Abstract
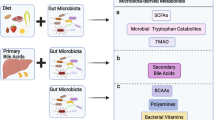
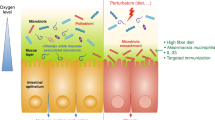
The study of the intestinal microbiota has begun to shift from cataloging individual members of the commensal community to understanding their contributions to the physiology of the host organism in health and disease. Here, we review the effects of the microbiome on innate and adaptive immunological players from epithelial cells and antigen-presenting cells to innate lymphoid cells and regulatory T cells. We discuss recent studies that have identified diverse microbiota-derived bioactive molecules and their effects on inflammation within the intestine and distally at sites as anatomically remote as the brain. Finally, we highlight new insights into how the microbiome influences the host response to infection, vaccination and cancer, as well as susceptibility to autoimmune and neurodegenerative disorders.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sender, R., Fuchs, S. & Milo, R. Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell 164, 337–340 (2016).
Gilbert, J.A. et al. Microbiome-wide association studies link dynamic microbial consortia to disease. Nature 535, 94–103 (2016).
Honda, K. & Littman, D.R. The microbiota in adaptive immune homeostasis and disease. Nature 535, 75–84 (2016).
Sonnenburg, J.L. & Bäckhed, F. Diet-microbiota interactions as moderators of human metabolism. Nature 535, 56–64 (2016).
Bittinger, K. et al. Improved characterization of medically relevant fungi in the human respiratory tract using next-generation sequencing. Genome Biol. 15, 487 (2014).
Iliev, I.D. et al. Interactions between commensal fungi and the C-type lectin receptor dectin-1 influence colitis. Science 336, 1314–1317 (2012).
Iliev, I.D. & Leonardi, I. Fungal dysbiosis: immunity and interactions at mucosal barriers. Nat. Rev. Immunol. http://dx.doi.org/10.1038/nri.2017.55 (2017).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Human Microbiome Project Consortium. A framework for human microbiome research. Nature 486, 215–221 (2012).
Browne, H.P. et al. Culturing of 'unculturable' human microbiota reveals novel taxa and extensive sporulation. Nature 533, 543–546 (2016).
Schloss, P.D., Iverson, K.D., Petrosino, J.F. & Schloss, S.J. The dynamics of a family's gut microbiota reveal variations on a theme. Microbiome 2, 25 (2014).
Yissachar, N. et al. An intestinal organ culture system uncovers a role for the nervous system in microbe-immune crosstalk. Cell 168, 1135–1148.e12 (2017).
Longman, R.S. & Littman, D.R. The functional impact of the intestinal microbiome on mucosal immunity and systemic autoimmunity. Curr. Opin. Rheumatol. 27, 381–387 (2015).
Roy, S. & Trinchieri, G. Microbiota: a key orchestrator of cancer therapy. Nat. Rev. Cancer 17, 271–285 (2017).
Smith, P.A. The tantalizing links between gut microbes and the brain. Nature 526, 312–314 (2015).
Hooper, L.V., Littman, D.R. & Macpherson, A.J. Interactions between the microbiota and the immune system. Science 336, 1268–1273 (2012).
Peterson, L.W. & Artis, D. Intestinal epithelial cells: regulators of barrier function and immune homeostasis. Nat. Rev. Immunol. 14, 141–153 (2014).
Blander, J.M. Death in the intestinal epithelium-basic biology and implications for inflammatory bowel disease. FEBS J. 283, 2720–2730 (2016).
Cummings, R.J. et al. Different tissue phagocytes sample apoptotic cells to direct distinct homeostasis programs. Nature 539, 565–569 (2016).
Donaldson, G.P., Lee, S.M. & Mazmanian, S.K. Gut biogeography of the bacterial microbiota. Nat. Rev. Microbiol. 14, 20–32 (2016).
Chu, H. et al. Gene-microbiota interactions contribute to the pathogenesis of inflammatory bowel disease. Science 352, 1116–1120 (2016).
Bunker, J.J. et al. Innate and adaptive humoral responses coat distinct commensal bacteria with immunoglobulin A. Immunity 43, 541–553 (2015).
Leone, V. et al. Effects of diurnal variation of gut microbes and high-fat feeding on host circadian clock function and metabolism. Cell Host Microbe 17, 681–689 (2015).
Thaiss, C.A. et al. Microbiota diurnal rhythmicity programs host transcriptome oscillations. Cell 167, 1495–1510.e12 (2016).
