Abstract
Cell-surface-receptor pathways amplify weak, rare and local stimuli to induce cellular responses. This task is accomplished despite signaling components that segregate into nanometer-scale membrane domains. Here we describe a 'catch-and-release' mechanism that amplified and dispersed stimuli by releasing activated kinases from receptors lacking intrinsic catalytic activity. Specifically, we discovered a cycle of recruitment, activation and release for Zap70 kinases at phosphorylated T cell antigen receptors (TCRs). This turned the TCR into a 'catalytic unit' that amplified antigenic stimuli. Zap70 released from the TCR remained at the membrane, translocated, and phosphorylated spatially distinct substrates. The mechanisms described here are based on widely used protein domains and post-translational modifications; therefore, many membrane-associated pathways might employ similar mechanisms for signal amplification and dispersion.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Weiss, A. & Littman, D.R. Signal transduction by lymphocyte antigen receptors. Cell 76, 263–274 (1994).
Irvine, D.J., Purbhoo, M.A., Krogsgaard, M. & Davis, M.M. Direct observation of ligand recognition by T cells. Nature 419, 845–849 (2002).
Purbhoo, M.A., Irvine, D.J., Huppa, J.B. & Davis, M.M. T cell killing does not require the formation of a stable mature immunological synapse. Nat. Immunol. 5, 524–530 (2004).
Huse, M. et al. Spatial and temporal dynamics of T cell receptor signaling with a photoactivatable agonist. Immunity 27, 76–88 (2007).
van der Merwe, P.A. & Dushek, O. Mechanisms for T cell receptor triggering. Nat. Rev. Immunol. 11, 47–55 (2011).
Valitutti, S., Müller, S., Cella, M., Padovan, E. & Lanzavecchia, A. Serial triggering of many T-cell receptors by a few peptide-MHC complexes. Nature 375, 148–151 (1995).
McKeithan, T.W. Kinetic proofreading in T-cell receptor signal transduction. Proc. Natl. Acad. Sci. USA 92, 5042–5046 (1995).
Wülfing, C. et al. Costimulation and endogenous MHC ligands contribute to T cell recognition. Nat. Immunol. 3, 42–47 (2002).
Krogsgaard, M. et al. Agonist/endogenous peptide-MHC heterodimers drive T cell activation and sensitivity. Nature 434, 238–243 (2005).
Cooper, J.A. & Qian, H. A mechanism for SRC kinase-dependent signaling by noncatalytic receptors. Biochemistry 47, 5681–5688 (2008).
O'Donoghue, G.P., Pielak, R.M., Smoligovets, A.A., Lin, J.J. & Groves, J.T. Direct single molecule measurement of TCR triggering by agonist pMHC in living primary T cells. eLife 2, e00778 (2013).
Wang, H. et al. ZAP-70: an essential kinase in T-cell signaling. Cold Spring Harb. Perspect. Biol. 2, a002279 (2010).
Veillette, A., Caron, L., Fournel, M. & Pawson, T. Regulation of the enzymatic function of the lymphocyte-specific tyrosine protein kinase p56lck by the non-catalytic SH2 and SH3 domains. Oncogene 7, 971–980 (1992).
Stepanek, O. et al. Coreceptor scanning by the T cell receptor provides a mechanism for T cell tolerance. Cell 159, 333–345 (2014).
Baniyash, M., Garcia-Morales, P., Luong, E., Samelson, L.E. & Klausner, R.D. The T cell antigen receptor zeta chain is tyrosine phosphorylated upon activation. J. Biol. Chem. 263, 18225–18230 (1988).
Hatada, M.H. et al. Molecular basis for interaction of the protein tyrosine kinase ZAP-70 with the T-cell receptor. Nature 377, 32–38 (1995).
Watts, J.D. et al. Identification by electrospray ionization mass spectrometry of the sites of tyrosine phosphorylation induced in activated Jurkat T cells on the protein tyrosine kinase ZAP-70. J. Biol. Chem. 269, 29520–29529 (1994).
