Abstract
Innate lymphoid cells (ILCs) are increasingly appreciated as important participants in homeostasis and inflammation. Substantial plasticity and heterogeneity among ILC populations have been reported. Here we have delineated the heterogeneity of human ILCs through single-cell RNA sequencing of several hundreds of individual tonsil CD127+ ILCs and natural killer (NK) cells. Unbiased transcriptional clustering revealed four distinct populations, corresponding to ILC1 cells, ILC2 cells, ILC3 cells and NK cells, with their respective transcriptomes recapitulating known as well as unknown transcriptional profiles. The single-cell resolution additionally divulged three transcriptionally and functionally diverse subpopulations of ILC3 cells. Our systematic comparison of single-cell transcriptional variation within and between ILC populations provides new insight into ILC biology during homeostasis, with additional implications for dysregulation of the immune system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
17 March 2016
In the version of this article initially published, two labels along the horizontal axis of Figure 1c were switched, so the data for donor a were presented for donor c (and vice versa). The error has been corrected in the HTML and PDF versions of the article.
References
Eberl, G. et al. An essential function for the nuclear receptor RORγ(t) in the generation of fetal lymphoid tissue inducer cells. Nat. Immunol. 5, 64–73 (2004).
Mebius, R.E., Rennert, P. & Weissman, I.L. Developing lymph nodes collect CD4+CD3−LTβ+ cells that can differentiate to APC, NK cells, and follicular cells but not T or B cells. Immunity 7, 493–504 (1997).
Cupedo, T. et al. Human fetal lymphoid tissue-inducer cells are interleukin 17-producing precursors to RORC+CD127+ natural killer-like cells. Nat. Immunol. 10, 66–74 (2009).
Artis, D. & Spits, H. The biology of innate lymphoid cells. Nature 517, 293–301 (2015).
Bernink, J.H. et al. Human type 1 innate lymphoid cells accumulate in inflamed mucosal tissues. Nat. Immunol. 14, 221–229 (2013).
Crellin, N.K., Trifari, S., Kaplan, C.D., Cupedo, T. & Spits, H. Human NKp44+IL-22+ cells and LTi-like cells constitute a stable RORC+ lineage distinct from conventional natural killer cells. J. Exp. Med. 207, 281–290 (2010).
Mjösberg, J.M. et al. Human IL-25- and IL-33-responsive type 2 innate lymphoid cells are defined by expression of CRTH2 and CD161. Nat. Immunol. 12, 1055–1062 (2011).
Spits, H. et al. Innate lymphoid cells–a proposal for uniform nomenclature. Nat. Rev. Immunol. 13, 145–149 (2013).
Bernink, J.H. et al. Interleukin-12 and -23 control plasticity of cd127+ group 1 and group 3 innate lymphoid cells in the intestinal lamina propria. Immunity 43, 146–160 (2015).
Klose, C.S. et al. Differentiation of type 1 ILCs from a common progenitor to all helper-like innate lymphoid cell lineages. Cell 157, 340–356 (2014).
Constantinides, M.G., McDonald, B.D., Verhoef, P.A. & Bendelac, A. A committed precursor to innate lymphoid cells. Nature 508, 397–401 (2014).
Glatzer, T. et al. RORγt+ innate lymphoid cells acquire a proinflammatory program upon engagement of the activating receptor NKp44. Immunity 38, 1223–1235 (2013).
Boyd, A., Ribeiro, J.M. & Nutman, T.B. Human CD117 (cKit)+ innate lymphoid cells have a discrete transcriptional profile at homeostasis and are expanded during filarial infection. PLoS ONE 9, e108649 (2014).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Deng, Q., Ramskold, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014).
Treutlein, B. et al. Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375 (2014).
Patel, A.P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Shalek, A.K. et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498, 236–240 (2013).
Mahata, B. et al. Single-cell RNA sequencing reveals T helper cells synthesizing steroids de novo to contribute to immune homeostasis. Cell Rep. 7, 1130–1142 (2014).
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013).
Krijthe, J. Rtsne: T-distributed stochastic neighbor embedding using Barnes-Hut implementation. R package version 0.9 (http://CRAN.R-project.org/package=Rtsne).
Taniguchi, Y. et al. Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329, 533–538 (2010).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Robinette, M.L. et al. Transcriptional programs define molecular characteristics of innate lymphoid cell classes and subsets. Nat. Immunol. 16, 306–317 (2015).
van de Pavert, S.A. et al. Maternal retinoids control type 3 innate lymphoid cells and set the offspring immunity. Nature 508, 123–127 (2014).
