Abstract
The manner in which regulatory T cells (Treg cells) control lymphocyte homeostasis is not fully understood. We identified two Treg cell populations with differing degrees of self-reactivity and distinct regulatory functions. We found that GITRhiPD-1hiCD25hi (Triplehi) Treg cells were highly self-reactive and controlled lympho-proliferation in peripheral lymph nodes. GITRloPD-1loCD25lo (Triplelo) Treg cells were less self-reactive and limited the development of colitis by promoting the conversion of CD4+ Tconv cells into induced Treg cells (iTreg cells). Although Foxp3-deficient (Scurfy) mice lacked Treg cells, they contained Triplehi-like and Triplelo-like CD4+ T cells zsuper> T cells infiltrated the skin, whereas Scurfy TripleloCD4+ T cells induced colitis and wasting disease. These findings indicate that the affinity of the T cell antigen receptor for self antigen drives the differentiation of Treg cells into distinct subsets with non-overlapping regulatory activities.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Fontenot, J.D., Gavin, M.A. & Rudensky, A.Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Khattri, R., Cox, T., Yasayko, S.A. & Ramsdell, F. An essential role for Scurfin in CD4+CD25+ T regulatory cells. Nat. Immunol. 4, 337–342 (2003).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061 (2003).
Bennett, C.L. et al. The immune dysregulation, polyendocrinopathy, enteropathy, X-linked syndrome (IPEX) is caused by mutations of FOXP3. Nat. Genet. 27, 20–21 (2001).
Josefowicz, S.Z., Lu, L.F. & Rudensky, A.Y. Regulatory T cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012).
Ohkura, N., Kitagawa, Y. & Sakaguchi, S. Development and maintenance of regulatory T cells. Immunity 38, 414–423 (2013).
Harrison, O.J. & Powrie, F.M. Regulatory T cells and immune tolerance in the intestine. Cold Spring Harb. Perspect. Biol. 5, 5 (2013).
Jordan, M.S. et al. Thymic selection of CD4+CD25+ regulatory T cells induced by an agonist self-peptide. Nat. Immunol. 2, 301–306 (2001).
Apostolou, I., Sarukhan, A., Klein, L. & von Boehmer, H. Origin of regulatory T cells with known specificity for antigen. Nat. Immunol. 3, 756–763 (2002).
Klein, L. & Jovanovic, K. Regulatory T cell differentiation: turning harmful into useful. Immunity 37, 441–443 (2012).
Chen, W. et al. Conversion of peripheral CD4+CD25- naive T cells to CD4+CD25+ regulatory T cells by TGF-beta induction of transcription factor Foxp3. J. Exp. Med. 198, 1875–1886 (2003).
Zheng, Y. et al. Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature 463, 808–812 (2010).
Samstein, R.M., Josefowicz, S.Z., Arvey, A., Treuting, P.M. & Rudensky, A.Y. Extrathymic generation of regulatory T cells in placental mammals mitigates maternal-fetal conflict. Cell 150, 29–38 (2012).
Yadav, M. et al. Neuropilin-1 distinguishes natural and inducible regulatory T cells among regulatory T cell subsets. in vivo. J Exp Med 209, 1713–1722 (2012).
Haribhai, D. et al. A requisite role for induced regulatory T cells in tolerance based on expanding antigen receptor diversity. Immunity 35, 109–122 (2011).
Smigiel, K.S. et al. CCR7 provides localized access to IL-2 and defines homeostatically distinct regulatory T cell subsets. J. Exp. Med. 211, 121–136 (2014).
Huehn, J. et al. Developmental stage, phenotype, and migration distinguish naive- and effector/memory-like CD4+ regulatory T cells. J. Exp. Med. 199, 303–313 (2004).
Miyara, M. et al. Functional delineation and differentiation dynamics of human CD4+ T cells expressing the FoxP3 transcription factor. Immunity 30, 899–911 (2009).
Levine, A.G., Arvey, A., Jin, W. & Rudensky, A.Y. Continuous requirement for the TCR in regulatory T cell function. Nat. Immunol. 15, 1070–1078 (2014).