Chan, A.C. et al. Activation of ZAP-70 kinase activity by phosphorylation of tyrosine 493 is required for lymphocyte antigen receptor function. EMBO J. 14, 2499–2508 (1995).
Brdicka, T., Kadlecek, T.A., Roose, J.P., Pastuszak, A.W. & Weiss, A. Intramolecular regulatory switch in ZAP-70: analogy with receptor tyrosine kinases. Mol. Cell. Biol. 25, 4924–4933 (2005).
Horejsí, V., Zhang, W. & Schraven, B. Transmembrane adaptor proteins: organizers of immunoreceptor signalling. Nat. Rev. Immunol. 4, 603–616 (2004).
Klammt, C. et al. T cell receptor dwell times control the kinase activity of Zap70. Nat. Immunol. 16, 961–969 (2015).
Dustin, M.L. & Groves, J.T. Receptor signaling clusters in the immune synapse. Annu. Rev. Biophys. 41, 543–556 (2012).
Klammt, C. & Lillemeier, B.F. How membrane structures control T cell signaling. Front. Immunol. 3, 291 (2012).
Lillemeier, B.F., Pfeiffer, J.R., Surviladze, Z., Wilson, B.S. & Davis, M.M. Plasma membrane-associated proteins are clustered into islands attached to the cytoskeleton. Proc. Natl. Acad. Sci. USA 103, 18992–18997 (2006).
Lillemeier, B.F. et al. TCR and Lat are expressed on separate protein islands on T cell membranes and concatenate during activation. Nat. Immunol. 11, 90–96 (2010).
Sherman, E. et al. Functional nanoscale organization of signaling molecules downstream of the T cell antigen receptor. Immunity 35, 705–720 (2011).
Bunnell, S.C. et al. T cell receptor ligation induces the formation of dynamically regulated signaling assemblies. J. Cell Biol. 158, 1263–1275 (2002).
Yokosuka, T. et al. Newly generated T cell receptor microclusters initiate and sustain T cell activation by recruitment of Zap70 and SLP-76. Nat. Immunol. 6, 1253–1262 (2005).
Cambi, A. et al. Organization of the integrin LFA-1 in nanoclusters regulates its activity. Mol. Biol. Cell 17, 4270–4281 (2006).
Varma, R., Campi, G., Yokosuka, T., Saito, T. & Dustin, M.L. T cell receptor-proximal signals are sustained in peripheral microclusters and terminated in the central supramolecular activation cluster. Immunity 25, 117–127 (2006).
Monks, C.R., Freiberg, B.A., Kupfer, H., Sciaky, N. & Kupfer, A. Three-dimensional segregation of supramolecular activation clusters in T cells. Nature 395, 82–86 (1998).
Grakoui, A. et al. The immunological synapse: a molecular machine controlling T cell activation. Science 285, 221–227 (1999).
Hashimoto-Tane, A. et al. Dynein-driven transport of T cell receptor microclusters regulates immune synapse formation and T cell activation. Immunity 34, 919–931 (2011).
Deindl, S. et al. Structural basis for the inhibition of tyrosine kinase activity of ZAP-70. Cell 129, 735–746 (2007).
Yan, Q. et al. Structural basis for activation of ZAP-70 by phosphorylation of the SH2-kinase linker. Mol. Cell. Biol. 33, 2188–2201 (2013).
Park, M.J. et al. SH2 domains serve as lipid-binding modules for pTyr-signaling proteins. Mol. Cell 62, 7–20 (2016).
Keshvara, L.M., Isaacson, C., Harrison, M.L. & Geahlen, R.L. Syk activation and dissociation from the B-cell antigen receptor is mediated by phosphorylation of tyrosine 130. J. Biol. Chem. 272, 10377–10381 (1997).
Zhang, Y. et al. Tyr130 phosphorylation triggers Syk release from antigen receptor by long-distance conformational uncoupling. Proc. Natl. Acad. Sci. USA 105, 11760–11765 (2008).
