Abstract
The aggregation of hypertrophic macrophages constitutes the basis of all granulomatous diseases, such as tuberculosis or sarcoidosis, and is decisive for disease pathogenesis. However, macrophage-intrinsic pathways driving granuloma initiation and maintenance remain elusive. We found that activation of the metabolic checkpoint kinase mTORC1 in macrophages by deletion of the gene encoding tuberous sclerosis 2 (Tsc2) was sufficient to induce hypertrophy and proliferation, resulting in excessive granuloma formation in vivo. TSC2-deficient macrophages formed mTORC1-dependent granulomatous structures in vitro and showed constitutive proliferation that was mediated by the neo-expression of cyclin-dependent kinase 4 (CDK4). Moreover, mTORC1 promoted metabolic reprogramming via CDK4 toward increased glycolysis while simultaneously inhibiting NF-κB signaling and apoptosis. Inhibition of mTORC1 induced apoptosis and completely resolved granulomas in myeloid TSC2-deficient mice. In human sarcoidosis patients, mTORC1 activation, macrophage proliferation and glycolysis were identified as hallmarks that correlated with clinical disease progression. Collectively, TSC2 maintains macrophage quiescence and prevents mTORC1-dependent granulomatous disease with clinical implications for sarcoidosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Cambier, C.J., Falkow, S. & Ramakrishnan, L. Host evasion and exploitation schemes of Mycobacterium tuberculosis. Cell 159, 1497–1509 (2014).
Russell, D.G., Cardona, P.J., Kim, M.J., Allain, S. & Altare, F. Foamy macrophages and the progression of the human tuberculosis granuloma. Nat. Immunol. 10, 943–948 (2009).
Orme, I.M. & Basaraba, R.J. The formation of the granuloma in tuberculosis infection. Semin. Immunol. 26, 601–609 (2014).
Hams, E., Aviello, G. & Fallon, P.G. The schistosoma granuloma: friend or foe? Front. Immunol. 4, 89 (2013).
Iannuzzi, M.C. & Fontana, J.R. Sarcoidosis: clinical presentation, immunopathogenesis, and therapeutics. J. Am. Med. Assoc. 305, 391–399 (2011).
Petersen, H.J. & Smith, A.M. The role of the innate immune system in granulomatous disorders. Front. Immunol. 4, 120 (2013).
Beegle, S.H., Barba, K., Gobunsuy, R. & Judson, M.A. Current and emerging pharmacological treatments for sarcoidosis: a review. Drug Des. Devel. Ther. 7, 325–338 (2013).
Zissel, G. & Müller-Quernheim, J. Cellular players in the immunopathogenesis of sarcoidosis. Clin. Chest Med. 36, 549–560 (2015).
Lockstone, H.E. et al. Gene set analysis of lung samples provides insight into pathogenesis of progressive, fibrotic pulmonary sarcoidosis. Am. J. Respir. Crit. Care Med. 181, 1367–1375 (2010).
Ehlers, S. & Schaible, U.E. The granuloma in tuberculosis: dynamics of a host-pathogen collusion. Front. Immunol. 3, 411 (2013).
Ramakrishnan, L. Revisiting the role of the granuloma in tuberculosis. Nat. Rev. Immunol. 12, 352–366 (2012).
Orme, I.M., Robinson, R.T. & Cooper, A.M. The balance between protective and pathogenic immune responses in the TB-infected lung. Nat. Immunol. 16, 57–63 (2015).
Ehlers, S., Kutsch, S., Ehlers, E.M., Benini, J. & Pfeffer, K. Lethal granuloma disintegration in mycobacteria-infected TNFRp55-/- mice is dependent on T cells and IL-12. J. Immunol. 165, 483–492 (2000).
North, R.J. & Jung, Y.J. Immunity to tuberculosis. Annu. Rev. Immunol. 22, 599–623 (2004).
Regev, D. et al. Heme oxygenase-1 promotes granuloma development and protects against dissemination of mycobacteria. Lab. Invest. 92, 1541–1552 (2012).
Dorhoi, A. & Kaufmann, S.H. Pathology and immune reactivity: understanding multidimensionality in pulmonary tuberculosis. Semin. Immunopathol. 38, 153–166 (2016).
Laplante, M. & Sabatini, D.M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Inoki, K., Li, Y., Zhu, T., Wu, J. & Guan, K.L. TSC2 is phosphorylated and inhibited by Akt and suppresses mTOR signalling. Nat. Cell Biol. 4, 648–657 (2002).