Schmidt, A.M. et al. Regulatory T cells require TCR signaling for their suppressive function. J. Immunol. 194, 4362–4370 (2015).
Hsieh, C.S., Zheng, Y., Liang, Y., Fontenot, J.D. & Rudensky, A.Y. An intersection between the self-reactive regulatory and nonregulatory T cell receptor repertoires. Nat. Immunol. 7, 401–410 (2006).
Lee, H.-M., Bautista, J.L., Scott-Browne, J., Mohan, J.F. & Hsieh, C.-S. A broad range of self-reactivity drives thymic regulatory T cell selection to limit responses to self. Immunity 37, 475–486 (2012).
Pacholczyk, R. et al. Nonself-antigens are the cognate specificities of Foxp3+ regulatory T cells. Immunity 27, 493–504 (2007).
Hsieh, C.S. et al. Recognition of the peripheral self by naturally arising CD25+ CD4+ T cell receptors. Immunity 21, 267–277 (2004).
Pacholczyk, R., Ignatowicz, H., Kraj, P. & Ignatowicz, L. Origin and T cell receptor diversity of Foxp3+CD4+CD25+ T cells. Immunity 25, 249–259 (2006).
Lathrop, S.K. et al. Peripheral education of the immune system by colonic commensal microbiota. Nature 478, 250–254 (2011).
Yadav, M., Stephan, S. & Bluestone, J.A. Peripherally induced Tregs: role in immune homeostasis and autoimmunity. Front. Immunol. 4, 232 (2013).
Yang, E., Zou, T., Leichner, T.M., Zhang, S.L. & Kambayashi, T. Both retention and recirculation contribute to long-lived regulatory T-cell accumulation in the thymus. Eur. J. Immunol. 44, 2712–2720 (2014).
Cowan, J.E., McCarthy, N.I. & Anderson, G. CCR7 controls thymus recirculation, but not production and emigration, of Foxp3(+) T cells. Cell Rep. 14, 1041–1048 (2016).
Kim, K.S. et al. Dietary antigens limit mucosal immunity by inducing regulatory T cells in the small intestine. Science 351, 858–863 (2016).
Moran, A.E. et al. T cell receptor signal strength in Treg and iNKT cell development demonstrated by a novel fluorescent reporter mouse. J. Exp. Med. 208, 1279–1289 (2011).
Mandl, J.N., Monteiro, J.P., Vrisekoop, N. & Germain, R.N. T cell-positive selection uses self-ligand binding strength to optimize repertoire recognition of foreign antigens. Immunity 38, 263–274 (2013).
Kuczma, M. et al. Foxp3-deficient regulatory T cells do not revert into conventional effector CD4+ T cells but constitute a unique cell subset. J. Immunol. 183, 3731–3741 (2009).
Lin, W. et al. Regulatory T cell development in the absence of functional Foxp3. Nat. Immunol. 8, 359–368 (2007).
Wagner, N. et al. Critical role for beta7 integrins in formation of the gut-associated lymphoid tissue. Nature 382, 366–370 (1996).
Killebrew, J.R. et al. A self-reactive TCR drives the development of Foxp3+ regulatory T cells that prevent autoimmune disease. J. Immunol. 187, 861–869 (2011).
Thornton, A.M. et al. Expression of Helios, an Ikaros transcription factor family member, differentiates thymic-derived from peripherally induced Foxp3+ T regulatory cells. J. Immunol. 184, 3433–3441 (2010).
Weiss, J.M. et al. Neuropilin 1 is expressed on thymus-derived natural regulatory T cells, but not mucosa-generated induced Foxp3+ T reg cells. J Exp Med 209, 1723–1742 (2012).
Kim, J.M., Rasmussen, J.P. & Rudensky, A.Y. Regulatory T cells prevent catastrophic autoimmunity throughout the lifespan of mice. Nat. Immunol. 8, 191–197 (2007).