Thaiss, C.A., Zeevi, D., Levy, M., Segal, E. & Elinav, E. A day in the life of the meta-organism: diurnal rhythms of the intestinal microbiome and its host. Gut Microbes 6, 137–142 (2015).
Kau, A.L. et al. Functional characterization of IgA-targeted bacterial taxa from undernourished Malawian children that produce diet-dependent enteropathy. Sci. Transl. Med. 7, 276ra24 (2015)This defining study used Bug-FACS to identify IgA-reactive microbiota from mice colonized with human microbiota from twins discordant for malnutrition. This study illustrates the utility of Bug-FACS in identifying immunologically relevant microbiota in human disease.
Palm, N.W. et al. Immunoglobulin A coating identifies colitogenic bacteria in inflammatory bowel disease. Cell 158, 1000–1010 (2014). This study, along with ref. 22 , describes the method of IgA-seq to sort and sequence IgA-coated microbiota. Culture libraries created from IgA-sorted microbiota were used to evaluate the effects of these microbiota in vivo.
Planer, J.D. et al. Development of the gut microbiota and mucosal IgA responses in twins and gnotobiotic mice. Nature 534, 263–266 (2016).
Moor, K. et al. High-avidity IgA protects the intestine by enchaining growing bacteria. Nature 544, 498–502 (2017).
Geva-Zatorsky, N. et al. Mining the human gut microbiota for immunomodulatory organisms. Cell 168, 928–943.e911 (2017). In this study, a systematic approach using both immunological phenotyping and transcriptional profiling was used to define the effects of 53 human-gut commensal bacteria on a wide range of gut immune responses.
Viladomiu, M. et al. IgA-coated E. coli enriched in Crohn's disease spondyloarthritis promote TH17-dependent inflammation. Sci. Transl. Med. 9, eaaf9655 (2017). Using IgA-seq to provide insight into microbiota that might have systemic inflammatory effects, this study analyzed samples from people with Crohn's disease–associated spondyloarthritis and has identified the ability of adherent-invasive E. coli to induce inflammatory T H 17 cells.
Tan, T.G. et al. Identifying species of symbiont bacteria from the human gut that, alone, can induce intestinal Th17 cells in mice. Proc. Natl. Acad. Sci. USA 113, E8141–E8150 (2016). This study used a gnotobiotic mouse platform to screen 39 human-gut symbionts and has identified the ubiquitous symbiont Bifidobacteria adolescentis as a notable inducer of T H 17 cells.
Atarashi, K. et al. Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500, 232–236 (2013).
Fung, T.C. et al. Lymphoid-tissue-resident commensal bacteria promote members of the IL-10 cytokine family to establish mutualism. Immunity 44, 634–646 (2016).
Kunisawa, J. & Kiyono, H. Alcaligenes is commensal bacteria habituating in the gut-associated lymphoid tissue for the regulation of intestinal IgA responses. Front. Immunol. 3, 65 (2012).
Obata, T. et al. Indigenous opportunistic bacteria inhabit mammalian gut-associated lymphoid tissues and share a mucosal antibody-mediated symbiosis. Proc. Natl. Acad. Sci. USA 107, 7419–7424 (2010).
Sonnenberg, G.F. & Artis, D. Innate lymphoid cells in the initiation, regulation and resolution of inflammation. Nat. Med. 21, 698–708 (2015).
Sato, S. et al. Transcription factor Spi-B-dependent and -independent pathways for the development of Peyer's patch M cells. Mucosal Immunol. 6, 838–846 (2013).
Satoh-Takayama, N. et al. The chemokine receptor CXCR6 controls the functional topography of interleukin-22 producing intestinal innate lymphoid cells. Immunity 41, 776–788 (2014).
Stockinger, B., Di Meglio, P., Gialitakis, M. & Duarte, J.H. The aryl hydrocarbon receptor: multitasking in the immune system. Annu. Rev. Immunol. 32, 403–432 (2014).
Lindemans, C.A. et al. Interleukin-22 promotes intestinal-stem-cell-mediated epithelial regeneration. Nature 528, 560–564 (2015).
Hallen-Adams, H.E. & Suhr, M.J. Fungi in the healthy human gastrointestinal tract. Virulence 8, 352–358 (2016).
Liguori, G. et al. Fungal dysbiosis in mucosa-associated microbiota of Crohn's disease patients. J. Crohns Colitis 10, 296–305 (2016).