Bezman, N.A. et al. Molecular definition of the identity and activation of natural killer cells. Nat. Immunol. 13, 1000–1009 (2012).
Wang, F., Tian, Z. & Wei, H. Genomic expression profiling of NK cells in health and disease. Eur. J. Immunol. 45, 661–678 (2015).
Holmes, M.L. et al. Peripheral natural killer cell maturation depends on the transcription factor Aiolos. EMBO J. 33, 2721–2734 (2014).
Barnig, C. et al. Lipoxin A4 regulates natural killer cell and type 2 innate lymphoid cell activation in asthma. Sci. Transl. Med. 5, 174ra126 (2013).
Xue, L. et al. Prostaglandin D2 activates group 2 innate lymphoid cells through chemoattractant receptor-homologous molecule expressed on TH2 cells. J. Allergy Clin. Immunol. 133, 1184–1194 (2014).
Gentek, R. et al. Modulation of signal strength switches Notch from an inducer of T Cells to an inducer of ILC2. Front. Immunol. 4, 334 (2013).
Mielke, L.A. et al. TCF-1 controls ILC2 and NKp46+RORγt+ innate lymphocyte differentiation and protection in intestinal inflammation. J. Immunol. 191, 4383–4391 (2013).
Yang, Q. et al. T cell factor 1 is required for group 2 innate lymphoid cell generation. Immunity 38, 694–704 (2013).
Larabee, J.L., Shakir, S.M., Barua, S. & Ballard, J.D. Increased cAMP in monocytes augments Notch signaling mechanisms by elevating RBP-J and transducin-like enhancer of Split (TLE). J. Biol. Chem. 288, 21526–21536 (2013).
Bandyopadhyay, S., Valdor, R. & Macian, F. Tle4 regulates epigenetic silencing of gamma interferon expression during effector T helper cell tolerance. Mol. Cell. Biol. 34, 233–245 (2014).
Fuchs, A. et al. Intraepithelial type 1 innate lymphoid cells are a unique subset of IL-12- and IL-15-responsive IFN-γ-producing cells. Immunity 38, 769–781 (2013).
Possot, C. et al. Notch signaling is necessary for adult, but not fetal, development of RORγt+ innate lymphoid cells. Nat. Immunol. 12, 949–958 (2011).
Lee, J.S. et al. AHR drives the development of gut ILC22 cells and postnatal lymphoid tissues via pathways dependent on and independent of Notch. Nat. Immunol. 13, 144–151 (2012).
Hoorweg, K. et al. Functional differences between human NKp44− and NKp44+ RORC+ innate lymphoid cells. Front. Immunol. 3, 72 (2012).
Hepworth, M.R. et al. Group 3 innate lymphoid cells mediate intestinal selection of commensal bacteria-specific CD4+ T cells. Science 348, 1031–1035 (2015).
Hepworth, M.R. et al. Innate lymphoid cells regulate CD4+ T-cell responses to intestinal commensal bacteria. Nature 498, 113–117 (2013).
Roederer, M., Nozzi, J.L. & Nason, M.C. SPICE: exploration and analysis of post-cytometric complex multivariate datasets. Cytometry 79, 167–174 (2011).
De Smedt, M. et al. Notch signaling induces cytoplasmic CD3ɛ expression in human differentiating NK cells. Blood 110, 2696–2703 (2007).
Huang, Y. et al. IL-25-responsive, lineage-negative KLRG1hi cells are multipotential 'inflammatory' type 2 innate lymphoid cells. Nat. Immunol. 16, 161–169 (2015).
Rohland, N. & Reich, D. Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res. 22, 939–946 (2012).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Ramsköld, D., Wang, E.T., Burge, C.B. & Sandberg, R. An abundance of ubiquitously expressed genes revealed by tissue transcriptome sequence data. PLoS Comput. Biol. 5, e1000598 (2009).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185 (2012).
Leek, J.T., Johnson, W.E., Parker, H.S., Jaffe, A.E. & Storey, J.D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Suzuki, R. & Shimodaira, H. Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22, 1540–1542 (2006).
Kharchenko, P.V., Silberstein, L. & Scadden, D.T. Bayesian approach to single-cell differential expression analysis. Nat. Methods 11, 740–742 (2014).
Wickham, H. The split-apply-combine strategy for data analysis. J. Stat. Softw. 40, 1–29 (2011).
