Abstract
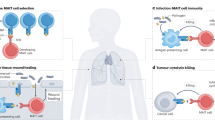
The identification of pattern-recognition receptors that selectively respond to evolutionarily conserved chemical (often pathogen-derived) moieties has provided key insight into how innate immune cells facilitate rapid and relatively specific antimicrobial immune activity. In contrast, relatively slower adaptive immune responses rely on T cell clonal expansion that develops in response to variable peptides bound to the groove of classical major histocompatibility complex (MHC) proteins. For certain nonclassical 'MHC-like' class Ib proteins, such as H2-M3 and CD1d, their respective binding grooves seem to have been adapted to present to T cells unique molecular patterns analogous to those involved in innate signaling. Here we propose that another MHC class Ib protein, MR1, which is required for the gut flora–dependent development of mucosa-associated invariant T cells, presents either a microbe-produced or a microbe-induced pattern.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Colmone, A. & Wang, C.R. H2–M3-restricted T cell response to infection. Microbes Infect. 8, 2277–2283 (2006).
Kinjo, Y. & Kronenberg, M. Vα14i NKT cells are innate lymphocytes that participate in the immune response to diverse microbes. J. Clin. Immunol. 25, 522–533 (2005).
Mattner, J. et al. Exogenous and endogenous glycolipid antigens activate NKT cells during microbial infections. Nature 434, 525–529 (2005).
Ploss, A., Tran, A., Menet, E., Leiner, I. & Pamer, E.G. Cross-recognition of N-formylmethionine peptides is a general characteristic of H2–M3-restricted CD8+ T cells. Infect. Immun. 73, 4423–4426 (2005).
Brutkiewicz, R.R. CD1d ligands: the good, the bad, and the ugly. J. Immunol. 177, 769–775 (2006).
Hashimoto, K., Hirai, M. & Kurosawa, Y. A gene outside the human MHC related to classical HLA class I genes. Science 269, 693–695 (1995).
Shiina, T. et al. Genomic anatomy of a premier major histocompatibility complex paralogous region on chromosome 1q21-q22. Genome Res. 11, 789–802 (2001).
Riegert, P., Wanner, V. & Bahram, S. Genomics, isoforms, expression, and phylogeny of the MHC class I-related MR1 gene. J. Immunol. 161, 4066–4077 (1998).
Yamaguchi, H., Hirai, M., Kurosawa, Y. & Hashimoto, K. A highly conserved major histocompatibility complex class I-related gene in mammals. Biochem. Biophys. Res. Commun. 238, 697–702 (1997).
Walter, L. & Gunther, E. Isolation and molecular characterization of the rat MR1 homologue, a non-MHC-linked class I-related gene. Immunogenetics 47, 477–482 (1998).
Parra-Cuadrado, J.F. et al. A study on the polymorphism of human MHC class I-related MR1 gene and identification of an MR1-like pseudogene. Tissue Antigens 56, 170–172 (2000).
Yamaguchi, H., Kurosawa, Y. & Hashimoto, K. Expanded genomic organization of conserved mammalian MHC class I-related genes, human MR1 and its murine ortholog. Biochem. Biophys. Res. Commun. 250, 558–564 (1998).
Hughes, A.L. & Nei, M. Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 335, 167–170 (1988).
Wang, C.R., Lambracht, D., Wonigeit, K., Howard, J.C. & Lindahl, K.F. Rat RT1 orthologs of mouse H2-M class Ib genes. Immunogenetics 42, 63–67 (1995).
Doyle, C.K., Davis, B.K., Cook, R.G., Rich, R.R. & Rodgers, J.R. Hyperconservation of the N-formyl peptide binding site of M3: evidence that M3 is an old eutherian molecule with conserved recognition of a pathogen-associated molecular pattern. J. Immunol. 171, 836–844 (2003).
Miller, M.M. et al. Characterization of two avian MHC-like genes reveals an ancient origin of the CD1 family. Proc. Natl. Acad. Sci. USA 102, 8674–8679 (2005).
Miley, M.J. et al. Biochemical features of the MHC-related protein 1 consistent with an immunological function. J. Immunol. 170, 6090–6098 (2003).
Yamaguchi, H. & Hashimoto, K. Association of MR1 protein, an MHC class I-related molecule, with β2-microglobulin. Biochem. Biophys. Res. Commun. 290, 722–729 (2002).
Treiner, E. et al. Selection of evolutionarily conserved mucosal-associated invariant T cells by MR1. Nature 422, 164–169 (2003).
Chiu, N.M., Chun, T., Fay, M., Mandal, M. & Wang, C.R. The majority of H2–M3 is retained intracellularly in a peptide-receptive state and traffics to the cell surface in the presence of N-formylated peptides. J. Exp. Med. 190, 423–434 (1999).
Zwickey, H.L. & Potter, T.A. Antigen secreted from noncytosolic Listeria monocytogenes is processed by the classical MHC class I processing pathway. J. Immunol. 162, 6341–6350 (1999).
Chun, T. et al. Functional roles of TAP and tapasin in the assembly of M3-N-formylated peptide complexes. J. Immunol. 167, 1507–1514 (2001).
Tilloy, F. et al. An invariant T cell receptor α chain defines a novel TAP-independent major histocompatibility complex class Ib-restricted α/β T cell subpopulation in mammals. J. Exp. Med. 189, 1907–1921 (1999).
Treiner, E. et al. Mucosal-associated invariant T (MAIT) cells: an evolutionarily conserved T cell subset. Microbes Infect. 7, 552–559 (2005).
