Abstract
Aire is a transcriptional regulator that induces promiscuous expression of thousands of genes encoding tissue-restricted antigens (TRAs) in medullary thymic epithelial cells (mTECs). While the target genes of Aire are well characterized, the transcriptional programs that regulate its own expression have remained elusive. Here we comprehensively analyzed both cis-acting and trans-acting regulatory mechanisms and found that the Aire locus was insulated by the global chromatin organizer CTCF and was hypermethylated in cells and tissues that did not express Aire. In mTECs, however, Aire expression was facilitated by concurrent eviction of CTCF, specific demethylation of exon 2 and the proximal promoter, and the coordinated action of several transcription activators, including Irf4, Irf8, Tbx21, Tcf7 and Ctcfl, which acted on mTEC-specific accessible regions in the Aire locus.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Nossal, G.J. Cellular mechanisms of immunologic tolerance. Annu. Rev. Immunol. 1, 33–62 (1983).
Klein, L., Kyewski, B., Allen, P.M. & Hogquist, K.A. Positive and negative selection of the T cell repertoire: what thymocytes see (and don't see). Nat. Rev. Immunol. 14, 377–391 (2014).
Aschenbrenner, K. et al. Selection of Foxp3+ regulatory T cells specific for self antigen expressed and presented by Aire+ medullary thymic epithelial cells. Nat. Immunol. 8, 351–358 (2007).
Yang, S., Fujikado, N., Kolodin, D., Benoist, C. & Mathis, D. Immune tolerance. Regulatory T cells generated early in life play a distinct role in maintaining self-tolerance. Science 348, 589–594 (2015).
Anderson, M.S. et al. The cellular mechanism of Aire control of T cell tolerance. Immunity 23, 227–239 (2005).
Derbinski, J., Schulte, A., Kyewski, B. & Klein, L. Promiscuous gene expression in medullary thymic epithelial cells mirrors the peripheral self. Nat. Immunol. 2, 1032–1039 (2001).
Derbinski, J. et al. Promiscuous gene expression in thymic epithelial cells is regulated at multiple levels. J. Exp. Med. 202, 33–45 (2005).
Gavanescu, I., Kessler, B., Ploegh, H., Benoist, C. & Mathis, D. Loss of Aire-dependent thymic expression of a peripheral tissue antigen renders it a target of autoimmunity. Proc. Natl. Acad. Sci. USA 104, 4583–4587 (2007).
DeVoss, J. et al. Spontaneous autoimmunity prevented by thymic expression of a single self-antigen. J. Exp. Med. 203, 2727–2735 (2006).
Anderson, M.S. et al. Projection of an immunological self shadow within the thymus by the aire protein. Science 298, 1395–1401 (2002).
Nagamine, K. et al. Positional cloning of the APECED gene. Nat. Genet. 17, 393–398 (1997).
Sansom, S.N. et al. Population and single-cell genomics reveal the Aire dependency, relief from Polycomb silencing, and distribution of self-antigen expression in thymic epithelia.1918–1931 (2014).
Meredith, M., Zemmour, D., Mathis, D. & Benoist, C. Aire controls gene expression in the thymic epithelium with ordered stochasticity. Nat. Immunol. 16, 942–949 (2015).
Heino, M. et al. RNA and protein expression of the murine autoimmune regulator gene (Aire) in normal, RelB-deficient and in NOD mouse. Eur. J. Immunol. 30, 1884–1893 (2000).
Gardner, J.M. et al. Deletional tolerance mediated by extrathymic Aire-expressing cells. Science 321, 843–847 (2008).
Nishikawa, Y. et al. Biphasic Aire expression in early embryos and in medullary thymic epithelial cells before end-stage terminal differentiation. J. Exp. Med. 207, 963–971 (2010).
Yamano, T. et al. Thymic B cells are licensed to present self antigens for central T cell tolerance induction. Immunity 42, 1048–1061 (2015).
Chin, R.K. et al. Lymphotoxin pathway directs thymic Aire expression. Nat. Immunol. 4, 1121–1127 (2003).
Hikosaka, Y. et al. The cytokine RANKL produced by positively selected thymocytes fosters medullary thymic epithelial cells that express autoimmune regulator. Immunity 29, 438–450 (2008).
