Abstract
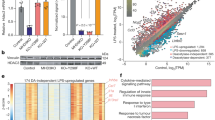
The DNA methyltransferase Dnmt3a has high expression in terminally differentiated macrophages; however, its role in innate immunity remains unknown. Here we report that deficiency in Dnmt3a selectively impaired the production of type I interferons triggered by pattern-recognition receptors (PRRs), but not that of the proinflammatory cytokines TNF and IL-6. Dnmt3a-deficient mice exhibited enhanced susceptibility to viral challenge. Dnmt3a did not directly regulate the transcription of genes encoding type I interferons; instead, it increased the production of type I interferons through an epigenetic mechanism by maintaining high expression of the histone deacetylase HDAC9. In turn, HDAC9 directly maintained the deacetylation status of the key PRR signaling molecule TBK1 and enhanced its kinase activity. Our data add mechanistic insight into the crosstalk between epigenetic modifications and post-translational modifications in the regulation of PRR signaling and activation of antiviral innate immune responses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Smale, S.T., Tarakhovsky, A. & Natoli, G. Chromatin contributions to the regulation of innate immunity. Annu. Rev. Immunol. 32, 489–511 (2014).
Álvarez-Errico, D., Vento-Tormo, R., Sieweke, M. & Ballestar, E. Epigenetic control of myeloid cell differentiation, identity and function. Nat. Rev. Immunol. 15, 7–17 (2015).
Foster, S.L., Hargreaves, D.C. & Medzhitov, R. Gene-specific control of inflammation by TLR-induced chromatin modifications. Nature 447, 972–978 (2007).
Saeed, S. et al. Epigenetic programming of monocyte-to-macrophage differentiation and trained innate immunity. Science 345, 1251086 (2014).
Ramirez-Carrozzi, V.R. et al. Selective and antagonistic functions of SWI/SNF and Mi-2beta nucleosome remodeling complexes during an inflammatory response. Genes Dev. 20, 282–296 (2006).
Zhang, Q. et al. Tet2 is required to resolve inflammation by recruiting Hdac2 to specifically repress IL-6. Nature 525, 389–393 (2015).
Jones, P.A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Anderson, P. Post-transcriptional regulons coordinate the initiation and resolution of inflammation. Nat. Rev. Immunol. 10, 24–35 (2010).
Mowen, K.A. & David, M. Unconventional post-translational modifications in immunological signaling. Nat. Immunol. 15, 512–520 (2014).
Piccirillo, C.A., Bjur, E., Topisirovic, I., Sonenberg, N. & Larsson, O. Translational control of immune responses: from transcripts to translatomes. Nat. Immunol. 15, 503–511 (2014).
Verdin, E. & Ott, M. 50 years of protein acetylation: from gene regulation to epigenetics, metabolism and beyond. Nat. Rev. Mol. Cell Biol. 16, 258–264 (2015).
Cao, W., Bao, C., Padalko, E. & Lowenstein, C.J. Acetylation of mitogen-activated protein kinase phosphatase-1 inhibits Toll-like receptor signaling. J. Exp. Med. 205, 1491–1503 (2008).
Munshi, N. et al. Acetylation of HMG I(Y) by CBP turns off IFN beta expression by disrupting the enhanceosome. Mol. Cell 2, 457–467 (1998).
Nusinzon, I. & Horvath, C.M. Interferon-stimulated transcription and innate antiviral immunity require deacetylase activity and histone deacetylase 1. Proc. Natl. Acad. Sci. USA 100, 14742–14747 (2003).
Fitzgerald, K.A. et al. IKKepsilon and TBK1 are essential components of the IRF3 signaling pathway. Nat. Immunol. 4, 491–496 (2003).
Liu, S. et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science 347, aaa2630 (2015).
Cao, X. Self-regulation and cross-regulation of pattern-recognition receptor signalling in health and disease. Nat. Rev. Immunol. 16, 35–50 (2016).
Wang, C. et al. The E3 ubiquitin ligase Nrdp1 'preferentially' promotes TLR-mediated production of type I interferon. Nat. Immunol. 10, 744–752 (2009).
Li, S., Wang, L., Berman, M., Kong, Y.Y. & Dorf, M.E. Mapping a dynamic innate immunity protein interaction network regulating type I interferon production. Immunity 35, 426–440 (2011).
Cui, J. et al. NLRP4 negatively regulates type I interferon signaling by targeting the kinase TBK1 for degradation via the ubiquitin ligase DTX4. Nat. Immunol. 13, 387–395 (2012).
Wu, C., Macleod, I. & Su, A.I. BioGPS and MyGene.info: organizing online, gene-centric information. Nucleic Acids Res. 41, D561–D565 (2013).