Suhr, M.J., Banjara, N. & Hallen-Adams, H.E. Sequence-based methods for detecting and evaluating the human gut mycobiome. Lett. Appl. Microbiol. 62, 209–215 (2016).
Hoarau, G. et al. Bacteriome and mycobiome interactions underscore microbial dysbiosis in familial Crohn's disease. MBio 7, e01250–16 (2016).
Li, Q. et al. Dysbiosis of gut fungal microbiota is associated with mucosal inflammation in Crohn's disease. J. Clin. Gastroenterol. 48, 513–523 (2014).
Sokol, H. et al. Fungal microbiota dysbiosis in IBD. Gut 66, 1039–1048 (2016).
Lamas, B. et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 22, 598–605 (2016).
Tang, C. et al. Inhibition of Dectin-1 signaling ameliorates colitis by inducing Lactobacillus-mediated regulatory T cell expansion in the intestine. Cell Host Microbe 18, 183–197 (2015).
Lewis, J.D. et al. Inflammation, antibiotics, and diet as environmental stressors of the gut microbiome in pediatric Crohn's disease. Cell Host Microbe 18, 489–500 (2015). This study shows that inflammation, antibiotics and diet independently affect the gut microbiota in people with Crohn's disease and provides evidence of an association between antibiotic use and fungal overgrowth.
Wheeler, M.L. et al. Immunological consequences of intestinal fungal dysbiosis. Cell Host Microbe 19, 865–873 (2016). This study shows that targeted fungal-community dysbiosis has local and systemic effects on immunity and inflammation.
Fan, D. et al. Activation of HIF-1α and LL-37 by commensal bacteria inhibits Candida albicans colonization. Nat. Med. 21, 808–814 (2015). This study shows that commensal bacteria can promote resistance to C. albicans colonization by increasing the H1F-1α-mediated expression of the antimicrobial peptide LL-37.
Chudnovskiy, A. et al. Host-protozoan interactions protect from mucosal infections through activation of the inflammasome. Cell 167, 444–456.e14 (2016).
Escalante, N.K. et al. The common mouse protozoa Tritrichomonas muris alters mucosal T cell homeostasis and colitis susceptibility. J. Exp. Med. 213, 2841–2850 (2016).
Kernbauer, E., Ding, Y. & Cadwell, K. An enteric virus can replace the beneficial function of commensal bacteria. Nature 516, 94–98 (2014). This paper demonstrated that colonization with a single symbiotic eukaryotic virus can reverse some of the physiological and immunological defects observed in germ-free or antibiotic-exposed mice.
Kuss, S.K. et al. Intestinal microbiota promote enteric virus replication and systemic pathogenesis. Science 334, 249–252 (2011).
Kane, M. et al. Successful transmission of a retrovirus depends on the commensal microbiota. Science 334, 245–249 (2011).
Monaco, C.L. et al. Altered virome and bacterial microbiome in human immunodeficiency virus-associated acquired immunodeficiency syndrome. Cell Host Microbe 19, 311–322 (2016).
Norman, J.M. et al. Disease-specific alterations in the enteric virome in inflammatory bowel disease. Cell 160, 447–460 (2015).
Handley, S.A. et al. Pathogenic simian immunodeficiency virus infection is associated with expansion of the enteric virome. Cell 151, 253–266 (2012).
Ramanan, D. et al. Helminth infection promotes colonization resistance via type 2 immunity. Science 352, 608–612 (2016).
Osborne, L.C. et al. Coinfection. Virus-helminth coinfection reveals a microbiota-independent mechanism of immunomodulation. Science 345, 578–582 (2014).
Reese, T.A. et al. Helminth infection reactivates latent γ-herpesvirus via cytokine competition at a viral promoter. Science 345, 573–577 (2014).
Wu, G.D. et al. Comparative metabolomics in vegans and omnivores reveal constraints on diet-dependent gut microbiota metabolite production. Gut 65, 63–72 (2016).
Desai, M.S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353.e21 (2016).
Corrêa-Oliveira, R., Fachi, J.L., Vieira, A., Sato, F.T. & Vinolo, M.A. Regulation of immune cell function by short-chain fatty acids. Clin. Transl. Immunology 5, e73 (2016).
Kaiko, G.E. et al. The colonic crypt protects stem cells from microbiota-derived metabolites. Cell 165, 1708–1720 (2016).
Kelly, C.J. et al. Crosstalk between microbiota-derived short-chain fatty acids and intestinal epithelial HIF augments tissue barrier function. Cell Host Microbe 17, 662–671 (2015).