LaFlam, T.N. et al. Identification of a novel cis-regulatory element essential for immune tolerance. J. Exp. Med. 212, 1993–2002 (2015).
Jones, P.A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Kont, V. et al. DNA methylation signatures of the AIRE promoter in thymic epithelial cells, thymomas and normal tissues. Mol. Immunol. 49, 518–526 (2011).
Rasmussen, K.D. & Helin, K. Role of TET enzymes in DNA methylation, development, and cancer. Genes Dev. 30, 733–750 (2016).
Gordon, J. et al. Specific expression of lacZ and cre recombinase in fetal thymic epithelial cells by multiplex gene targeting at the Foxn1 locus. BMC Dev. Biol. 7, 69 (2007).
Singh, H., Glasmacher, E., Chang, A.B. & Vander Lugt, B. The molecular choreography of IRF4 and IRF8 with immune system partners. Cold Spring Harb. Symp. Quant. Biol. 78, 101–104 (2013).
Su, M.A. et al. Mechanisms of an autoimmunity syndrome in mice caused by a dominant mutation in Aire. J. Clin. Invest. 118, 1712–1726 (2008).
Lara-Astiaso, D. et al. Chromatin state dynamics during blood formation. Science 345, 943–949 (2014).
Phillips, J.E. & Corces, V.G. CTCF: master weaver of the genome. Cell 137, 1194–1211 (2009).
Sleutels, F. et al. The male germ cell gene regulator CTCFL is functionally different from CTCF and binds CTCF-like consensus sites in a nucleosome composition-dependent manner. Epigenetics Chromatin 5, 8 (2012).
Verbeek, S. et al. An HMG-box-containing T-cell factor required for thymocyte differentiation. Nature 374, 70–74 (1995).
Dawlaty, M.M. et al. Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 9, 166–175 (2011).
Blecher-Gonen, R. et al. High-throughput chromatin immunoprecipitation for genome-wide mapping of in vivo protein-DNA interactions and epigenomic states. Nat. Protoc. 8, 539–554 (2013).
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Mayor, C. et al. VISTA : visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16, 1046–1047 (2000).
Kuo, H.-C. et al. DBCAT: database of CpG islands and analytical tools for identifying comprehensive methylation profiles in cancer cells. J. Comput. Biol. 18, 1013–1017 (2011).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Rosenbloom, K.R. et al. ENCODE data in the UCSC Genome Browser: year 5 update. Nucleic Acids Res. 41, D56–D63 (2013).
Acknowledgements
We thank M. Anderson (University of California at San Francisco) for Aire-Igrp-GFP mice; D. Graf (University of Zurich) for B6.Foxn1-Cre mice (obtained with the consent of N. Manley (University of Georgia)); H. Clevers (Hubrecht Institute) for Tcf7−/− mice; and Y. Peleg, G. Yona and V. Krupalnik for experimental expertise and help. Supported by the Israel Science Foundation (1825/10 and 1376/13), the Sy Syms Foundation, the Dr. Celia Zwillenberg-Fridman and Dr. Lutz Fridman Career Development Chair (J.A.), the Weizmann-German Cancer Research Center PhD fellowship program (Y.H., M.D., J.A., M.F.), the German Cancer Research Center–Israeli Ministry of Science and Technology foundation for German-Israeli co-operation (2431 to J.A. and M.F.), the Agence Nationale de Recherche (2011-CHEX-001-R12004KK to M.Gi.) and the European Federation of Immunological Societies fellowship program (M.R.B.).
Author information
Authors and Affiliations
Contributions
Y.H. and J.A. designed the study and wrote the manuscript; Y.H. performed most of the experimental work; S.N. performed several experiments, including ChIP, protein immunoprecipitation and Aire intracellular staining; C.B. performed the 'indexing-first' ChIP and ATAC-Seq experiments; M.R.B set up and conducted luciferase-based assays and analyzed the Tbx21-mutant mice; L.W. and S.V. constructed the Tet1fl/flTet2fl/flTet3fl/fl mice; M.D. assisted in performing the DNA bisulfite experiments; and S.B.-H., A.S., M.E.-B., Y.G., B.L., E.D., S.B.-D., M.G., J.H.H., A.B., F.L., I.A., M.F. and J.A. helped in performing, analyzing and/or designing some of the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Aire expression in various immune-cell populations.
