Abstract
Intestinal T cells and group 3 innate lymphoid cells (ILC3 cells) control the composition of the microbiota and gut immune responses. Within the gut, ILC3 subsets coexist that either express or lack the natural cytoxicity receptor (NCR) NKp46. We identified here the transcriptional signature associated with the transcription factor T-bet–dependent differentiation of NCR− ILC3 cells into NCR+ ILC3 cells. Contrary to the prevailing view, we found by conditional deletion of the key ILC3 genes Stat3, Il22, Tbx21 and Mcl1 that NCR+ ILC3 cells were redundant for the control of mouse colonic infection with Citrobacter rodentium in the presence of T cells. However, NCR+ ILC3 cells were essential for cecal homeostasis. Our data show that interplay between intestinal ILC3 cells and adaptive lymphocytes results in robust complementary failsafe mechanisms that ensure gut homeostasis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Dudakov, J.A., Hanash, A.M. & van den Brink, M.R. Interleukin-22: immunobiology and pathology. Annu. Rev. Immunol. 33, 747–785 (2015).
Spits, H. et al. Innate lymphoid cells–a proposal for uniform nomenclature. Nat. Rev. Immunol. 13, 145–149 (2013).
McKenzie, A.N., Spits, H. & Eberl, G. Innate lymphoid cells in inflammation and immunity. Immunity 41, 366–374 (2014).
Diefenbach, A., Colonna, M. & Koyasu, S. Development, differentiation, and diversity of innate lymphoid cells. Immunity 41, 354–365 (2014).
Montaldo, E., Juelke, K. & Romagnani, C. Group 3 innate lymphoid cells (ILC3s): Origin, differentiation, and plasticity in humans and mice. Eur. J. Immunol. 45, 2171–2182 (2015).
Eberl, G., Di Santo, J.P. & Vivier, E. The brave new world of innate lymphoid cells. Nat. Immunol. 16, 1–5 (2015).
Eberl, G., Colonna, M., Di Santo, J.P. & McKenzie, A.N. Innate lymphoid cells: a new paradigm in immunology. Science 348, aaa6566 (2015).
Mebius, R.E., Rennert, P. & Weissman, I.L. Developing lymph nodes collect CD4+CD3− LTβ+ cells that can differentiate to APC, NK cells, and follicular cells but not T or B cells. Immunity 7, 493–504 (1997).
Constantinides, M.G. et al. PLZF expression maps the early stages of ILC1 lineage development. Proc. Natl. Acad. Sci. USA 112, 5123–5128 (2015).
Klose, C.S. et al. A T-bet gradient controls the fate and function of CCR6-RORγt+ innate lymphoid cells. Nature 494, 261–265 (2013).
Rankin, L.C. et al. The transcription factor T-bet is essential for the development of NKp46+ innate lymphocytes via the Notch pathway. Nat. Immunol. 14, 389–395 (2013).
Kirchberger, S. et al. Innate lymphoid cells sustain colon cancer through production of interleukin-22 in a mouse model. J. Exp. Med. 210, 917–931 (2013).
Satoh-Takayama, N. et al. Microbial flora drives interleukin 22 production in intestinal NKp46+ cells that provide innate mucosal immune defense. Immunity 29, 958–970 (2008).
Cella, M. et al. A human natural killer cell subset provides an innate source of IL-22 for mucosal immunity. Nature 457, 722–725 (2009).
Sonnenberg, G.F., Fouser, L.A. & Artis, D. Functional biology of the IL-22-IL-22R pathway in regulating immunity and inflammation at barrier surfaces. Adv. Immunol. 107, 1–29 (2010).
Zheng, Y. et al. Interleukin-22 mediates early host defense against attaching and effacing bacterial pathogens. Nat. Med. 14, 282–289 (2008).
Satoh-Takayama, N. et al. The chemokine receptor CXCR6 controls the functional topography of interleukin-22 producing intestinal innate lymphoid cells. Immunity 41, 776–788 (2014).
Artis, D. & Spits, H. The biology of innate lymphoid cells. Nature 517, 293–301 (2015).
van de Pavert, S.A. & Vivier, E. Differentiation and function of group 3 innate lymphoid cells, from embryo to adult. Int. Immunol. doi:10.1093/intimm/dxv052 (22 September 2015).