Kibe, R. et al. Upregulation of colonic luminal polyamines produced by intestinal microbiota delays senescence in mice. Sci. Rep. 4, 4548 (2014).
Levy, M. et al. Microbiota-modulated metabolites shape the intestinal microenvironment by regulating NLRP6 inflammasome signaling. Cell 163, 1428–1443 (2015).
Zelante, T. et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 39, 372–385 (2013).
Schiering, C. et al. Feedback control of AHR signalling regulates intestinal immunity. Nature 542, 242–245 (2017).
Zhu, W. et al. Gut microbial metabolite TMAO enhances platelet hyperreactivity and thrombosis risk. Cell 165, 111–124 (2016).
Wang, Z. et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–63 (2011).
Koeth, R.A. et al. Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat. Med. 19, 576–585 (2013).
Dumas, M.E. et al. Metabolic profiling reveals a contribution of gut microbiota to fatty liver phenotype in insulin-resistant mice. Proc. Natl. Acad. Sci. USA 103, 12511–12516 (2006).
Levin, B.J. et al. A prominent glycyl radical enzyme in human gut microbiomes metabolizes trans-4-hydroxy-l-proline. Science 355, eaai8386 (2017). This study describes a novel chemically guided functional profiling–coupled protein sequence-similarity network with quantitative metagenomics analysis, which enabled the discovery and functional characterization of the GRE superfamily in the microbiome.
Donia, M.S. et al. A systematic analysis of biosynthetic gene clusters in the human microbiome reveals a common family of antibiotics. Cell 158, 1402–1414 (2014).
Guo, C.J. et al. Discovery of reactive microbiota-derived metabolites that inhibit host proteases. Cell 168, 517–526.e18 (2017).
Manoury, B. Proteases: essential actors in processing antigens and intracellular toll-like receptors. Front. Immunol. 4, 299 (2013).
Kim, Y.G. et al. Gut dysbiosis promotes M2 macrophage polarization and allergic airway inflammation via fungi-induced PGE2 . Cell Host Microbe 15, 95–102 (2014).
Vatanen, T. et al. Variation in microbiome LPS immunogenicity contributes to autoimmunity in humans. Cell 165, 842–853 (2016)Bacteroides species in the microbiota of children from Finland and Estonia with high susceptibility to autoimmunity produce a type of LPS that inhibits innate immune signaling and endotoxin tolerance. These properties may interfere with early immunological education and contribute to the development of type 1 diabetes.
Bach, J.F. & Chatenoud, L. The hygiene hypothesis: an explanation for the increased frequency of insulin-dependent diabetes. Cold Spring Harb. Perspect. Med. 2, a007799 (2012).
von Mutius, E. & Vercelli, D. Farm living: effects on childhood asthma and allergy. Nat. Rev. Immunol. 10, 861–868 (2010).
Calderon-Gomez, E. et al. Commensal-specific CD4+ cells from patients with Crohn′s disease have a T-helper 17 inflammatory profile. Gastroenterology 151, 489–500 e3 (2016).
Campisi, L. et al. Apoptosis in response to microbial infection induces autoreactive TH17 cells. Nat. Immunol. 17, 1084–1092 (2016).
Hand, T.W. et al. Acute gastrointestinal infection induces long-lived microbiota-specific T cell responses. Science 337, 1553–1556 (2012).
Blander, J.M., Torchinsky, M.B. & Campisi, L. Revisiting the old link between infection and autoimmune disease with commensals and T helper 17 cells. Immunol. Res. 54, 50–68 (2012).
Yang, Y. et al. Focused specificity of intestinal TH17 cells towards commensal bacterial antigens. Nature 510, 152–156 (2014).
Wu, H.J. et al. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity 32, 815–827 (2010).
Lee, Y.K., Menezes, J.S., Umesaki, Y. & Mazmanian, S.K. Proinflammatory T-cell responses to gut microbiota promote experimental autoimmune encephalomyelitis. Proc. Natl. Acad. Sci. USA 108 (Suppl. 1), 4615–4622 (2011).
Asquith, M.J. et al. Perturbed mucosal immunity and dysbiosis accompany clinical disease in a rat model of spondyloarthritis. Arthritis Rheumatol. 68, 2151–2162 (2016).
Ciccia, F. et al. Dysbiosis and zonulin upregulation alter gut epithelial and vascular barriers in patients with ankylosing spondylitis. Ann. Rheum. Dis. 76, 1123–1132 (2017).