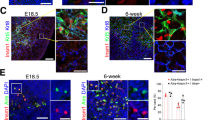
(a) Representative flow cytometry analyses of the expression of Aire.GFP (x-axis) and MHC class II (MHCII) (y-axis) in cell populations obtained from (n=3) 6-7week-old Aire.GFP mice, gated on thymic CD45+ EpCAM- CD11c- (T cells); CD45+ EpCAM- CD11c+ (Dendritic Cells; DCs); CD45- EpCAM+ Ly51+ (cortical thymic epithelial cells; cTECs); CD45- EpCAM+ Ly51- (medullary thymic epithelial cells; mTECs) or CD45+ EpCAM- CD19+ (B cells) populations. Numbers in outlined areas indicate %GFP+ cells, gated on an age-matched WT mouse. Data are representative of two independent experiments with similar results. (b) Quantitative PCR analysis of mean Aire expression in various sorted cell types obtained from same mice as indicated in (a). Results are normalized to the expression of Hprt and are presented relative to the expression in mature mTECs. (c) Genomic sequence of the predicted CpG islands. Color coded are the CG pairs. In light green are mTEC-specific differentially demethylated and in yellow are immature (MHCIIlo) mTEC-specific differentially demethylated cytosine residues.
Supplementary Figure 2 Quantitative PCR analysis verifies the mTEChi-cell-specific transcriptional regulator signature.
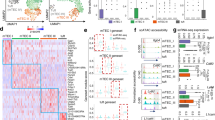
Clustered heat map depicting relative mRNA expression of indicated transcription regulators in sorted EpCAM+ Ly51neg-lo mature (MHCIIhi) vis-à-vis immature (MHCIllo) medullary or cortical (Ly51hi) thymic epithelial cells obtained from 6-7 weeks old mice (n=3). Shown are relative signal values normalized to expression of Hprt and relative to cTECs. Colors represent high (yellow) or low (blue) expression levels. Data are representative of at least three independent experiments with similar results.
Supplementary Figure 3 Aire reporter assays highlight potential mTEC-specific Aire regulators.
(a) Outline of the Aire promoter-based luciferase and RFP screening systems. (b) Representative flow cytometry analysis showing Aire.RFP reporter activity (x-axis) in HEK 293FT cells co-transfected with the Aire.RFP reporter vector alongside either an empty vector or expression vectors encoding all ~50 candidate mTEChi–specific transcription regulator genes for 48h. Numbers outlined present %RFPhi cells. Data are representative of at least three independent experiments with similar results. (c) Clustered heat map depicting mRNA expression profiles of candidate Aire-regulators that are predominantly expressed in antigen presenting cells or T cells and are predicted to activate the Aire promoter by either the Luciferase or the RFP reporter assays (Fig. 3e). Shown are normalized relative expression values of indicated immune cell populations. Colors represent high (yellow) or low (blue) expression levels. DCs, Dentiritic cells; T γδ, gamma delta T cells; T DP, Double positive (CD4+ CD8+) T cells; NK, Natural Killer cells.
Supplementary Figure 4 AIRE and AIRE-dependent genes are induced following expression of various transcriptional regulators.
(a+b) AIRE mRNA expression in HEK 293FT cells, treated with or without 5-Aza, encoding highlighted transcriptional regulators (a) or their combinations (b), and tested 48h later. (c-e) qPCR analyses showing mRNA expression of the AIRE-dependent genes ALOX12 (c) and KRT14 (d) or AIRE independent gene CAND1 (e) in 5-Aza treated cells transfected with the indicated factors and tested 48h later. (f-i) qPCR analyses showing AIRE mRNA levels in 5-Aza treated HEK 293FT cells transfected with expression vectors of Myb (f), Tbx21 (g), Tcf7 (h) or Tox4 (i) alongside short-listed candidates and measured 48h later; Results are normalized to the expression of HPRT and are presented relative to the expression in untreated cells transfected with an empty expression vector. Data in all experiments are representative of at least two independent experiments with similar results (mean and S.E.M of n = 2 biological replicates).