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis (Springer New York, 2009).
Neuwirth, E. RColorBrewer: ColorBrewer palettes. R package version 1.0–5. http://CRAN.R-project.org/package=RColorBrewer (2011).
Warnes, G.R. et al. gplots: Various R programming tools for plotting data. R package version 2.14.2. http://CRAN.R-project.org/package=gplots (2014).
Acknowledgements
We thank I. Douagi and R. Månsson for support with FACSAria sorting; J. Michaelsson, M. Karlsson and Y. Bryceson for discussions and critical reading of the manuscript; and M. Holm (Uppsala University) for Python scripts and input on coding. Supported by the Karolinska Institutet (J.M., M.F. and J.T.), the Swedish Research Council (J.M., M.F. and R.S.), the Swedish Cancer Society (J.M. and M.F.), the Swedish Society for Medical Research (J.M. and M.F.), the Swedish Foundation for Strategic Research (J.M., M.F. and R.S.), Torsten Söderberg's Foundation (J.M. and M.F.), the Jonas Söderquist Foundation (J.M. and M.F.), the European Union's Horizon 2020 research and innovation program (Marie Sklodowska-Curie 655677 for V.K.) and the Stockholm County Government (J.T.).
Author information
Authors and Affiliations
Contributions
Å.K.B. contributed to study design, performed the computational analyses of transcriptome data, analyzed and interpreted data and co-wrote the manuscript; M.F. designed the study, performed experiments, analyzed and interpreted data and co-wrote the manuscript; S.P. contributed to study design, generated all scRNA-seq libraries, interpreted data and contributed to manuscript writing; V.K. performed experiments, analyzed and interpreted data and contributed to manuscript writing; J.T. analyzed and interpreted data and contributed to manuscript writing; D.F. provided clinical samples, interpreted clinical data and contributed to manuscript writing; R.S. contributed to study design, supervised the computational analyses, interpreted data and co-wrote the manuscript; and J.M. designed the study, performed experiments, interpreted data and co-wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Overview of quality control (QC) filtering.
Histograms showing number of reads (a), percent uniquely mapping reads (b), fraction mismatches (c), fraction exon mapping reads (d), fraction of reads mapping to a region at the 10% most 3prime end of each transcript (e) and number of mRNA reads (f). Blue bars represents empty well controls and red bars represent wells containing one cell. Black bold lines indicates filtering cutoffs where cells above/below the black line have been removed. Data were generated in three independent experiments with one tonsil donor each (n=798).
Supplementary Figure 2 Overview of scRNA-seq data filtering and normalization.
a) Detection of biologically variable transcripts over technical noise with ERCC spike-in RNAs highlighted in black, human transcripts in blue (variable) or red (non-variable). Violin plots showing distributions of: b) Forward scattering (FSC), c) ratio of cell RNA to ERCC spike-in RNA (ERCC-ratio) and d) number of detected transcripts. e) PCA based on 847 variable transcripts, before (upper panel) and after (lower panel) batch normalization, cells colored either by surface phenotype (left panel) or donor (left panel) origin. Data were generated in three independent experiments with one tonsil donor each (total number of cells per cell population: ILC1, n=112; ILC2, n=143; ILC3, n=320; NK cells, n=73).
Supplementary Figure 3 Pairwise comparisons of scRNA-seq profiles.
a) Pairwise comparison of the mean expression of the top 50 differentially expressed transcripts for the two cell populations in question. Cells are colored according to cluster definition described in main Fig. 2. Cells where cluster definition and surface phenotype were in agreement are shown with a star (*). Cells that deviated in the clustering are either highlighted as cross (x) if they were defined as the other phenotype in the comparison, and as a circle (o) if the cell had another phenotype than the 2 cell populations in question. b) ERCC-ratio plots for each plot shown in a demonstrating the ratio of cellular RNA to ERCC spike-in RNA for each cell. Data were generated in three independent experiments with one tonsil donor each. Total number of cells per cell population: ILC1, n=112; ILC2, n=143; ILC3, n=320; NK cells, n=73.
Supplementary Figure 4 Comparison of protein expression versus RNA expression.
Each plot shows the surface protein expression intensity (y-axis) vs. log2(RPKM) RNA expression (x-axis) as measured by flow cytometry (indexed flow cytometric sorting data collection) and scRNA-seq, respectively. Both quantities were normalized on a scale from 0-1. Correlation values, both Spearman and Pearson are shown in the titles. Data were generated in three independent experiments with one tonsil donor each (n=648).