Szabo, M. et al. Fine-tuning of proximal TCR signaling by ZAP-70 tyrosine residues in Jurkat cells. Int. Immunol. 24, 79–87 (2012).
Lovering, F. et al. Identification of type-II inhibitors using kinase structures. Chem. Biol. Drug Des. 80, 657–664 (2012).
Isakov, N. et al. ZAP-70 binding specificity to T cell receptor tyrosine-based activation motifs: the tandem SH2 domains of ZAP-70 bind distinct tyrosine-based activation motifs with varying affinity. J. Exp. Med. 181, 375–380 (1995).
Bu, J.Y., Shaw, A.S. & Chan, A.C. Analysis of the interaction of ZAP-70 and syk protein-tyrosine kinases with the T-cell antigen receptor by plasmon resonance. Proc. Natl. Acad. Sci. USA 92, 5106–5110 (1995).
Evans, R. et al. Integrins in immunity. J. Cell Sci. 122, 215–225 (2009).
Oh, D. et al. Fast rebinding increases dwell time of Src homology 2 (SH2)-containing proteins near the plasma membrane. Proc. Natl. Acad. Sci. USA 109, 14024–14029 (2012).
Huppa, J.B. et al. TCR-peptide-MHC interactions in situ show accelerated kinetics and increased affinity. Nature 463, 963–967 (2010).
Pawson, T., Gish, G.D. & Nash, P. SH2 domains, interaction modules and cellular wiring. Trends Cell Biol. 11, 504–511 (2001).
Pawson, T. & Nash, P. Assembly of cell regulatory systems through protein interaction domains. Science 300, 445–452 (2003).
Wilson, B.S., Pfeiffer, J.R. & Oliver, J.M. Observing FcɛRI signaling from the inside of the mast cell membrane. J. Cell Biol. 149, 1131–1142 (2000).
Mattila, P.K. et al. The actin and tetraspanin networks organize receptor nanoclusters to regulate B cell receptor-mediated signaling. Immunity 38, 461–474 (2013).
O'Brien, R. et al. Alternative modes of binding of proteins with tandem SH2 domains. Protein Sci. 9, 570–579 (2000).
Wolters, D.A., Washburn, M.P. & Yates, J.R. III. An automated multidimensional protein identification technology for shotgun proteomics. Anal. Chem. 73, 5683–5690 (2001).
He, L., Diedrich, J., Chu, Y.Y. & Yates, J.R. III. Extracting accurate precursor information for tandem mass spectra by RawConverter. Anal. Chem. 87, 11361–11367 (2015).
Peng, J., Elias, J.E., Thoreen, C.C., Licklider, L.J. & Gygi, S.P. Evaluation of multidimensional chromatography coupled with tandem mass spectrometry (LC/LC-MS/MS) for large-scale protein analysis: the yeast proteome. J. Proteome Res. 2, 43–50 (2003).
Xu, T. et al. ProLuCID: An improved SEQUEST-like algorithm with enhanced sensitivity and specificity. J. Proteomics 129, 16–24 (2015).
Tabb, D.L., McDonald, W.H. & Yates, J.R. III. DTASelect and Contrast: tools for assembling and comparing protein identifications from shotgun proteomics. J. Proteome Res. 1, 21–26 (2002).
Acknowledgements
We thank J. Williamson (the Scripps Research Institute) for use of the BLI instrument; the Salk Biophotonics Core for assistance with FRAP acquisition; the Salk Flow Cytometry Core Facility for assistance with flow-cytometry-based analysis of calcium flux and cell sorting; Jolene Diedrich and James Moresco for mass spectrometry analyses; M. Hetzer, T. Hunter, P. Hogan, G. Lemke, and J. Karlseder for critical reading of the manuscript; A. Weiss (University of California San Francisco, San Francisco, California, USA) for parental Jurkat cell lines; M.M. Davis (Stanford University, Stanford, California) for HEK293T cells; and the Nomis Foundation, the Waitt Foundation and the James B. Pendleton Charitable Trust for support. Supported by the US National Institutes of Health (1DP2GM105455-01 to B.F.L. and T32 CA009370 to Z.B.K.), the Salk Institute Cancer Center core facilities (funded by CA014195 from the National Cancer Institute), the Mass Spectrometry Core of the Salk Institute (supported by P30 014195 from the National Cancer Institute Cancer Center Support Grants of the US National Institutes of Health) and the Helmsley Center for Genomic Medicine.