Robinette, M.L. et al. Transcriptional programs define molecular characteristics of innate lymphoid cell classes and subsets. Nat. Immunol. 16, 306–317 (2015).
Sciumé, G. et al. Distinct requirements for T-bet in gut innate lymphoid cells. J. Exp. Med. 209, 2331–2338 (2012).
Guo, X. et al. Induction of innate lymphoid cell-derived interleukin-22 by the transcription factor STAT3 mediates protection against intestinal infection. Immunity 40, 25–39 (2014).
Sathe, P. et al. Innate immunodeficiency following genetic ablation of Mcl1 in natural killer cells. Nat. Commun. 5, 4539 (2014).
Qiu, J. et al. Group 3 innate lymphoid cells inhibit T-cell-mediated intestinal inflammation through aryl hydrocarbon receptor signaling and regulation of microflora. Immunity 39, 386–399 (2013).
Guo, X. et al. Innate lymphoid cells control early colonization resistance against intestinal pathogens through ID2-dependent regulation of the microbiota. Immunity 42, 731–743 (2015).
Basu, R. et al. Th22 cells are an important source of IL-22 for host protection against enteropathogenic bacteria. Immunity 37, 1061–1075 (2012).
Ahlfors, H. et al. IL-22 fate reporter reveals origin and control of IL-22 production in homeostasis and infection. J. Immunol. 193, 4602–4613 (2014).
Vonarbourg, C. et al. Regulated expression of nuclear receptor RORγt confers distinct functional fates to NK cell receptor-expressing RORγt+ innate lymphocytes. Immunity 33, 736–751 (2010).
Dhiman, R. et al. IL-22 produced by human NK cells inhibits growth of Mycobacterium tuberculosis by enhancing phagolysosomal fusion. J. Immunol. 183, 6639–6645 (2009).
Guo, H. & Topham, D.J. Interleukin-22 (IL-22) production by pulmonary Natural killer cells and the potential role of IL-22 during primary influenza virus infection. J. Virol. 84, 7750–7759 (2010).
Male, V. et al. Immature NK cells, capable of producing IL-22, are present in human uterine mucosa. J. Immunol. 185, 3913–3918 (2010).
Tang, Q. et al. Development of IL-22-producing NK lineage cells from umbilical cord blood hematopoietic stem cells in the absence of secondary lymphoid tissue. Blood 117, 4052–4055 (2011).
Xu, X. et al. Conventional NK cells can produce IL-22 and promote host defense in Klebsiella pneumoniae pneumonia. J. Immunol. 192, 1778–1786 (2014).
Macho-Fernandez, E. et al. Lymphotoxin β receptor signaling limits mucosal damage through driving IL-23 production by epithelial cells. Mucosal Immunol. 8, 403–413 (2015).
Sonnenberg, G.F. et al. Innate lymphoid cells promote anatomical containment of lymphoid-resident commensal bacteria. Science 336, 1321–1325 (2012).
Sonnenberg, G.F., Monticelli, L.A., Elloso, M.M., Fouser, L.A. & Artis, D. CD4+ lymphoid tissue-inducer cells promote innate immunity in the gut. Immunity 34, 122–134 (2011).
Hepworth, M.R. et al. Innate lymphoid cells regulate CD4+ T-cell responses to intestinal commensal bacteria. Nature 498, 113–117 (2013).
Gasteiger, G. & Rudensky, A.Y. Interactions between innate and adaptive lymphocytes. Nat. Rev. Immunol. 14, 631–639 (2014).
Wiles, S. et al. Organ specificity, colonization and clearance dynamics in vivo following oral challenges with the murine pathogen Citrobacter rodentium. Cell. Microbiol. 6, 963–972 (2004).
Smith, H.F. et al. Comparative anatomy and phylogenetic distribution of the mammalian cecal appendix. J. Evol. Biol. 22, 1984–1999 (2009).
Frisch, M., Pedersen, B.V. & Andersson, R.E. Appendicitis, mesenteric lymphadenitis, and subsequent risk of ulcerative colitis: cohort studies in Sweden and Denmark. Br. Med. J. 338, b716 (2009).
Alonzi, T. et al. Essential role of STAT3 in the control of the acute-phase response as revealed by inducible gene inactivation [correction of activation] in the liver. Mol. Cell. Biol. 21, 1621–1632 (2001).
