Abstract
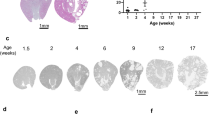
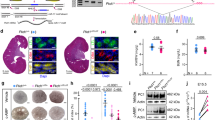
Hepatocyte nuclear factor-1β (HNF-1β) is a transcription factor required for the expression of several renal cystic genes and whose prenatal deletion leads to polycystic kidney disease (PKD)1. We show here that inactivation of Hnf1b from postnatal day 10 onward does not elicit cystic dilations in tubules after their proliferative morphogenetic elongation is over. Cystogenic resistance is intrinsically linked to the quiescent state of cells. In fact, when Hnf1b deficient quiescent cells are forced to proliferate by an ischemia-reperfusion injury, they give rise to cysts, owing to loss of oriented cell division. Remarkably, in quiescent cells, the transcription of crucial cystogenic target genes is maintained even in the absence of HNF-1β. However, their expression is lost as soon as cells proliferate and the chromatin of target genes acquires heterochromatin marks. These results unveil a previously undescribed aspect of gene regulation. It is well established that transcription is shut off during the mitotic condensation of chromatin2,3. We propose that transcription factors such as HNF-1β might be involved in reprogramming gene expression after transcriptional silencing is induced by mitotic chromatin condensation. Notably, HNF-1β remains associated with the mitotically condensed chromosomal barrels. This association suggests that HNF-1β is a bookmarking factor that is necessary for reopening the chromatin of target genes after mitotic silencing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gresh, L. et al. A transcriptional network in polycystic kidney disease. EMBO J. 23, 1657–1668 (2004).
Gottesfeld, J.M. & Forbes, D.J. Mitotic repression of the transcriptional machinery. Trends Biochem. Sci. 22, 197–202 (1997).
Komura, J., Ikehata, H. & Ono, T. Chromatin fine structure of the c-MYC insulator element/DNase I-hypersensitive site I is not preserved during mitosis. Proc. Natl. Acad. Sci. USA 104, 15741–15746 (2007).
Torres, V.E., Harris, P.C. & Pirson, Y. Autosomal dominant polycystic kidney disease. Lancet 369, 1287–1301 (2007).
Fischer, E., Gresh, L., Reimann, A. & Pontoglio, M. Cystic kidney diseases: learning from animal models. Nephrol. Dial. Transplant. 19, 2700–2702 (2004).
Guay-Woodford, L.M. Murine models of polycystic kidney disease: molecular and therapeutic insights. Am. J. Physiol. Renal Physiol. 285, F1034–F1049 (2003).
Davenport, J.R. et al. Disruption of intraflagellar transport in adult mice leads to obesity and slow-onset cystic kidney disease. Curr. Biol. 17, 1586–1594 (2007).
Lantinga-van Leeuwen, I.S. et al. Kidney-specific inactivation of the Pkd1 gene induces rapid cyst formation in developing kidneys and a slow onset of disease in adult mice. Hum. Mol. Genet. 16, 3188–3196 (2007).
Piontek, K., Menezes, L.F., Garcia-Gonzalez, M.A., Huso, D.L. & Germino, G.G. A critical developmental switch defines the kinetics of kidney cyst formation after loss of Pkd1. Nat. Med. 13, 1490–1495 (2007).
Patel, V. et al. Acute kidney injury and aberrant planar cell polarity induce cyst formation in mice lacking renal cilia. Hum. Mol. Genet. 17, 1578–1590 (2008).
Takakura, A., Contrino, L., Beck, A.W. & Zhou, J. Pkd1 inactivation induced in adulthood produces focal cystic disease. J. Am. Soc. Nephrol. 19, 2351–2363 (2008).
Coffinier, C., Barra, J., Babinet, C. & Yaniv, M. Expression of the vHNF-1/HNF-1β homeoprotein gene during mouse organogenesis. Mech. Dev. 89, 211–213 (1999).
Lindner, T.H. et al. A novel syndrome of diabetes mellitus, renal dysfunction and genital malformation associated with a partial deletion of the pseudo-POU domain of hepatocyte nuclear factor-1β. Hum. Mol. Genet. 8, 2001–2008 (1999).
Ulinski, T. et al. Renal phenotypes related to hepatocyte nuclear factor-1β (TCF2) mutations in a pediatric cohort. J. Am. Soc. Nephrol. 17, 497–503 (2006).
Kühn, R., Schwenk, F., Aguet, M. & Rajewsky, K. Inducible gene targeting in mice. Science 269, 1427–1429 (1995).
Coffinier, C. et al. Bile system morphogenesis defects and liver dysfunction upon targeted deletion of HNF-1β. Development 129, 1829–1838 (2002).
Soriano, P. Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat. Genet. 21, 70–71 (1999).
Igarashi, P. & Somlo, S. Genetics and pathogenesis of polycystic kidney disease. J. Am. Soc. Nephrol. 13, 2384–2398 (2002).
Marquez, M.G., Cabrera, I., Serrano, D.J. & Sterin-Speziale, N. Cell proliferation and morphometric changes in the rat kidney during postnatal development. Anat. Embryol. (Berl.) 205, 431–440 (2002).
Nony, P.A. & Schnellmann, R.G. Mechanisms of renal cell repair and regeneration after acute renal failure. J. Pharmacol. Exp. Ther. 304, 905–912 (2003).
Fischer, E. et al. Defective planar cell polarity in polycystic kidney disease. Nat. Genet. 38, 21–23 (2006).