Author information
Authors and Affiliations
Contributions
Z.B.K. and B.F.L. designed the study and wrote the manuscript; Z.B.K. performed SPT, Ca2+-flux measurements, phospho-flow cytometry and statistical analyses; L.N. performed FRAP experiments; L.N. and A.B. established expression systems for recombinant proteins; and B.F.L. performed kinase and adhesion assays.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 SPT analyses of Zap70 at the T cell plasma membrane.
(a) Total internal reflection fluorescent (TIRF) excitation at 64-68 degrees limits single molecule detection to 100-150 nm above the glass surface. (b) Distribution of single molecule localization precisions from all trajectories of murine wild-type Zap70 (WT; black) and the tandem SH2 module (tSH2; brown) fused to PATagRFP in primary 5c.c7 T cells. (c) Average Trajectory lifetimes. (d) The average diffusion rate (τ = 500 ms). (e) Average trajectory spans (distance between the two furthest points in a trajectory). (f) Distribution of single molecule localization precisions from all trajectories in P116 Jurkat T cell lines expressing different forms of human Zap70 fused to PATagRFP. (g) Average trajectory lifetimes. (h) The average diffusion rate (τ = 500 ms). (i) Average trajectory spans. The T cell receptor (specifically CD3ɛ) was tracked on poly-L-lysine surfaces to reference free movement within the plasma membrane. (j) Zap70 trajectories in a fixed cell expressing WT overlaid with diffraction limited TCR microclusters (gray pixels). (k) Representative tracks of Zap70’s tSH2 module on non-stimulatory PLL surface. Data represents at least 3 independent experiments and 18 different cells for each condition. Significance was assessed with a two-tailed unpaired t-test between indicated samples (* p<0.05, ** p<0.01, *** p<0.001).
Supplementary Figure 2 Inducible phosphorylation of Tyr126 in Zap70.
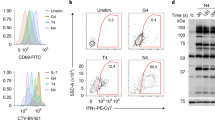
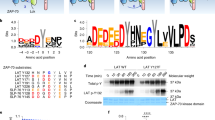
(a) Detection of Y126 phosphorylation upon OKT3/OKT4 stimulation by mass spectrometry. Chromatogram of p-Y126 containing peptide (sequence shown above) for non-stimulated (top) and stimulated (bottom) T cells. Graph (right) shows the percentage of p-Y126 phosphorylated peptide. (b) Western Blot analyses of lysates from non- and OKT3-stimulated P116 Jurkat T cells expressing WT, Y126E and Y126F using a rabbit serum raised against a peptide containing p-Y126.
Supplementary Figure 3 Effects of endogenous Zap70 on the mobility of Zap70 mutants.