Morelli, A.E. & Thomson, A.W. Tolerogenic dendritic cells and the quest for transplant tolerance. Nat. Rev. Immunol. 7, 610–621 (2007).
Liu, K. et al. In vivo analysis of dendritic cell development and homeostasis. Science 324, 392–397 (2009).
Tadokoro, C.E. et al. Regulatory T cells inhibit stable contacts between CD4+ T cells and dendritic cells in vivo. J. Exp. Med. 203, 505–511 (2006).
Mottet, C., Uhlig, H.H. & Powrie, F. Cutting edge: cure of colitis by CD4+CD25+ regulatory T cells. J. Immunol. 170, 3939–3943 (2003).
Round, J.L. & Mazmanian, S.K. Inducible Foxp3+ regulatory T-cell development by a commensal bacterium of the intestinal microbiota. Proc. Natl. Acad. Sci. USA 107, 12204–12209 (2010).
Haribhai, D. et al. Alternatively activated macrophages boost induced regulatory T and Th17 cell responses during immunotherapy for colitis. J. Immunol. 196, 3305–3317 (2016).
Sharma, R., Sung, S.S., Gaskin, F., Fu, S.M. & Ju, S.T. A novel function of IL-2: chemokine/chemoattractant/retention receptor genes induction in Th subsets for skin and lung inflammation. J. Autoimmun. 38, 322–331 (2012).
Zelante, T., Fric, J., Wong, A.Y. & Ricciardi-Castagnoli, P. Interleukin-2 production by dendritic cells and its immuno-regulatory functions. Front. Immunol. 3, 161 (2012).
Kurts, C. et al. CD4+ T cell help impairs CD8+ T cell deletion induced by cross-presentation of self-antigens and favors autoimmunity. J. Exp. Med. 186, 2057–2062 (1997).
Koehli, S., Naeher, D., Galati-Fournier, V., Zehn, D. & Palmer, E. Optimal T-cell receptor affinity for inducing autoimmunity. Proc. Natl. Acad. Sci. USA 111, 17248–17253 (2014).
Barnden, M.J., Allison, J., Heath, W.R. & Carbone, F.R. Defective TCR expression in transgenic mice constructed using cDNA-based alpha- and beta-chain genes under the control of heterologous regulatory elements. Immunol. Cell Biol. 76, 34–40 (1998).
Lin, W. et al. Allergic dysregulation and hyperimmunoglobulinemia E in Foxp3 mutant mice. J. Allergy Clin. Immunol. 116, 1106–1115 (2005).
Huseby, E.S. et al. How the T cell repertoire becomes peptide and MHC specific. Cell 122, 247–260 (2005).
Malissen, M. et al. Altered T cell development in mice with a targeted mutation of the CD3-epsilon gene. EMBO J. 14, 4641–4653 (1995).
Wang, Y. et al. Th2 lymphoproliferative disorder of LatY136F mutant mice unfolds independently of TCR-MHC engagement and is insensitive to the action of Foxp3+ regulatory T cells. J. Immunol. 180, 1565–1575 (2008).
Zehn, D. & Bevan, M.J. T cells with low avidity for a tissue-restricted antigen routinely evade central and peripheral tolerance and cause autoimmunity. Immunity 25, 261–270 (2006).
Stadinski, B.D. et al. A role for differential variable gene pairing in creating T cell receptors specific for unique major histocompatibility ligands. Immunity 35, 694–704 (2011).
Vanguri, V., Govern, C.C., Smith, R. & Huseby, E.S. Viral antigen density and confinement time regulate the reactivity pattern of CD4 T-cell responses to vaccinia virus infection. Proc. Natl. Acad. Sci. USA 110, 288–293 (2013).
White, A., Jenkinson, E. & Anderson, G. Reaggregate thymus cultures. J. Vis. Exp. 18, 905 (2008).
Aronesty, E. ea-utils: “Command-line tools for processing biological sequencing data”. 2011 [cited]Available from: http://code.google.com/p/ea-utils.
Yang, X. et al. TCRklass: a new K-string-based algorithm for human and mouse TCR repertoire characterization. J. Immunol. 194, 446–454 (2015).