Gong, Y. et al. HNF-1β regulates transcription of the PKD modifier gene Kif12. J. Am. Soc. Nephrol. 20, 41–47 (2009).
Pontoglio, M., Faust, D.M., Doyen, A., Yaniv, M. & Weiss, M.C. Hepatocyte nuclear factor 1alpha gene inactivation impairs chromatin remodeling and demethylation of the phenylalanine hydroxylase gene. Mol. Cell. Biol. 17, 4948–4956 (1997).
Ma, Z. et al. Mutations of HNF-1β inhibit epithelial morphogenesis through dysregulation of SOCS-3. Proc. Natl. Acad. Sci. USA 104, 20386–20391 (2007).
Takakura, A. et al. Renal injury is a third hit promoting rapid development of adult polycystic kidney disease. Hum. Mol. Genet. 18, 2523–2531 (2009).
Bhunia, A.K. et al. PKD1 induces p21(waf1) and regulation of the cell cycle via direct activation of the JAK-STAT signaling pathway in a process requiring PKD2. Cell 109, 157–168 (2002).
Li, X. et al. Polycystin-1 and polycystin-2 regulate the cell cycle through the helix-loop-helix inhibitor Id2. Nat. Cell Biol. 7, 1202–1212 (2005).
Bukanov, N.O., Smith, L.A., Klinger, K.W., Ledbetter, S.R. & Ibraghimov-Beskrovnaya, O. Long-lasting arrest of murine polycystic kidney disease with CDK inhibitor roscovitine. Nature 444, 949–952 (2006).
Runembert, I. et al. Recovery of Na-glucose cotransport activity after renal ischemia is impaired in mice lacking vimentin. Am. J. Physiol. Renal Physiol. 287, F960–F968 (2004).
Park, K.M., Kim, J.I., Ahn, Y., Bonventre, A.J. & Bonventre, J.V. Testosterone is responsible for enhanced susceptibility of males to ischemic renal injury. J. Biol. Chem. 279, 52282–52292 (2004).
Pontoglio, M. et al. Hepatocyte nuclear factor 1 inactivation results in hepatic dysfunction, phenylketonuria and renal Fanconi syndrome. Cell 84, 575–585 (1996).
Nelson, J.D., Denisenko, O. & Bomsztyk, K. Protocol for the fast chromatin immunoprecipitation (ChIP) method. Nat. Protoc. 1, 179–185 (2006).
Acknowledgements
We are grateful to S. Bettiol, J.B. Weitzman and M. Yaniv for critical advice. We are grateful to R. Sandford (Cambridge Institute of Medical Research) for Pkd2-specific antibodies and to B. Viollet, (Institut Cochin, Paris) for the GFP–HNF-4α fusion construct. MxCre mice were kindly provided by K. Rajewsky (Harvard Medical School, Boston), and the ROSA26R strain was provided by P. Soriano (Fred Hutchinson Cancer Research Center, Seattle). We thank E. Perret and P. Roux from the “Plate-Forme d'Imagerie Dynamique,” Institut Pasteur, Paris) for assistance and advice. This work was supported by Centre National de la Recherche Scientifique, Institut National de la Santé et de la Recherche Médicale, Université Paris Descartes, Polycystic Kidney Disease Foundation, Société de Néphrologie, Agence Nationale Recherche, Fondation de la Recherche Médicale. F.V. was a recipient of a fellowship from Société de Néphrologie and from Fundación La Caixa.
Author information
Authors and Affiliations
Contributions
F.V., experimental mouse model development, X-gal staining, immunofluorescence analysis, gene expression analysis; chromatin immunoprecipitation studies, cell culture and time-lapse video microscopy. S.L.C., renal ishemia-reperfusion injury, histological studies, immunohistochemistry, mitotic angle measurements and gene expression studies. E.F., experimental design, mitotic angle measurements and experimental mouse model development. C.C., GFP fusion proteins and cell culture studies. S.G., bioinformatic analysis of genomic HNF-1 binding sites. A.D., mouse breeding and genotyping. P.I., cell line expressing dominant-negative HNF-1β. F.T. supervised the studies and wrote the paper. M.P. supervised the project and wrote the paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Table 1, Supplementary Data and Supplementary Methods (PDF 1683 kb)
Supplementary Video 1
Localization of HNF1β and HNF4α GFP fusion proteins during mitosis. Contrary to the vast majority of transcription factors (exemplified here by HNF4α) HNF1β remains associated with the chromatin during mitosis. (To play, view as slideshow.) (PPT 10368 kb)
Rights and permissions
About this article
Cite this article
Verdeguer, F., Le Corre, S., Fischer, E. et al. A mitotic transcriptional switch in polycystic kidney disease. Nat Med 16, 106–110 (2010). https://doi.org/10.1038/nm.2068
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2068
This article is cited by
-
Mechanisms of ion transport regulation by HNF1β in the kidney: beyond transcriptional regulation of channels and transporters
Pflügers Archiv - European Journal of Physiology (2022)
-
HNF-1β as an immunohistochemical marker for distinguishing chromophobe renal cell carcinoma and hybrid oncocytic tumors from renal oncocytoma
Virchows Archiv (2021)
-
mTOR and S6K1 drive polycystic kidney by the control of Afadin-dependent oriented cell division
Nature Communications (2020)
-
A changing paradigm of transcriptional memory propagation through mitosis
Nature Reviews Molecular Cell Biology (2019)
-
Autosomal dominant tubulointerstitial kidney disease
Nature Reviews Disease Primers (2019)