Yoshimoto, T. & Paul, W.E. CD4pos, NK1.1pos T cells promptly produce interleukin 4 in response to in vivo challenge with anti-CD3. J. Exp. Med. 179, 1285–1295 (1994).
Stetson, D.B. et al. Constitutive cytokine mRNAs mark natural killer (NK) and NK T cells poised for rapid effector function. J. Exp. Med. 198, 1069–1076 (2003).
Treiner, E. & Lantz, O. CD1d- and MR1-restricted invariant T cells: of mice and men. Curr. Opin. Immunol. 18, 519–526 (2006).
Kawachi, I., Maldonado, J., Strader, C. & Gilfillan, S. MR1-restricted Vα19i mucosal-associated invariant T cells are innate T cells in the gut lamina propria that provide a rapid and diverse cytokine response. J. Immunol. 176, 1618–1627 (2006).
Croxford, J.L., Miyake, S., Huang, Y.Y., Shimamura, M. & Yamamura, T. Invariant Vα19i T cells regulate autoimmune inflammation. Nat. Immunol. 7, 987–994 (2006).
Shimamura, M. & Huang, Y.Y. Presence of a novel subset of NKT cells bearing an invariant Vα19.1-Jα26 TCR α chain. FEBS Lett. 516, 97–100 (2002).
Xu, H., Chun, T., Choi, H.J., Wang, B. & Wang, C.R. Impaired response to Listeria in H2–M3-deficient mice reveals a nonredundant role of MHC class Ib-specific T cells in host defense. J. Exp. Med. 203, 449–459 (2006).
Bendelac, A. Positive selection of mouse NK1+ T cells by CD1-expressing cortical thymocytes. J. Exp. Med. 182, 2091–2096 (1995).
Urdahl, K.B., Sun, J.C. & Bevan, M.J. Positive selection of MHC class Ib-restricted CD8+ T cells on hematopoietic cells. Nat. Immunol. 3, 772–779 (2002).
Cannarile, M.A., Decanis, N., van Meerwijk, J.P. & Brocker, T. The role of dendritic cells in selection of classical and nonclassical CD8+ T cells in vivo. J. Immunol. 173, 4799–4805 (2004).
Lodoen, M.B. & Lanier, L.L. Viral modulation of NK cell immunity. Nat. Rev. Microbiol. 3, 59–69 (2005).
Gasser, S. & Raulet, D.H. Activation and self-tolerance of natural killer cells. Immunol. Rev. 214, 130–142 (2006).
Kelley, L.A., MacCallum, R.M. & Sternberg, M.J. Enhanced genome annotation using structural profiles in the program 3D-PSSM. J. Mol. Biol. 299, 499–520 (2000).
O'Callaghan, C.A. et al. Structural features impose tight peptide binding specificity in the nonclassical MHC molecule HLA-E. Mol. Cell 1, 531–541 (1998).
Wang, C.R., Lindahl, K.F. & Deisenhofer, J. Crystal structure of the MHC class Ib molecule H2–M3. Res. Immunol. 147, 313–321 (1996).
Sanchez, L.M., Chirino, A.J. & Bjorkman, P. Crystal structure of human ZAG, a fat-depleting factor related to MHC molecules. Science 283, 1914–1919 (1999).
Okamoto, N. et al. Synthetic α-mannosyl ceramide as a potent stimulant for an NKT cell repertoire bearing the invariant Vα19-Jα26 TCR α chain. Chem. Biol. 12, 677–683 (2005).
Koch, M. et al. The crystal structure of human CD1d with and without α-galactosylceramide. Nat. Immunol. 6, 819–826 (2005).
Huang, S. et al. Evidence for MR1 antigen presentation to mucosal-associated invariant T cells. J. Biol. Chem. 280, 21183–21193 (2005).
Jayawardena-Wolf, J., Benlagha, K., Chiu, Y.H., Mehr, R. & Bendelac, A. CD1d endosomal trafficking is independently regulated by an intrinsic CD1d-encoded tyrosine motif and by the invariant chain. Immunity 15, 897–908 (2001).
Illes, Z., Shimamura, M., Newcombe, J., Oka, N. & Yamamura, T. Accumulation of Vα7.2-Jα33 invariant T cells in human autoimmune inflammatory lesions in the nervous system. Int. Immunol. 16, 223–230 (2004).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Jeanmougin, F., Thompson, J.D., Gouy, M., Higgins, D.G. & Gibson, T.J. Multiple sequence alignment with Clustal X. Trends Biochem. Sci. 23, 403–405 (1998).
Acknowledgements
We thank S. Gilfillan for discussions and M. Miley for help with computational studies. Supported by the National Institute of Allergy and Infectious Diseases.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Hansen, T., Huang, S., Arnold, P. et al. Patterns of nonclassical MHC antigen presentation. Nat Immunol 8, 563–568 (2007). https://doi.org/10.1038/ni1475
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni1475
This article is cited by
-
Donor-unrestricted T cells in the human CD1 system
Immunogenetics (2016)
-
Structure and function of the non-classical major histocompatibility complex molecule MR1
Immunogenetics (2016)
-
MAIT cells are depleted early but retain functional cytokine expression in HIV infection
Immunology & Cell Biology (2015)
-
Unusual evolutionary conservation and further species-specific adaptations of a large family of nonclassical MHC class Ib genes across different degrees of genome ploidy in the amphibian subfamily Xenopodinae
Immunogenetics (2014)
-
Evolution of nonclassical MHC-dependent invariant T cells
Cellular and Molecular Life Sciences (2014)