Eberl, M., Jomaa, H. & Hayday, A.C. Integrated immune responses to infection - cross-talk between human γδ T cells and dendritic cells. Immunology 112, 364–368 (2004).
Nieuwenhuis, E.E. et al. Disruption of T helper 2-immune responses in Epstein-Barr virus-induced gene 3-deficient mice. Proc. Natl. Acad. Sci. USA 99, 16951–16956 (2002).
Narni-Mancinelli, E. et al. Fate mapping analysis of lymphoid cells expressing the NKp46 cell surface receptor. Proc. Natl. Acad. Sci. USA 108, 18324–18329 (2011).
Thakur, C. et al. Increased expression of mdig predicts poorer survival of the breast cancer patients. Gene 535, 218–224 (2014).
Liao, Y., Smyth, G.K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Ritchie, M.E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, 3 (2004).
McCarthy, D.J. & Smyth, G.K. Testing significance relative to a fold-change threshold is a TREAT. Bioinformatics 25, 765–771 (2009).
Yu, Z. & Morrison, M. Improved extraction of PCR-quality community DNA from digesta and fecal samples. Biotechniques 36, 808–812 (2004).
Acknowledgements
We thank M. Camilleri, J. Vella, E. Loza and L. Inglis and the staff of the animal and flow cytometry facilities of The Walter and Eliza Hall Institute of Medical Research and the Centre d'Immunologie de Marseille-Luminy for technical assistance; S. Kaech (Yale University) for mice with loxP-flanked alleles encoding T-bet; and L. Chasson and C. Laprie (Centre d'Immunologie de Marseille-Luminy) for expertise in histology and for assigning scores to tissue sections. Supported by the National Health and Medical Research Council of Australia (G.T.B., Se.C. and N.H.; Dora Lush Postgraduate Research Scholarship to L.C.R.); the Australian Research Council (Future Fellowship to G.T.B.); the Victorian State Government Operational Infrastructure Support and Australian Government National Health and Medical Research Council Independent Research Institutes Infrustructure Support; the European Research Council (THINK Advanced Grant), the Ligue Nationale contre le Cancer (Equipe Labellisée), INSERM, CNRS and Aix-Marseille University to Centre d'Immunologie de Marseille-Luminy (all for the E.V. laboratory); and the Institut Universitaire de France (E.V.).
Author information
Authors and Affiliations
Contributions
C.S., L.A.M. and Y.K. contributed equally to this work; L.C.R., M.J.H.G.-M., C.S., L.A.M. and Y.K. designed the research, performed experiments and analyzed the data; A.F., E.W., S.M., L.M., J.G., T.P. and A.T.P. performed experiments and analyzed the data; W.S. and G.K.S. performed bioinformatics analyses of RNA-seq data; O.L., D.D., M.L., J.-C.R., C.A., S.U., Se.C. and N.D.H. provided reagents; and G.T.B. and E.V. designed the research, supervised the study, and wrote the manuscript with the help of the other co-authors.
Corresponding authors
Ethics declarations
Competing interests
E.V. is a cofounder and shareholder of Innate Pharma, and T.S. and C.A. are employees of Sanofi.
Integrated supplementary information
Supplementary Figure 1 Characterization of Il22eGFP/eGFPNcr1-iCre mice.
(a) Schematic representation of the Il22eGFP allele. (b) Spleen and liver, (c) small intestine and colon were isolated from Il22eGFP/eGFPNcr1-iCre mice. The frequency of major lymphoid populations was assessed by flow cytometry. Data shown are from (b) one of two independent experiments (n = 4 mice per group in each) and (c) 2 pooled independent experiments (n=4-8 mice per group in total). (d) Small intestinal lymphocytes were stimulated in vitro with IL-23 for 4-5 hr in the presence of brefeldin A to elicit IL-22 production. The frequency of IL-22+ NCR− ILC3 was assessed by flow cytometry. Data shown are from one of two independent experiments (n = 3 mice per group in each). Statistical analyses were performed with unpaired Student’s t-test. n.s., not statistically significant (P > 0.05)
Supplementary Figure 2 IL-22 production by ILC3 subsets in Stat3fl/flNcr1-iCre mice.