Chen, W. et al. Induction of Siglec-G by RNA viruses inhibits the innate immune response by promoting RIG-I degradation. Cell 152, 467–478 (2013).
Servant, M.J. et al. Identification of the minimal phosphoacceptor site required for in vivo activation of interferon regulatory factor 3 in response to virus and double-stranded RNA. J. Biol. Chem. 278, 9441–9447 (2003).
Chau, T.L. et al. Are the IKKs and IKK-related kinases TBK1 and IKK-ɛ similarly activated? Trends Biochem. Sci. 33, 171–180 (2008).
Ma, X. et al. Molecular basis of Tank-binding kinase 1 activation by transautophosphorylation. Proc. Natl. Acad. Sci. USA 109, 9378–9383 (2012).
Wu, H. et al. Dnmt3a-dependent nonpromoter DNA methylation facilitates transcription of neurogenic genes. Science 329, 444–448 (2010).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Jones, P.L. et al. Methylated DNA and MeCP2 recruit histone deacetylase to repress transcription. Nat. Genet. 19, 187–191 (1998).
Nan, X. et al. Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. Nature 393, 386–389 (1998).
Viré, E. et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439, 871–874 (2006).
Clément, J.F., Meloche, S. & Servant, M.J. The IKK-related kinases: from innate immunity to oncogenesis. Cell Res. 18, 889–899 (2008).
Tu, D. et al. Structure and ubiquitination-dependent activation of TANK-binding kinase 1. Cell Rep. 3, 747–758 (2013).
Wang, L., Li, S. & Dorf, M.E. NEMO binds ubiquitinated TANK-binding kinase 1 (TBK1) to regulate innate immune responses to RNA viruses. PLoS One 7, e43756 (2012).
Wu, H. & Zhang, Y. Reversing DNA methylation: mechanisms, genomics, and biological functions. Cell 156, 45–68 (2014).
Ramirez-Carrozzi, V.R. et al. A unifying model for the selective regulation of inducible transcription by CpG islands and nucleosome remodeling. Cell 138, 114–128 (2009).
Marr, A.K. et al. Leishmania donovani infection causes distinct epigenetic DNA methylation changes in host macrophages. PLoS Pathog. 10, e1004419 (2014).
Rowles, D.L. et al. DNA methyltransferase DNMT3A associates with viral proteins and impacts HSV-1 infection. Proteomics 15, 1968–1982 (2015).
Chen, C. et al. DNA methyltransferases 1 and 3B are required for hepatitis C virus infection in cell culture. Virology 441, 57–65 (2013).
Vivekanandan, P., Daniel, H.D., Kannangai, R., Martinez-Murillo, F. & Torbenson, M. Hepatitis B virus replication induces methylation of both host and viral DNA. J. Virol. 84, 4321–4329 (2010).
Bahar Halpern, K., Vana, T. & Walker, M.D. Paradoxical role of DNA methylation in activation of FoxA2 gene expression during endoderm development. J. Biol. Chem. 289, 23882–23892 (2014).
Zhou, V.W., Goren, A. & Bernstein, B.E. Charting histone modifications and the functional organization of mammalian genomes. Nat. Rev. Genet. 12, 7–18 (2011).
Northrup, D.L. & Zhao, K. Application of ChIP-Seq and related techniques to the study of immune function. Immunity 34, 830–842 (2011).
Shakespear, M.R., Halili, M.A., Irvine, K.M., Fairlie, D.P. & Sweet, M.J. Histone deacetylases as regulators of inflammation and immunity. Trends Immunol. 32, 335–343 (2011).
Aung, H.T. et al. LPS regulates proinflammatory gene expression in macrophages by altering histone deacetylase expression. FASEB J. 20, 1315–1327 (2006).
Halili, M.A. et al. Differential effects of selective HDAC inhibitors on macrophage inflammatory responses to the Toll-like receptor 4 agonist LPS. J. Leukoc. Biol. 87, 1103–1114 (2010).
Nusinzon, I. & Horvath, C.M. Positive and negative regulation of the innate antiviral response and beta interferon gene expression by deacetylation. Mol. Cell. Biol. 26, 3106–3113 (2006).
Zhu, J., Coyne, C.B. & Sarkar, S.N. PKC alpha regulates Sendai virus-mediated interferon induction through HDAC6 and β-catenin. EMBO J. 30, 4838–4849 (2011).
Saul, V.V., Niedenthal, R., Pich, A., Weber, F. & Schmitz, M.L. SUMO modification of TBK1 at the adaptor-binding C-terminal coiled-coil domain contributes to its antiviral activity. Biochim. Biophys. Acta 1853, 136–143 (2015).
Zhao, W. Negative regulation of TBK1-mediated antiviral immunity. FEBS Lett. 587, 542–548 (2013).