Chi, H. Regulation and function of mTOR signalling in T cell fate decisions. Nat. Rev. Immunol. 12, 325–338 (2012).
Wang, Y. et al. Tuberous sclerosis 1 (Tsc1)-dependent metabolic checkpoint controls development of dendritic cells. Proc. Natl. Acad. Sci. USA 110, E4894–E4903 (2013).
Weichhart, T., Hengstschläger, M. & Linke, M. Regulation of innate immune cell function by mTOR. Nat. Rev. Immunol. 15, 599–614 (2015).
Byles, V. et al. The TSC-mTOR pathway regulates macrophage polarization. Nat. Commun. 4, 2834 (2013).
Pan, H., O'Brien, T.F., Zhang, P. & Zhong, X.P. The role of tuberous sclerosis complex 1 in regulating innate immunity. J. Immunol. 188, 3658–3666 (2012).
Fang, C. et al. Tsc1 is a Critical Regulator of Macrophage Survival and Function. Cell. Physiol. Biochem. 36, 1406–1418 (2015).
Hernandez, O., Way, S., McKenna, J. III & Gambello, M.J. Generation of a conditional disruption of the Tsc2 gene. Genesis 45, 101–106 (2007).
Prokop, S., Heppner, F.L., Goebel, H.H. & Stenzel, W. M2 polarized macrophages and giant cells contribute to myofibrosis in neuromuscular sarcoidosis. Am. J. Pathol. 178, 1279–1286 (2011).
Misharin, A.V., Morales-Nebreda, L., Mutlu, G.M., Budinger, G.R. & Perlman, H. Flow cytometric analysis of macrophages and dendritic cell subsets in the mouse lung. Am. J. Respir. Cell Mol. Biol. 49, 503–510 (2013).
Wong, D.J. et al. Module map of stem cell genes guides creation of epithelial cancer stem cells. Cell Stem Cell 2, 333–344 (2008).
Bertoli, C., Skotheim, J.M. & de Bruin, R.A. Control of cell cycle transcription during G1 and S phases. Nat. Rev. Mol. Cell Biol. 14, 518–528 (2013).
Hochegger, H., Takeda, S. & Hunt, T. Cyclin-dependent kinases and cell-cycle transitions: does one fit all? Nat. Rev. Mol. Cell Biol. 9, 910–916 (2008).
Sherr, C.J., Beach, D. & Shapiro, G.I. Targeting CDK4 and CDK6: from discovery to tTherapy. Cancer Discov. 6, 353–367 (2016).
Howell, J.J., Ricoult, S.J., Ben-Sahra, I. & Manning, B.D. A growing role for mTOR in promoting anabolic metabolism. Biochem. Soc. Trans. 41, 906–912 (2013).
Lebwohl, D. et al. Development of everolimus, a novel oral mTOR inhibitor, across a spectrum of diseases. Ann. NY Acad. Sci. 1291, 14–32 (2013).
Pforte, A. et al. Proliferating alveolar macrophages in BAL and lung function changes in interstitial lung disease. Eur. Respir. J. 6, 951–955 (1993).
Petzmann, S. et al. Enhanced proliferation and decreased apoptosis in lung lavage cells of sarcoidosis patients. Sarcoidosis Vasc. Diffuse Lung Dis. 23, 190–200 (2006).
Lan, H.Y., Nikolic-Paterson, D.J., Mu, W. & Atkins, R.C. Local macrophage proliferation in multinucleated giant cell and granuloma formation in experimental Goodpasture's syndrome. Am. J. Pathol. 147, 1214–1220 (1995).
Huaux, F. et al. IL-1α induces CD11b(low) alveolar macrophage proliferation and maturation during granuloma formation. J. Pathol. 235, 698–709 (2015).
Sieweke, M.H. & Allen, J.E. Beyond stem cells: self-renewal of differentiated macrophages. Science 342, 1242974 (2013).
Robbins, C.S. et al. Local proliferation dominates lesional macrophage accumulation in atherosclerosis. Nat. Med. 19, 1166–1172 (2013).
Haase, J. et al. Local proliferation of macrophages in adipose tissue during obesity-induced inflammation. Diabetologia 57, 562–571 (2014).
Fingar, D.C. & Blenis, J. Target of rapamycin (TOR): an integrator of nutrient and growth factor signals and coordinator of cell growth and cell cycle progression. Oncogene 23, 3151–3171 (2004).