Colwell, F.S. et al. Estimates of biogenic methane production rates in deep marine sediments at Hydrate Ridge, Cascadia margin. Appl. Environ. Microbiol. 74, 3444–3452 (2008).
Shannon, C.E. A mathematical theory of communication. Bell Syst. Tech. J. 27, 379–423 (1948).
Simpson, E.H. Measurement of diversity. Nature 163, 688–688 (1949).
Acknowledgements
We thank U. Schneider for animal husbandry, E. Traunecker and T. Krebs for cell sorting, and G. DeLibero, L. Jeker and O. Stepanek for reviewing the manuscript. This study was funded by grants 310030-149972/1 [SNF], Sybilla [EU FP7], and TerraIncognita [ERC] (E.P.); RO1-DK095077, U19 AI109858 and UMass DERC grant DK32520 (E.S.H.); T32 AI 007349 (B.D.S.); Federal Ministry of Education and Research grant (BMBF), German Center for Diabetes Research (grant DZD e.V., FKZ01GI0924) and Center for Regenerative Therapies Dresden, Cluster of Excellence grant FZT 111 (K.K.), Programme Grant from MRC (G.A.); Project IBS-R005-D1 from the Inst. for Basic Science, Korean Ministry of Science (C.D.S.) and Oncosuisse KFS-3169 (L.M.T.).
Author information
Authors and Affiliations
Contributions
L.W. and E.P. conceived and designed the experiments. L.W. performed all of experiments except for re-aggregated thymic organ cultures, which were carried out by C.G.K.; analysis of Treg cells in Foxp3.RFP/GFP mice, which was carried out by S.S. and K.K.; deep sequencing and analysis of TCR clonotypes, which were carried by B.D.S. and E.S.H.; analysis of thymic Treg cells in Foxp3-RFP/Rag-GFP dual reporter mice, which was carried out by N.I.M. and G.A.; analysis of Treg cells in in GF, AF and SPF mice, which was carried out by J.Y.L. and C.D.S.; and evaluation of histological sections, which was carried out by L.M.T. The manuscript was written by L.W. and E.P. All of the authors read the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of GITRintPD-1int Treg cells.
(a) CD4+ T cells from B6 LNs were analyzed for Foxp3, GITR, PD-1 and CD25 protein expression by flow cytometry. Gates (left plot) show Triplehi (GITRhiPD-1hiCD25hi, red), Tripleint (GITRintPD-1intCD25int, grey) and Triplelo Treg cell (GITRloPD-1loCD25lo, brown) frequencies among total CD4+Foxp3+ cells. Histogram and bar graph (right) shows CD25 expression on Triplehi, Tripleint and Triplelo Treg cells (n=4 mice). (b-d) Expression of homing and activation markers on B6 LN Triplehi (red), Tripleint (grey) and Triplelo Treg cells (brown) analyzed by flow cytometry (n= 4 mice). (e) Triplehi (red), Tripleint (grey) and Triplelo (brown) Treg cells were analyzed for CD5 (n=4 mice) and Nur77-GFP (n=2 mice) expression by flow cytometry. (f) In vivo proliferation of Triplehi (red), Tripleint (grey), Triplelo (brown) Treg cells and CD4+ Tconv cells (blue) isolated from B6 LNs. Percentages of proliferating (BrdU+) cells are shown (n=4 mice). Bar graphs show mean ± s.e.m. Data is taken from 2-3 independent experiments.
Supplementary Figure 2 Nrp-1 and Helios expression in Triplehi and Triplelo Treg cells from SPF, GF and AF mice.
Lymph node cells from SPF, germ free (GF) and antigen free (AF) B6 mice were isolated, and Triplehi (red) and Triplelo (brown) Treg cells were analyzed for Nrp-1 (top) and Helios (bottom) expression by flow cytometry. Bar graphs show geometric mean of fluorescence intensity of Nrp-1 or Helios protein expression in Triplehi and Triplelo Treg cells (n=2 mice) NS= not significant (Kruskal-Wallis Test). Bar graphs display mean ± s.e.m. Data is taken from one experiment.