(a) Confocal images of microclusters from fluorescence recovery after photobleaching (FRAP) experiments of GFP tagged Zap70 variants stably expressed in P116 (top) and E6.1 (bottom) Jurkat T cells. Scale bars represents 10 μm. (b) Microclusters of GFP tagged tSH2, tSH2-Y126E and tSH2-Y126F transiently expressed in P116 (top) and E6.1 (bottom) Jurkat T cells. (c) Relative expression of Zap70-GFP constructs stably expressed in E6.1 cells with endogenous Zap70 and P116 exogenously expressing wild-type Zap70. (d) Trajectories of WT, tSH2, Y126E and Y126F fused to PATagRFP in E6.1 Jurkat T cells expressing endogenous Zap70. Trajectories are overlaid on diffraction-limited images of TCR microclusters (CD3ζ-GFP; gray; 500 nm scale bar). Pixels are 160×160 nm. (e) Comparison of mean square displacement of Zap70 variants in P116 (cyan), E6.1 (red), and P116 + wild-type Zap70 Jurkat T cells (purple), with P116 WT curve as reference (black solid line). (f) FRAP recovery curves for WT, tSH2, Y126E and Y126F fused to GFP in P116 (cyan), E6.1 Jurkat T cells (red), and P116 + wild-type Zap70 Jurkat T cells (purple). The data for WT in P116 Jurkat T cells is inserted as reference in each plot (solid black curve). (g) The average diffusion rate (τ = 500 ms) from SPT. (h) Bar graphs of average mobile fraction from FRAP. (i) average recovery rates (t1/2) from FRAP. Tracking data represents averages from at least ten cells from 3 independent experiments with ± s.e.m. Pixels are 160×160 nm. FRAP data represent averages with s.e.m. from at least 11 different cells from 3 or more independent experiments. Error bars for mobile fractions and recovery rates (t1/2) represent the 95% confidence interval.
Supplementary Figure 4 Inhibition of Zap70’s release at late time points requires Zap70’s kinase activity.
(a) Confocal images of microclusters from fluorescence recovery after photobleaching (FRAP) experiments of GFP tagged Zap70 ATP binding mutants (specifically K369A and K369R) stably expressed in P116 (left) and E6.1 (right) Jurkat T cells. (b) Kinase dead Zap70s were analyzed in Zap70 deficient P116 (left) and E6.1 Jurkat T cells (right). Early time points (<10 min; black lines) are compared to data at late time points (>30 min; gray lines). (c) Bar graphs of average mobile fraction and average recovery rates (t1/2). FRAP data represent averages with s.e.m. from at least 15 different cells from 3 or more independent experiments. Error bars for mobile fractions and recovery rates t1/2 represent the 95% confidence interval.
Supplementary Figure 5 Zap70’s release from the TCR controls downstream signaling.
(a) Flow cytometry analysis of original (left) and expression matched (right) WT, Y126E and Y126F expressing P116 Jurkat T cell lines. Original cell lines were sorted for overlapping expression level (gray bar) and cultivated for one week before re-analyses. (b) Western-blot analyses of Y493 phosphorylation (p-Y493, left) and total Zap70 protein (right) in non-sorted P116 Jurkat T cell lines after stimulation with different concentrations of OKT3. (c) Western-blot analyses of Lat phosphorylation (p-Y171 + p-Y191; left) and total Lat protein (right) in P116 Jurkat T cell lines after stimulation with different concentrations of OKT3.
Supplementary Figure 6 IL-2 secretion and Ca2+ flux of Jurkat T cell lines.
(a) IL-2 secretion of P116 Jurkat T cell lines after stimulation with SEE pulsed Raji B cells. (b) IL-2 secretion of WT, Y126E and Y126F expressing P116 Jurkat T cell lines and E6.1 Jurkat T cells after stimulation with OKT3. Data represents averages ± s.e.m. from at least two independent measurements with duplicates. (c) Calcium flux of P116 Jurkat T cell lines stably expressing WT, Y126E and Y126F with 10 μg/ml, 1 μg/ml and 0.1 μg/ml OKT3. Ca-Flux was analyzed for T cells with matched GFP expression. The P116 Zap70 null cells and Kinase dead Zap70 mutants (K369A and K369R) are shown for 10 μg/ml OKT3 stimulation. Curves are population averages of the violet/blue ratio of Indo1 ratiometric calcium dye. (d) ER calcium release of sorted P116 WT, Y126E, and Y126F cell lines assessed by ratiometric Fura2 imaging on activating surfaces (10 μg/ml OKT3) in the absence of extracellular calcium. Curves represent averages ± s.e.m. from at least three independent experiments with at least ten cells analyzed from each experiment.
Supplementary Figure 7 Alteration of Zap70’s release from the TCR reduces adhesion to ICAM-1 after T cell stimulation.