Teng, F. et al. Gut microbiota drive autoimmune arthritis by promoting differentiation and migration of Peyer's patch T follicular helper cells. Immunity 44, 875–888 (2016).
Cryan, J.F. & Dinan, T.G. Mind-altering microorganisms: the impact of the gut microbiota on brain and behaviour. Nat. Rev. Neurosci. 13, 701–712 (2012).
Sharon, G., Sampson, T.R., Geschwind, D.H. & Mazmanian, S.K. The central nervous system and the gut microbiome. Cell 167, 915–932 (2016).
Gabanyi, I. et al. Neuro-immune interactions drive tissue programming in intestinal macrophages. Cell 164, 378–391 (2016).
Rieder, R., Wisniewski, P.J., Alderman, B.L. & Campbell, S.C. Microbes and mental health: a review. Brain Behav. Immun. http://dx.doi.org/10.1016/j.bbi.2017.01.016 (2017).
Yano, J.M. et al. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell 161, 264–276 (2015)Indigenous spore-forming microbes from the gut microbiota produce metabolites that promote host serotonin biosynthesis in the gastrointestinal tract and affect gastrointestinal motility and hemostasis.
Reigstad, C.S. et al. Gut microbes promote colonic serotonin production through an effect of short-chain fatty acids on enterochromaffin cells. FASEB J. 29, 1395–1403 (2015).
O'Mahony, S.M., Clarke, G., Borre, Y.E., Dinan, T.G. & Cryan, J.F. Serotonin, tryptophan metabolism and the brain-gut-microbiome axis. Behav. Brain Res. 277, 32–48 (2015).
Erny, D. et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat. Neurosci. 18, 965–977 (2015).
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 (2016).
Sampson, T.R. et al. Gut microbiota regulate motor deficits and neuroinflammation in a model of Parkinson′s disease. Cell 167, 1469–1480.e12 (2016). SCFAs from gut microbes modulate microglia, are required for neuroinflammatory responses. They are also required for the hallmark α-synuclein-dependent motor and gastrointestinal deficits and brain pathology in a model of Parkinson's disease. The microbiota from people with Parkinson's disease induces motor dysfunction in this model.
Rothhammer, V. et al. Type I interferons and microbial metabolites of tryptophan modulate astrocyte activity and central nervous system inflammation via the aryl hydrocarbon receptor. Nat. Med. 22, 586–597 (2016).
Schirmer, M. et al. Linking the human gut microbiome to inflammatory cytokine production capacity. Cell 167, 1125–1136.e8 (2016)This study investigates how differences in the microbiome contribute to variations in the human inflammatory response and demonstrates that TNF and IFNγ responses are associated with microbial palmitoleic acid and tryptophan metabolism. This study also provides a database for microbial mediators that influence human cytokine responses.
Buffie, C.G. et al. Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile. Nature 517, 205–208 (2015).
Rangan, K.J. et al. A secreted bacterial peptidoglycan hydrolase enhances tolerance to enteric pathogens. Science 353, 1434–1437 (2016).
Sansone, C.L. et al. Microbiota-dependent priming of antiviral intestinal immunity in Drosophila. Cell Host Microbe 18, 571–581 (2015).
Khosravi, A. et al. Gut microbiota promote hematopoiesis to control bacterial infection. Cell Host Microbe 15, 374–381 (2014).
Ichinohe, T. et al. Microbiota regulates immune defense against respiratory tract influenza A virus infection. Proc. Natl. Acad. Sci. USA 108, 5354–5359 (2011).
Clarke, T.B. et al. Recognition of peptidoglycan from the microbiota by Nod1 enhances systemic innate immunity. Nat. Med. 16, 228–231 (2010).
Pamer, E.G. Resurrecting the intestinal microbiota to combat antibiotic-resistant pathogens. Science 352, 535–538 (2016).
Soares, M.P., Teixeira, L. & Moita, L.F. Disease tolerance and immunity in host protection against infection. Nat. Rev. Immunol. 17, 83–96 (2017).
Schieber, A.M. et al. Disease tolerance mediated by microbiome E. coli involves inflammasome and IGF-1 signaling. Science 350, 558–563 (2015). This study elegantly demonstrates that a strain of E. coli naturally colonizing the intestine in mice is sufficient to prevent wasting after infections, owing to the sustained inflammasome-dependent activation of the IGF1–PI3K–AKT pathway in skeletal muscle.