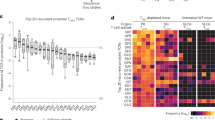
Supplementary Figure 3 Sorting strategy of CD4+ Tconv, Triplehi and Triplelo Treg cells from YAe62β-tg mice.
Splenocytes from YAe62β-tg TCRα+/KO mice were FACS sorted based upon the expression of Foxp3-GFP, PD-1 and GITR. (a) Gating strategy for FACS sorting of magnetically isolated CD4+ splenic populations from YAe62β-tg TCRα+/KO. (b) Frequency of Triplehi and Triplelo Treg cell populations among Foxp3+ CD4+ T cells in the 3 independent sorted samples (2 pooled mice per sample). Bars are the mean of the 3 samples. (c) Representative flow cytometric analysis of the CD4+ Tconv cell population following FACS sorting. (d) Representative flow cytometric analysis of the Triplehi Treg cell population following FACS sorting. (e) Representative flow cytometric analysis of the Triplelo Treg cell population following FACS sorting.
Supplementary Figure 4 Thymic Treg cell development in the presence of negatively selecting peptides.
Antigen affinity influences TCR downstream signaling. (a) Re-aggregate thymic organ cultures were established from B3K508TCR-tg, MHC II KO thymocytes, which are arrested at the CD4+ CD8+ DP stage and thymic epithelial cells from B6 mice. Peptides were added at the following concentrations: 20 μM P-1A (threshold affinity negative selector), 2 μM P2A (intermediate affinity negative selector), and 0.2 μM 3K (high affinity negative selector). After 7d in culture, thymocytes were stained for CD4 and Foxp3 and analyzed by flow cytometry. (b) Histograms shows flow cytometric analysis for CD25 (left) and Helios (right) protein expression of 3BK508TCR-tg Rag2–/– Foxp3+ CD4SP thymocytes, 48h after stimulation with P-1A (1μM, brown), P2A (1μM, grey) or 3K (1μM,red) peptide presented on mature B6 BMDCs in the presence of IL-2 and TGF-β. (c,d) Phosphorylation of CD3ζ, Erk and cJun in 3BK508TCR-tg CD4SP thymocytes stimulated with P-1A (grey line), P2A (brown line), 3K (red line) or no peptide (light grey filled) was determined after 90min, 24 and 48h by flow cytometry. (c) Representative histograms of fluorescence intensity in thymocytes stained for pCD3ζ, pJun and pErk following 24 and 48h of antigen stimulation and (d) geometric mean fluorescence intensity of pCD3ζ, pJun and pErk after 90min, 24h and 48h of peptide stimulation. (n=2 independent experiments)
Supplementary Figure 5 Sorting strategy to obtain Triplehi and Triplelo Treg cells and design of experimental lympho-proliferation.
(a) Magnetic bead enriched CD4+ T cells isolated from pooled LNs of 6-10 week old Foxp3eGFP mice were sorted for CD4+GFP+GITRhiPD-1hi (TriplehiTreg cells) and CD4+GFP+GITRloPD-1lo (TripleloTreg cells). Purity of each population was > 97%. (b) Lymphoproliferation induction by acute Treg cell ablation and B6 Treg cell treatment. 2.5x105 sorted B6 Treg cells, (unaffected by DTx) from pooled B6 LNs were injected intravenously (i.v.) into 6-10 week old Foxp3DTR mice. Three days later, host Foxp3+ cells were depleted intra peritoneal (i.p.) injection of DTx every other day for 10 days.
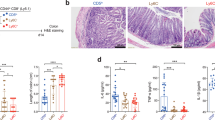
Supplementary Figure 6 Colitis-induction protocol and maintenance of Treg cell phenotype following adoptive transfer.