(a) Representative images of P116 Jurkat T cells stably expressing WT and mutant Zap70s adhered to ICAM-1 surfaces after stimulation with different concentrations of OKT3 (indicated above). Left column shows maximal adhesion upon treatment with MgCl2/EGTA. (b) Quantification of cell adhesion by extraction of crystal violet from adherent cells in relation to MgCl2/EGTA treatment. (c) Surface area covered by P116 Jurkat T cell lines in relation to MgCl2/EGTA treatment. Scale bars are 20 μm. Error bars represent s.e.m. from three independent experiments with duplicates. Three images of different regions quantified for each sample. Significance was assessed with a two-tailed unpaired t-test (* p<0.05, ** p<0.01, *** p<0.001).
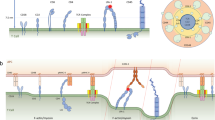
Supplementary Figure 8 The ‘catch-and-release’ model for TCR signal amplification and dispersion.
Proposed ‘Catch-And-Release’ model for early and late Zap70 signaling at the TCR. Signaling events are indicated by numbered circles. In brief, upon TCR engagement by pMHC, Zap70 kinase is recruited to the Lck-phosphorylated ITAMs of the TCR-CD3-complex. During early signaling Zap70 is Lck- and/or auto-transphosphorylated on Y126, activated and binds ATP. Altogether, this induces Zap70’s release from the TCR into the plane of the plasma membrane. Released Zap70 translocates to other membrane domains to phosphorylate its substrates (e.g. LAT). The vacated p-ITAM is available to recruit, activate and release additional Zap70 molecules. This turns the TCR into a ‘catalytic unit’ that amplifies antigenic stimuli. During later stages of T cell activation Zap70 release does not take place and Zap70 maintains TCR signaling while stably associated with p-ITAMs, probably through phosphorylation of Y315 and Y319.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–12 (PDF 2424 kb)
Single particle tracking of Zap70 WT in primary T cells.
Single molecules of wild-type Zap70 (WT) tagged with PATagRFP in 5c.c7 primary T cells on I-EK/MCC + CD80 surfaces during the first ten minutes of stimulation were visualized by TIRF microscopy (left panel). Snapshot of TCR microclusters (specifically CD3ζ-GFP; middle panel) and the merge of both channels (right panel) (MOV 6137 kb)
Single particle tracking of Zap70's tSH2 module in primary T cells.
Single molecules of the Zap70 tandem SH2 domain module (tSH2) tagged with PATagRFP in 5c.c7 primary T cells on I-EK/MCC + CD80 surfaces during the first ten minutes of stimulation were visualized by TIRF microscopy (left panel). Snapshot of TCR microclusters (specifically CD3ζ-GFP; middle panel) and the merge of both channels (right panel). (MOV 6012 kb)
Single particle tracking of CD3ɛ in P116 Jurkat T cells on PLL only
Single molecules of CD3ɛ tagged with PATagRFP in P116 Jurkat T cells on PLL surfaces were visualized by TIRF microscopy (left panel). Snapshot of Zap70 signal at the membrane (specifically Zap70-GFP; middle panel) and the merge of both channels (right panel). (MOV 10963 kb)
Single particle tracking of CD3ɛ in P116 Jurkat T cells.
Single molecules of CD3ɛ tagged with PATagRFP in P116 Jurkat T cells on OKT3 surfaces during the first ten minutes of stimulation were visualized by TIRF microscopy (left panel). Snapshot of Zap70 microclusters (specifically Zap70-GFP; middle panel) and the merge of both channels (right panel). (MOV 9316 kb)
Single particle tracking of wild-type Zap70 in P116 Jurkat T cells.
Single molecules of wild-type Zap70 (WT) tagged with PATagRFP in P116 Jurkat T cells on OKT3 surfaces during the first ten minutes of stimulation were visualized by TIRF microscopy (left panel). Snapshot of TCR microclusters (specifically CD3ζ-GFP; middle panel) and the merge of both channels (right panel). (MOV 9529 kb)
Single particle tracking of tSH2 in P116 Jurkat T cells.