Song, J., Salek-Ardakani, S., So, T. & Croft, M. The kinases aurora B and mTOR regulate the G1-S cell cycle progression of T lymphocytes. Nat. Immunol. 8, 64–73 (2007).
Miliani de Marval, P.L. et al. Transgenic expression of cyclin-dependent kinase 4 results in epidermal hyperplasia, hypertrophy, and severe dermal fibrosis. Am. J. Pathol. 159, 369–379 (2001).
Pearce, E.L. & Pearce, E.J. Metabolic pathways in immune cell activation and quiescence. Immunity 38, 633–643 (2013).
Rodríguez-Prados, J.C. et al. Substrate fate in activated macrophages: a comparison between innate, classic, and alternative activation. J. Immunol. 185, 605–614 (2010).
Hofmann, S. et al. Genome-wide association analysis reveals 12q13.3-q14.1 as new risk locus for sarcoidosis. Eur. Respir. J. 41, 888–900 (2013).
Teirstein, A.S. et al. Results of 188 whole-body fluorodeoxyglucose positron emission tomography scans in 137 patients with sarcoidosis. Chest 132, 1949–1953 (2007).
Judson, M.A. & Baughman, R.P. Worsening of pulmonary sarcoidosis. Curr. Opin. Pulm. Med. 20, 508–516 (2014).
Manzia, T.M. et al. Successful treatment of systemic de novo sarcoidosis with cyclosporine discontinuation and provision of rapamune after liver transplantation. Transpl. Int. 24, e69–e70 (2011).
Rosner, M. et al. Efficient siRNA-mediated prolonged gene silencing in human amniotic fluid stem cells. Nat. Protoc. 5, 1081–1095 (2010).
Van Noorden, C.J.F. & Frederiks, W.M. Enzyme Histochemistry: a Laboratory Manual of Current Methods (Oxford University Press, 1992).
Acknowledgements
We thank U. Reichart and T. Kolbe for support in mouse breeding. We are grateful to D. Georg for the possibility of using the XYLON Maxishot. We would also like to thank M. Stadler and H. Dolznig (Medical University of Vienna) for providing human-monocyte-derived macrophages. Tsc1+/+ MEFs were a kind gift of D.J. Kwiatkowski (Harvard Medical School). T.W. is supported by grants from the Austrian Science Fund (FWF) grant FWF-P27701-B20, the Else-Kröner-Fresenius-Stiftung (P2013_A149), and the Herzfelder'sche Familienstiftung. M.L. is supported by the [DOC] Doctoral Fellowship Programme of the Austrian Academy of Sciences. V.S. is funded by FWF SFB F28 and SBF F47. M. Müller is funded by FWF SFB F28. M. Mikula is supported by the FWF grant P25336-B13.
Author information
Authors and Affiliations
Contributions
The study was conceived by M.L. and T.W. The research was carried out by M.L., H.T.T.P., K.K., T.S., A.M., F.D., B.S., M.R., B.K., N.S., B.N., P.K. and T.W. Resources were provided by S.B., V.S., M. Mikula, M. Müller, W.W., A.H., M.S., M.H., M.J.G. and T.W. T.W. wrote the original draft of the manuscript. All of the authors reviewed and edited the manuscript. The study was supervised by T.W.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Phenotypic evaluation of Tsc2fl/fl,Lyz2-Cre mice.
(a) Mac-2 and p-S6 immunofluorescence of liver sections of 3 month-old Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre mice. Scale bar, 100 μm. (b) Lung section of a 3 month-old Tsc2fl/fl,Lyz2-Cre mouse evaluated by H&E staining. Scale bar, 50 μm. (c) Mediastinal and superficial cervical lymph nodes (LN) of 3 month-old mice were evaluated by H&E staining or by immunohistochemistry with Mac-2. Black scale bar, 200 μm, white scale bar 25 μm. (d) Lung sections of Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre mice were stained with immunohistochemistry for myeloperoxidase (MPO). Scale bar, 25 μm. Data are representative of three mice (a,c,d) or five mice (b) per genotype.
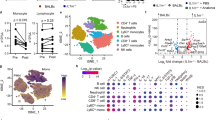
Supplementary Figure 2 Identification and characterization of the hypertrophic macrophages in the lungs of Tsc2fl/fl,Lyz2-Cre mice.