Supplementary Figure 5 Transcripts commonly expressed by CD127+ ILCs and NK cells.
Violin plots with expression distribution in each cell population on log2(rpkm) scale for ILC and NK specific transcripts. Coloring according to mean expression. a) Other transcripts commonly expressed by ILCs (according to SCDE; multiple-testing corrected p-value < 0.001). b) Transcription factors commonly expressed by ILCs (according to SCDE; multiple-testing corrected p-value < 0.05). c) Transcripts known to be expressed by NK cells d) Other transcripts expressed by NK cells (according to SCDE; multipletesting corrected p-value < 0.001 for both c and d). Data were generated in three independent experiments with one tonsil donor each (total number of cells per cell population: ILC1, n=112; ILC2, n=143; ILC3, n=320; NK cells, n=73).
Supplementary Figure 6 T cell–related transcripts and detection of differentially expressed genes.
a) Heatmap of T-cell related transcripts, expression in ILC1s (blue), ILC2s (cyan), NK cells (green) and ILC3s (red). Each vertical line in the heatmap represents the expression intensity in an individual cell. Color intensities according to log2(rpkm) values. b) Number of significantly differentially expressed genes (p-value < 0.001) in ILC1s (blue), ILC2s (cyan) and ILC3s (red) with random subsampling of 25,50,75 or 100 cells from each population. Error bars represents standard deviation from 10 iterations. Data were generated in three independent experiments with one tonsil donor each (total number of cells per cell population: ILC1, n=112; ILC2, n=143; ILC3, n=320; NK cells, n=73).
Supplementary Figure 7 Transcripts expressed in ILC3 subpopulations.
a) Top 20 transcript loadings for principal components 1 (PC1) and 2 (PC2) in PCA with ILC3s (Fig. 8a). b) t-SNE plots with the ILC3s colored according to expression intensity of some selected transcripts. Data were generated in three independent experiments with one tonsil donor each (n=320).
Supplementary Figure 8 Flow cytometry of ILC3 subpopulations.
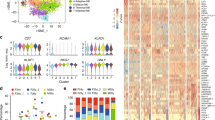
a) t-SNE plots based on surface marker intensities for 5 adult donors and 7 pediatric donors (n=20543) with intensities of the 4 markers used for t-SNE (NKp44, HLA-DR, CD62L and CD45RA) followed by donor distribution (red to yellow shades for pediatric, blue to green shades for adults). b) Bar charts show percentage of IL-2+, IL-22+, IL-17F+ and TNF+ cells from the indicated ILC3 subpopulations after IL-23 plus IL-1β (50 ng/ml each, for 12+6 hours) and/or PMA plus ionomycin (20 ng/ml plus 0.5 μM, for the last 6 hours). Bars show mean and SEM from 4-6 donors. **** p<0.0001, *** p<0.005, ** p<0.01 and * p<0.05 as calculated using oneway ANOVA and Tukey’s multi-comparisons test. c) Representative dot plots show intracellular IL-2 and IL-22 production by the different ILC3 subpopulations after the indicated stimulations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Table 1 (PDF 4224 kb)
Supplementary Data Set 1
Differentially regulated genes per ILC population (XLSX 10975 kb)
Supplementary Data Set 2
List of genes correlated to RORC and GATA3 (XLSX 53 kb)
Rights and permissions
About this article
Cite this article
Björklund, Å., Forkel, M., Picelli, S. et al. The heterogeneity of human CD127+ innate lymphoid cells revealed by single-cell RNA sequencing. Nat Immunol 17, 451–460 (2016). https://doi.org/10.1038/ni.3368
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3368
This article is cited by
-
Group 2 innate lymphoid cells and their surrounding environment
Inflammation and Regeneration (2023)
-
Group 3 innate lymphoid cells secret neutrophil chemoattractants and are insensitive to glucocorticoid via aberrant GR phosphorylation
Respiratory Research (2023)
-
Graphene oxide elicits microbiome-dependent type 2 immune responses via the aryl hydrocarbon receptor
Nature Nanotechnology (2023)
-
Immunomodulatory Biomaterials and Emerging Analytical Techniques for Probing the Immune Micro-Environment
Tissue Engineering and Regenerative Medicine (2023)
-
Single-cell RNA-seq of out-of-thaw mesenchymal stromal cells shows tissue-of-origin differences and inter-donor cell-cycle variations
Stem Cell Research & Therapy (2021)