Han, C. et al. Integrin CD11b negatively regulates TLR-triggered inflammatory responses by activating Syk and promoting degradation of MyD88 and TRIF via Cbl-b. Nat. Immunol. 11, 734–742 (2010).
Xu, S. et al. Constitutive MHC class I molecules negatively regulate TLR-triggered inflammatory responses via the Fps-SHP-2 pathway. Nat. Immunol. 13, 551–559 (2012).
Zhao, T. et al. The NEMO adaptor bridges the nuclear factor-κB and interferon regulatory factor signaling pathways. Nat. Immunol. 8, 592–600 (2007).
Li, Y. et al. The DNA methylome of human peripheral blood mononuclear cells. PLoS Biol. 8, e1000533 (2010).
Acknowledgements
We thank J. Zhu and N. Wang for mass-spectrometry analysis; Z. Chen (University of Texas Southwestern Medical Center) for the L929-shSTING cell line; B. Sun (Chinese Academy of Sciences) for SeV; and Q. Li (Chinese Academy of Medical Sciences) for HSV-1 virus (Kos strain). Supported by the National Key Basic Research Program of China (2013CB530502) and National Natural Science Foundation of China (31390431, 31370864 and 31200654).
Author information
Authors and Affiliations
Contributions
X. Li, Q.Z. Y.D., Y.L., D.Z., K.Z., Q.S., X. Liu, X.Z., N.L. and Q.W. performed the experiments; Q.Z. established the DNA methylomes and performed bioinformatics analysis; Z.C. did mass-spectrometry analysis; G.F. provided Dnmt3afl/fl mice; X.C. designed and supervised the research; and X.C., X. Li, Q.Z. and Q.W. analyzed data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Dnmt3afl/flLyz2-Cre mice have no abnormal differentiation of macrophages.
(a) Immunoblot analysis of Dnmt3a expression in mouse peritoneal macrophages from Dnmt3afl/flLyz2-cre and Dnmt3afl/fl mice stimulated with LPS for the indicated time. (b) Cell counts of peritoneal macrophages in Dnmt3afl/flLyz2-cre and Dnmt3afl/fl mice. (c and d) FACS analysis of F4/80, CD11b expression in Dnmt3afl/flLyz2-cre and Dnmt3afl/fl peritoneal macrophages. Data are representative of three independent experiments with similar results (a,c,d) or are from three independent experiments (b; means ± s.e.m.).
Supplementary Figure 2 Knockdown of Dnmt3a impairs TBK1-dependent production of type I interferons.
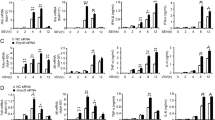
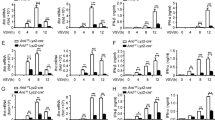
(a) Q-PCR analysis of Dnmt3a mRNA expression in peritoneal macrophages 48hr after transfection with Dnmt3a siRNA. (b) Immunoblot analysis of Dnmt3a expression in peritoneal macrophages treated as in a. (c) Q-PCR analysis of IFN-β, IFN-a, TNF and IL-6 mRNA expression in Dnmt3a-knockdown peritoneal macrophages infected with VSV, SeV, HSV-1 for 8h or analysis of IFN-β, IFN-a, TNF mRNA expression in Dnmt3a-knockdown peritoneal macrophages stimulated with LPS, Poly(I:C), CpG ODN for 2h, or analysis of IL-6 mRNA expression in Dnmt3a-knockdown peritoneal macrophages stimulated with LPS, Poly(I:C), CpG ODN for 4h. (d) ELISA assay of IFN-β, IFN-a, TNF and IL-6 in supernatants of Dnmt3a-knockdown peritoneal macrophages infected with VSV, SeV, HSV-1 for 12h or stimulated with LPS, Poly(I:C), CpG ODN for 8h. (e) Q-PCR analysis of IFN-β, IFN-a mRNA expression in L929 cells stably expressing a shRNA against GFP or STING transfected with a shRNA against Dnmt3a (shDnmt3a) or Control (shCtrl) and 36hr later infected with VSV, SeV, HSV-1 for 8h. * P < 0.01. (two-tailed Student’s t-test). Data are from three independent experiments (a,c,d,e; means ± s.e.m.) or are representative of three independent experiments with similar results (b).
Supplementary Figure 3 Dnmt3a does not directly regulate the transcription of genes encoding type I interferons.
(a) ChIP analysis of the binding of IRF3 or Dnmt3a to the Ifnb1 promoter in peritoneal macrophages infected with VSV for the indicated time. IgG was used as control. (b) MeDIP analysis of the DNA methylation level of the Ifnb1 promoter in peritoneal macrophages infected with VSV for the indicated time. Data are from three independent experiments (means ± s.e.m.).
Supplementary Figure 4 Knockdown of Dnmt3a selectively impairs phosphorylation of IRF3 in response to innate stimuli.