(a) Flow cytometry plots of lung single cell suspensions at 10 weeks. The hypertrophic macrophages are indicated as “hypertroph. MΦ”. (b) Flow cytometric analysis of the indicated surface markers of alveolar macrophages, interstitial macrophages, and Ly6Chi monocytes in Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre mice and their comparison to the hypertrophic macrophages in the lung of the Tsc2fl/fl,Lyz2-Cre mice. (c) Flow cytometric analysis of immune cell populations in the lung of 10 week-old Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre mice. Shown are means ± s.d. (d) Bone marrow-derived macrophages were unstimulated or stimulated with either 100 ng/ml LPS and 20 ng/ml IFNγ or with 10 ng/ml IL-4 for 48h. Expression of the indicated mRNAs normalized to β-actin is shown as means ± s.e.m. Data are representative of five mice (a,b) per genotype or cumulative of five (c,d) individual mice per genotype.
Supplementary Figure 3 Granulomatous aggregation in vitro is specific for TSC2-deficient macrophages.
(a) p-S6 immunofluorescence of growth factor-deprived Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre BMDM on day 7. Scale bar, 10 μm. (b) BMDM were deprived of growth factor overnight and re-stimulated with 10 ng/ml CSF1 or 100 ng/ml LPS for the indicated time points. Whole cell lysates were analyzed by immunoblotting. (c) Representative images of IL-4 stimulated (10 ng/ml) BMDM treated with solvent or 100 nM rapamycin for four days. Scale bar, 100 μm. (d) Cell volume of BMDM after differentiation. Shown are means ± s.e.m. (Tsc2fl/fl n= 4, Tsc2fl/fl,Lyz2-Cre n= 5). (e) Images of Tsc2fl/fl,Lyz2-Cre BMDM that were treated with solvent or 100 nM rapamycin for 24 h. Scale bar, 100 μm. (f) Images of lung single cell suspensions that were cultivated in CSF1 containing media for 7 days. Treatment with solvent or 100 nM rapamycin lasted for 48h. Scale bar, 100 μm. (g) Images of Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre dendritic cells (DC) differentiated for 6 six days that were cultured for additional 6 days in the presence of 20 ng/ml CSF2. Scale bar, 100 μm. Right panel, whole cell lysates of Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre DCs were analyzed by immunoblotting with the indicated antibodies. (h) Representative images of primary human foreskin and lung fibroblasts, as well as mouse embryonic fibroblasts (MEF) transfected with Tsc2 or non-target siRNAs or left untreated as indicated. Scale bar, 200 μm. Right panel, Knockdown efficiency was confirmed by immunoblotting using the indicated antibodies. Results are representative of three (a,c,e,f), four (g), or two (b) mice per genotype or cumulative for four to five (d) mice per genotype or for 2 independent experiments (lung and foreskin) or one experiment (h).
Supplementary Figure 4 Regulation of proliferation, self-renewal, and apoptosis by TSC2 in macrophages.
(a) Images of Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre BMDM on day 7. Scale bar, 100 μm. (b) Cell cycle analysis of growth-factor deprived Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre BMDM on day 7. Shown are means ± s.e.m. (n= 3). (c) IHC for Ki-67 in a paw section of a 6 month-old Tsc2fl/fl,Lyz2-Cre mice. Scale bar, 200 μm. (d) Cell cycle analysis of CSF1-stimulated BMDM treated with solvent or 100 nM rapamycin for 18 hours. Shown are means ± s.e.m. (n= 3). (e) Whole cell lysates of Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre BMDM were analyzed by immunoblotting with the indicated antibodies. (f) GSEA of the 'Self-renewal' gene signature (WONG_EMBRYONIC_STEM_CELL_CORE) in Tsc2fl/fl,Lyz2-Cre BMDM relative to Tsc2fl/fl BMDM. Data are from one experiment with four biological replicates per genotype. (g) IHC for survivin in lung sections of 3 month-old Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre mice. Scale bar, 100 μm. Results are representative of three mice per genotype (a,c,g), cumulative for three (b,d) mice per genotype, or are from one experiment with two (e) or four biological replicates per genotype (f).
Supplementary Figure 5 TSC2-deficiency promotes CDK4-dependent proliferation in macrophages.
