)
-
An alternative CYB5A transcript is expressed in aneuploid ALL and enriched in relapse
BMC Genomic Data (2022)