Supplementary Figure 5 mTECs with 50% Aire expression show normal TRA expression.
Quantitative PCR analysis of expression of Aire, several Aire-responsive TRAs (Ins2, Csn a, Mup4, Pcp4 and Spt1), and several Aire-neutral TRAs (Csn b and Gad67) in FACS-sorted mTEChi cells obtained from 6-8 week old Aire+/+, Aire+/– or Aire−/− mice (n=3). Results are normalized to the expression of Hprt and are presented relative to the expression values in Aire+/+ mTECs. Data are representative of two independent experiments with similar results (mean and S.E.M).
Supplementary Figure 6 Most candidate regulators bind the AIRE promoter independently of DNA-methylation status.
Chromatin immunoprecipitation followed by quantitative PCR assessing relative enrichment of indicated transfected candidates (vs. IgG control) along the AIRE TSS (corresponding to fragment 3 in Fig. 6b), in HEK 293FT cells treated 48h or untreated with 5-Aza. Results are normalized to values obtained from 5-Aza untreated cells. Data are representative of at least two independent experiments with similar results (mean and S.E.M of n = 2 biological replicates).
Supplementary Figure 7 Binding dynamics and endogenous expression of candidate transcriptional regulators.
(a-b) Chromatin immunoprecipitation followed by quantitative PCR assessing relative enrichment of CTCF following expression of Irf4/Irf8/Tbx21/Tcf7 (a) or of HA-tagged Irf4/Irf8/Tbx21/Tcf7 following expression of Ctcfl (b) in 5-Aza treated HEK 293FT cells transfected for 48h (n=3); (c-g) Quantitative PCR analyses showing endogenous expression of CTCF (c) HNF4G (d), IRF4 (e), IRF8 (f), TBX21 (g) and TCF7 (h) in 5-Aza treated HEK 293FT cells (pooled n=4) transfected with an siRNA set targeting expression of CTCF, and tested 48h later. Results are normalized to the expression of HPRT and are presented relative to the expression in the same cells transfected with a non-targeting siRNA set; Data are representative of at least two independent experiments with similar results. n.s. not significant, *P < 0.05, **P < 0.01 and ***P < 0.001 (Student's t-test, S.E.M).
Supplementary Figure 8 The molecular mechanisms that control the expression of Aire.
Aire is not expressed in the vast majority of cells in the body, except for a few cell types, primarily mature mTECs. To achieve such regulation, the Aire expression is regulated at multiple levels: First, the Aire locus is physically inaccessible and hypermethylated at specific CpG residues upstream and downstream to the TSS in cells and tissues that do not express it. Second, Aire locus is insulated by a global chromatin organizer - CTCF, which specifically binds at Aire’s TSS and downstream of the last Aire exon; Third, in mTECs, Aire expression is facilitated by concurrent eviction of CTCF, specific DNA demethylation and coordinated action of several transcription activators, including Irf4, Irf8, Tbx21, Tcf7 and Ctcfl, which act on mTEC-specific accessible regions in the Aire locus.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–3 (PDF 1340 kb)
Supplementary Table
Supplementary Table 4 (XLSX 8 kb)
Supplementary Table
Supplementary Table 5 (XLSX 12 kb)
Supplementary Table
Supplementary Table 6 (XLSX 12 kb)
Supplementary Table
Supplementary Table 7 (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Herzig, Y., Nevo, S., Bornstein, C. et al. Transcriptional programs that control expression of the autoimmune regulator gene Aire. Nat Immunol 18, 161–172 (2017). https://doi.org/10.1038/ni.3638
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3638
This article is cited by
-
Insm1 regulates mTEC development and immune tolerance
Cellular & Molecular Immunology (2023)
-
The function of brother of the regulator of imprinted sites in cancer development
Cancer Gene Therapy (2023)
-
Combined multidimensional single-cell protein and RNA profiling dissects the cellular and functional heterogeneity of thymic epithelial cells
Nature Communications (2023)
-
Thymic self-antigen expression for immune tolerance and surveillance
Inflammation and Regeneration (2022)
-
Novel antigen-presenting cell imparts Treg-dependent tolerance to gut microbiota
Nature (2022)