(a) BMDM were treated with 100 nM rapamycin or solvent control for 18 h. Whole cell lysates were analyzed by immunoblotting with the indicated antibodies. (b) Freshly differentiated C57BL/6J BMDM were starved from CSF1 for 0, 4 and 12 h. Cells that were deprived from CSF1 for 12 h were then restimulated with 40 ng/ml CSF1 for 12 or 24h. Whole cell lysates of the time points were analyzed by immunoblotting. (c) C57BL/6J BMDM were stimulated with 10, 20, 30 and 40 ng/ml CSF1 for 24 h. Whole cell lysates were analyzed by immunoblotting. (d) C57BL/6J BMDM were deprived of CSF1 for 12 h. Afterwards, they were either harvested or treated with 250 nM Torin1 or solvent control, and stimulated with 20 ng/ml CSF1 for 24h. Whole cell lysates of the time points were analyzed by immunoblotting. (e) Representative IHC of CDK4 in liver sections of 3 month-old Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre mice. Scale bar, 100 μm. (f) Human monocyte-derived macrophages were treated with 100 nM rapamycin or 250 nM Torin in the presence of 20 ng/ml CSF1. Whole cell lysates were analyzed by immunoblotting. (g) Analysis of CSF1-induced proliferation of Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre BMDM treated with solvent or the indicated amounts of SC-203873. Shown are means ± s.e.m. (n= 4). (h) BMDM were treated with 1 μM PD-0332991 or solvent control and stimulated with 10 ng/ml CSF1 for 18 h. Whole cell lysates of the time points were analyzed by immunoblotting. Results are representative of two (a-f,h) or cumulative for two (g) individual experiments.
Supplementary Figure 6 Deletion of TSC2 in macrophages promotes glycolysis and mitochondrial functions.
(a) Extracellular acidification rate (ECAR) and oxygen consumption rate (OCR) of Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre BMDM stimulated with 100 ng/ml LPS. Depicted time course was normalized to unstimulated samples (n=10/group). (b) Total amount of glucose in Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre BMDM in nmol/106 cells on day 7. Shown are boxplots with mean (n=4). (c) A mito stress-test was performed with Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre BMDM on the Seahorse XFe 24 analyzer. Concentrations were 0.8 μg/ml for oligomycin, 0.5 μM for FCCP, and 1 μM for rotenone and antimycin A. Shown are means ± s.d.; (n=11/group). Upper panel: absolute levels; lower panel: normalized to baseline recordings before compound injection. (d) Total amount of citric acid in Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre BMDM in nmol/106 cells on day 7. Shown are boxplots with mean (n=4). (e) Images of glyceraldehyde 3-phosphate dehydrogenase (GAPDH) activity on frozen liver sections of Tsc2fl/fl and Tsc2fl/fl,Lyz2-Cre mice in situ. Sections were additionally stained for Mac-2, p-S6, and DAPI. (f) Fraction (in %) of p-S6-positive Mac-2 macrophages in liver of Tsc2fl/fl (n=4) and Tsc2fl/fl,Lyz2-Cre (n=5) mice with high activities of GAPDH. *p < 0.05, ***p<0.001 (Student’s t test). Data are representative of two (c) or one (a,b,d,e,f) experiments. Scale bar, 100 μm.
Supplementary Figure 7 Expression of TSC1 and TSC2 mRNA in sarcoidosis patients.
Expression of TSC1 and TSC2 mRNA in progressive (n=7) relative to self-limiting (n=8) sarcoidosis from the microarray data of Lockstone et al. Shown are means ± s.e.m. *p < 0.05.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 1394 kb)
Supplementary Figure 8
Uncropped gels (PDF 383 kb)
Rights and permissions
About this article
Cite this article
Linke, M., Pham, H., Katholnig, K. et al. Chronic signaling via the metabolic checkpoint kinase mTORC1 induces macrophage granuloma formation and marks sarcoidosis progression. Nat Immunol 18, 293–302 (2017). https://doi.org/10.1038/ni.3655
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3655
This article is cited by
-
Serine metabolism in macrophage polarization
Inflammation Research (2024)
-
Identification of miRNA-mRNA-TF regulatory networks in peripheral blood mononuclear cells of type 1 diabetes
BMC Endocrine Disorders (2022)
-
Loss of myeloid Tsc2 predisposes to angiotensin II-induced aortic aneurysm formation in mice
Cell Death & Disease (2022)
-
Kutane Sarkoidose – eine granulomatöse Modellerkrankung
hautnah (2022)
-
The Evolving Landscape of Cutaneous Sarcoidosis: Pathogenic Insight, Clinical Challenges, and New Frontiers in Therapy
American Journal of Clinical Dermatology (2022)