(a,b) Immunoblot analysis of phosphorylated (p-) or total proteins in lysates of Dnmt3a-knockdown peritoneal macrophages infected with VSV or stimulated with LPS for the indicated time. Quantification of relative the protein presented levels is shown in the bottom panel. Numbers below lanes (top) indicate densitometry of the protein presented relative to β-actin expression in that same lane (below). (c,d) Immunoblot analysis of phosphorylated (p-) or total proteins in Dnmt3afl/flLyz2-cre and Dnmt3afl/fl peritoneal macrophages infected with VSV or stimulated with LPS for the indicated time. (e,f) Immunoblot analysis of phosphorylated (p-) or total proteins in lysates of Dnmt3a-knockdown peritoneal macrophages infected with VSV or stimulated with LPS for the indicated time. Data are representative of three independent experiments with similar results.
Supplementary Figure 5 Knockdown of HDAC9 does not inhibit the expression of TNF mRNA or IL-6 mRNA in macrophages.
(a) Q-PCR analysis of HDAC9 mRNA expression in peritoneal macrophages transfected with HDAC9 siRNA. (b) Immunoblot analysis of HDAC9 expression in peritoneal macrophages transfected as in (a). (c) Q-PCR analysis of IFN-β, TNF and IL-6 mRNA expression in HDAC9-knockdown peritoneal macrophages infected with VSV, or stimulated with LPS, cGAMP for the indicated time. * P < 0.01. (two-tailed Student’s t-test). Data are from three independent experiments (a,c; means ± s.e.m.) or are representative of three independent experiments with similar results (b).
Supplementary Figure 6 Dnmt3a does not affect the polyubiquitination of TBK1 in macrophages.
(a) Immunoblot analysis of K63-linked polyubiquitination of endogenous TBK1 in Dnmt3a-deficient and control peritoneal macrophages infected with VSV for 4 hours. (b) Immunoblot analysis of polyubiquitination of TBK1 in the HEK293T cells transiently transfected for 48 h with various combinations of tagged vectors (above lanes), followed by immunoprecipitation with anti-Flag or anti-HA agarose or analysis without immunoprecipitation. Data are representative of three independent experiments with similar results.
Supplementary Figure 7 Specificity of the antibody to TBK1 acetylated at Lys241.
Dot immunoblot analysis of K241 acetylation of TBK1 by using antibody specifically recognizing indicated peptide with acetylated K241 of TBK1 (TBK1 K241 Ace peptide) or pan anti-acetyl lysine antibody. Dot immunoblot analysis of non-acetylated K241 of TBK1 by using antibody recognizing the indicated peptide (TBK1 K241 control peptide) is the loading control. Data are representative of three independent experiments with similar results.
Supplementary Figure 8 Working model of Dnmt3a in promoting the production of type I interferons.
Dnmt3a enhances TLR3/4 and virus-induced type I IFN production by increasing TBK1 kinase activity through upregulating HDAC9 and promoting HDAC9-mediated deacetylation of TBK1.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 6–9 (PDF 1766 kb)
Supplementary Table 1
Genes with significant expression variation in Dnmt3a-knockdown peritoneal macrophages in microarray data (XLSX 27 kb)
Supplementary Table 2
Genes with significant expression variation in Dnmt3a-deficient peritoneal macrophages in RNA-seq data (XLSX 9 kb)
Supplementary Table 3
DMR location (XLSX 39 kb)
Supplementary Table 4
Genes with significant expression variation in HDAC9-knockdown peritoneal macrophages in microarray data (XLSX 19 kb)
Supplementary Table 5
Mass spectrometry analysis of lysine acetylated residues of mouse TBK1 in Dnmt3a-deficient peritoneal macrophages infected with VSV (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Li, X., Zhang, Q., Ding, Y. et al. Methyltransferase Dnmt3a upregulates HDAC9 to deacetylate the kinase TBK1 for activation of antiviral innate immunity. Nat Immunol 17, 806–815 (2016). https://doi.org/10.1038/ni.3464
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3464
This article is cited by
-
HDAC9-mediated epithelial cell cycle arrest in G2/M contributes to kidney fibrosis in male mice
Nature Communications (2023)
-
The HDAC inhibitor domatinostat induces type I interferon α in Merkel cell carcinoma by HES1 repression
Journal of Cancer Research and Clinical Oncology (2023)
-
TRIM18 is a critical regulator of viral myocarditis and organ inflammation
Journal of Biomedical Science (2022)
-
The role of TBK1 in cancer pathogenesis and anticancer immunity
Journal of Experimental & Clinical Cancer Research (2022)
-
Donor clonal hematopoiesis increases risk of acute graft versus host disease after matched sibling transplantation
Leukemia (2022)