Pickard, J.M. et al. Rapid fucosylation of intestinal epithelium sustains host-commensal symbiosis in sickness. Nature 514, 638–641 (2014).
Reichardt, N. et al. Phylogenetic distribution of three pathways for propionate production within the human gut microbiota. ISME J. 8, 1323–1335 (2014).
Fonseca, D.M. et al. Microbiota-dependent sequelae of acute infection compromise tissue-specific immunity. Cell 163, 354–366 (2015).
Kim, D. et al. Nod2-mediated recognition of the microbiota is critical for mucosal adjuvant activity of cholera toxin. Nat. Med. 22, 524–530 (2016).
Ruane, D. et al. Microbiota regulate the ability of lung dendritic cells to induce IgA class-switch recombination and generate protective gastrointestinal immune responses. J. Exp. Med. 213, 53–73 (2016).
Oh, J.Z. et al. TLR5-mediated sensing of gut microbiota is necessary for antibody responses to seasonal influenza vaccination. Immunity 41, 478–492 (2014).
Nakaya, H.I. et al. Systems biology of vaccination for seasonal influenza in humans. Nat. Immunol. 12, 786–795 (2011).
Kollmann, T.R., Kampmann, B., Mazmanian, S.K., Marchant, A. & Levy, O. Protecting the newborn and young infant from infectious diseases: lessons from immune ontogeny. Immunity 46, 350–363 (2017).
Valdez, Y., Brown, E.M. & Finlay, B.B. Influence of the microbiota on vaccine effectiveness. Trends Immunol. 35, 526–537 (2014).
Grivennikov, S.I. et al. Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature 491, 254–258 (2012).
Huber, S. et al. IL-22BP is regulated by the inflammasome and modulates tumorigenesis in the intestine. Nature 491, 259–263 (2012).
Ekbom, A., Helmick, C., Zack, M. & Adami, H.O. Ulcerative colitis and colorectal cancer: apopulation-based study. N. Engl. J. Med. 323, 1228–1233 (1990).
Arthur, J.C. et al. Intestinal inflammation targets cancer-inducing activity of the microbiota. Science 338, 120–123 (2012).
Kostic, A.D. et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 14, 207–215 (2013).
Wu, S. et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat. Med. 15, 1016–1022 (2009).
Pitt, J.M. et al. Fine-tuning cancer immunotherapy: optimizing the gut microbiome. Cancer Res. 76, 4602–4607 (2016).
Vétizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015).
Sivan, A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089 (2015). Refs. 132 and 133 show that the intestinal microbiota affects the outcome of checkpoint-blockade-based cancer immunotherapy.
Acknowledgements
The authors thank all their past and present laboratory members for their contributions. We thank our funding agencies for their support to our laboratories: NIH grants DK072201, DK111862, AI073899, AI123284 and AI127658, the Searle Scholars Program, the Burroughs Wellcome Fund, the American Cancer Society and the Leukemia & Lymphoma Society to J.M.B.; NIH grant DK099381, the Crohn's and Colitis Foundation Senior Research Award 346814 and the Charina Foundation to R.S.L.; NIH grants DK098310 and AI123819, and Kenneth Rainin Foundation Innovator and Breakthrough awards to I.D.I.; NIH grants DP5OD012116, AI123368 and DK110262, and the Crohn's and Colitis Foundation, the Searle Scholars Program and the American Asthma Foundation Scholar Award to G.F.S.; NIH grants AI061570, AI087990, AI074878, AI083480, AI095466, AI095608, AI102942 and AI097333, Burroughs Wellcome Fund and the Crohn's & Colitis Foundation of America to D.A.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Blander, J., Longman, R., Iliev, I. et al. Regulation of inflammation by microbiota interactions with the host. Nat Immunol 18, 851–860 (2017). https://doi.org/10.1038/ni.3780
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3780
This article is cited by
-
The gut-liver axis in hepatobiliary diseases
Inflammation and Regeneration (2024)
-
Gut microbial metabolite facilitates colorectal cancer development via ferroptosis inhibition
Nature Cell Biology (2024)
-
An immune-competent human gut microphysiological system enables inflammation-modulation by Faecalibacterium prausnitzii
npj Biofilms and Microbiomes (2024)
-
Dietary non-starch polysaccharides impair immunity to enteric nematode infection
BMC Biology (2023)
-
A diet rich in fermentable fiber promotes robust changes in the intestinal microbiota, mitigates intestinal permeability, and attenuates autoimmune uveitis
Scientific Reports (2023)