(a) To induce colitis, 6-10 week old T cell deficient CD3ε–/– mice received 3.2x105 sorted naive CD4+ Tconv cells (CD4+CD25–) isolated from B6 Ly5.1+ mice (B6Tconv). In some groups, 0.8 x105 sorted, Ly5.2+ Triplehi or Triplelo Treg cells from pooled B6 LNs were co-transferred along with Tconv cells. In experiments where iTreg cell generation was inhibited during colitis induction, 6-10 week old T cell deficient CD3ε–/– recipients were first injected i.v. with 3.2x105 sorted, naive CD4 Tconv cells isolated from Foxp3DTR, Ly5.1+ mice (Foxp3DTR Tconv cells). In some groups, 0.8 x105 sorted B6 TripleloTreg cells (unaffected by DTx) were co-transferred along with 3.2x105 sorted naive Foxp3DTRTconv cells. To deplete Tconv cell-derived iTreg cells, recipients were injected intraperitoneal every third day with DTx (10μg/kg). Mice were weighed weekly at the same time of day and were sacrificed when they lost > 20% of their initial body weight or at six weeks following adoptive transfer. (b) Flow cytometry analyzing GITR and PD-1 expression in LN cells isolated from mice that received Triplehi or TripleloTreg cells (described in a.The majority of transferred Triplelow Tregs maintain low PD-1 expression, although GITR expression is increased on Triplelow Tregs after six weeks in this lymphopenic environment. Data is from one experiment representative for four independent experiments with similar results.
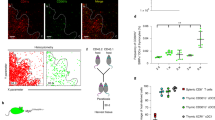
Supplementary Figure 7 Sorting strategy, experimental setup for adoptive transfer of Scurfy disease, pathology of adoptively transferred mice and α4β7 integrin expression on transferred Scurfy cells homing to mLNs.
(a) Sorting strategy to obtain Scurfy Triplehi and Scurfy Triplelo CD4+ T cells. Magnetic bead enriched CD4+ T cells isolated from pooled LNs from 2 week old, male Foxp3-deficient mice were sorted for CD4+GITRhiCD25hi (Scurfy Triplehi CD4+ T cells) and CD4+GITRloCD25lo (Scurfy TripleloCD4+ T cells) T cells. Purity of each population was > 97%. (b) To transfer scurfy disease, 5x105 sorted Scurfy Triplehi or Scurfy Triplelo CD4+ T cells were injected i.v. into 6-10 week old CD3ε–/– recipient mice. Mice were weighed weekly at the same time of day and were sacrificed when they lost > 20% of their initial body weight or at six weeks following adoptive transfer. Photographs of peripheral and mesenteric LNs (c) and skin (tail) and colons (d) from CD3ε–/– mice six weeks after receiving either Scurfy Triplehi or Scurfy Triplelo CD4+ T cells. (e) Flow cytometric analysis of α4β7 integrin expression on Scurfy CD4+ T cells isolated from mLNs of CD3ε–/– mice six weeks after receiving either Scurfy Triplehi (top panel, orange bar) or Scurfy Triplelo (lower panel, purple bar) CD4+ T cells (n=3 mice each group). Data are from one experiment (d). Images for (c, e) are from one experiment representative of 5 independent experiments with similar results. Mean ± s.e.m., *P<0.05 (unpaired, two-tailed t-test).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 1704 kb)
Rights and permissions
About this article
Cite this article
Wyss, L., Stadinski, B., King, C. et al. Affinity for self antigen selects Treg cells with distinct functional properties. Nat Immunol 17, 1093–1101 (2016). https://doi.org/10.1038/ni.3522
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3522
This article is cited by
-
Regulation of activated T cell survival in rheumatic autoimmune diseases
Nature Reviews Rheumatology (2022)
-
PD-L1–PD-1 interactions limit effector regulatory T cell populations at homeostasis and during infection
Nature Immunology (2022)
-
Asymmetric cell division shapes naive and virtual memory T-cell immunity during ageing
Nature Communications (2021)
-
Essential role of a ThPOK autoregulatory loop in the maintenance of mature CD4+ T cell identity and function
Nature Immunology (2021)
-
A mouse model that is immunologically tolerant to reporter and modifier proteins
Communications Biology (2020)