Single molecules of the Zap70 tandem SH2 domain module (tSH2) tagged with PATagRFP in P116 Jurkat T cells on OKT3 surfaces during the first ten minutes of stimulation were visualized by TIRF microscopy (left panel). Snapshot of TCR microclusters (specifically CD3ζ-GFP; middle panel) and the merge of both channels (right panel). (MOV 9865 kb)
Single particle tracking of wild-type Zap70 in fixed T cells.
Single molecules of wild-type Zap70 (WT) tagged with PATagRFP in fixed T cells were visualized by TIRF microscopy (left panel). Snapshot of TCR microclusters (specifically CD3ζ-GFP; middle panel) and the merge of both channels (right panel). (MOV 5420 kb)
Single particle tracking of tSH2 in P116 Jurkat T cells on PLL only.
Single molecules of the Zap70 tandem SH2 domain module (tSH2) tagged with PATagRFP in P116 Jurkat T cells on PLL surfaces were visualized by TIRF microscopy (left panel). Snapshot of TCR membrane signal (specifically CD3ζ-GFP; middle panel) and the merge of both channels (right panel). (MOV 1726 kb)
Single particle tracking of Y126E in P116 Jurkat T cells.
Single molecules of the Zap70 Y126 glutamic acid mutant (Y126E) tagged with PATagRFP in P116 Jurkat T cells on OKT3 surfaces during the first ten minutes of stimulation were visualized by TIRF microscopy (left panel). Snapshot of TCR microclusters (specifically CD3ζ-GFP; middle panel) and the merge of both channels (right panel). (MOV 10485 kb)
Single particle tracking of Y126F in P116 Jurkat T cells.
Single molecules of the Zap70 Y126 phenylalanine mutant (Y126F) tagged with PATagRFP in P116 Jurkat T cells on OKT3 surfaces during the first ten minutes of stimulation were visualized by TIRF microscopy (left panel). Snapshot of TCR microclusters (specifically CD3ζ-GFP; middle panel) and the merge of both channels (right panel). (MOV 9856 kb)
Single particle tracking of K369A in P116 Jurkat T cells.
Single molecules of the kinase dead Zap70 K369 alanine mutant (K369A) tagged with PATagRFP in P116 Jurkat T cells on OKT3 surfaces during the first ten minutes of stimulation were visualized by TIRF microscopy (left panel). Snapshot of TCR microclusters (specifically CD3ζ-GFP; middle panel) and the merge of both channels (right panel). (MOV 8900 kb)
Single particle tracking of K369R in P116 Jurkat T cells.
Single molecules of the kinase dead Zap70 K369 lysine mutant (K369R) tagged with PATagRFP in P116 Jurkat T cells on OKT3 surfaces during the first ten minutes of stimulation were visualized by TIRF microscopy (left panel). Snapshot of TCR microclusters (specifically CD3ζ-GFP; middle panel) and the merge of both channels (right panel). (MOV 10014 kb)
Rights and permissions
About this article
Cite this article
Katz, Z., Novotná, L., Blount, A. et al. A cycle of Zap70 kinase activation and release from the TCR amplifies and disperses antigenic stimuli. Nat Immunol 18, 86–95 (2017). https://doi.org/10.1038/ni.3631
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3631
This article is cited by
-
i-shaped antibody engineering enables conformational tuning of biotherapeutic receptor agonists
Nature Communications (2024)
-
Recent advances in anti-inflammatory active components and action mechanisms of natural medicines
Inflammopharmacology (2023)
-
Molecular dynamics simulations reveal membrane lipid interactions of the full-length lymphocyte specific kinase (Lck)
Scientific Reports (2022)
-
Host response transcriptomic analysis of Crimean-Congo hemorrhagic fever pathogenesis in the cynomolgus macaque model
Scientific Reports (2021)
-
Rapid statistical discrimination of fluorescence images of T cell receptors on immobilizing surfaces with different coating conditions
Scientific Reports (2021)