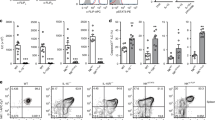
Male Stat3fl/flNcr1+/+ and Stat3fl/flNcr1-iCre mice were orally inoculated with 2 × 109 CFU of C. rodentium. Histograms show total numbers (left) and frequency (right) of IL-22 producing Lin−CD45+NCR+ ILC3 and Lin−CD45+NCR− ILC3 from the small intestine of Stat3fl/flNcr1+/+Rorc+/GFP and Stat3fl/flNcr1-iCreRorc+/GFP mice on day 8 of C. rodentium infection. Cells were stimulated in vitro with IL-23 for 4-5 hr in the presence of brefeldin A to elicit IL-22 production. Data shown are from one of two independent experiments (n=3-4 mice per group in each) and the mean ± s.d. values are shown. Statistical analyses were performed with unpaired Student’s t-test. n.s., not statistically significant (P > 0.05), **P < 0.01 and ***P < 0.001.
Supplementary Figure 3 C. rodentium infection in Tbx21fl/flNcr1-iCre mice.
(a) Numbers of Lin−CD45+ small intestine lymphocytes among naïve Tbx21fl/flNcr1-iCre and their respective controls. Counts were calculated from flow cytometry plots of intracellular RORγt and surface NKp46 expression. Numbers show the cell counts within the indicated gate. Data were pooled from 2 independent experiments (n = 3-4 mice per group in each) and the mean ± s.d. values are shown. Statistical analyses were performed with unpaired Student’s t-test. n.s., not statistically significant (P > 0.05), ***P < 0.001 and ****P < 0.0001 (b–d) Control and Tbx21fl/flNcr1-iCre mice were orally inoculated with 2 × 109 CFU of C. rodentium. (b) H&E staining of formalin-fixed paraffin-embedded sections (left micrographs) and histological scores (right histogram) for colons from wild-type and Tbx21fl/flNcr1-iCre mice, 8 days after infection. The scale bar represents 1000 µm. Histological scores were determined on 1-2 H&E sections per mouse colon. (c) Colon length and (d) C. rodentium bacterial loads (CFU) in the spleen (left histogram) and liver (right histogram) 8 days after infection. (b-d) Data are pooled from 2 independent experiments and are shown as means ± s.d. (n=5-9 mice per group in total). Statistical analyses were carried out with unpaired Student’s t-test. n.s., not statistically significant (P > 0.05).
Supplementary Figure 4 Gastrointestinal lymphocytes and C. rodentium infection in R26DTA/+Ncr1-iCre mice.
(a) Numbers of Lin−CD45+ small intestine lymphocytes from naïve R26DTA/+Ncr1-iCre and their respective Ncr1-iCre controls. Counts were obtained from flow cytometry plots showing intracellular RORγt and surface NKp46 expression. Numbers show cell counts within the indicated gate. Data were pooled from at least three independent experiments (n=2-4 mice per group in each) and are shown as the mean ± s.d. Statistical analyses were performed with unpaired Student’s t-test. **P < 0.01. (b) Characterization of colon ILCs in R26DTA/+Ncr1-iCre mice at steady state. Flow cytometry analyses (left panels) showing expression of RORγt and NKp46 within Lin−CD45.2+ cells from the colons of the indicated mice. Representative profiles show the frequency of the indicated populations (right panel). Data are pooled from two independent experiments (n=6 mice per group in total) and show the mean. Statistical analyses were carried out with unpaired Student’s t-test, ****P < 0.0001. (c-f) R26DTA/+Ncr1-iCre and Ncr1-iCre control mice were infected with 1010 CFU of C. rodentium. (c) The percentages of several myeloid and lymphoid populations were assessed in the colonic lamina propria, draining mesenteric lymph nodes and spleen from R26DTA/+Ncr1-iCre and Ncr1-iCre control mice, 12 days after infection. (d,e) Total cells were restimulated with PMA/ionomycin for 4 hours in the presence of brefeldin, to assess the percentage of IFN-γ- and IL-17A-producing CD4+ and CD8+ T cells, as indicated. Data shown are representative of two independent experiments. (f) Explants of distal colon were weighed, homogenized and incubated for 24 hours in complete RPMI. Cytokine levels were determined in the supernatants. Data shown are one of two independent experiments (n=6-7 mice per group in each) and show the mean.
Supplementary Figure 5 C. rodentium infection in Mcl1fl/flNcr1-iCre mice.
Mcl-1fl/flNcr1-iCre mice and controls were orally inoculated with 2 × 109 CFU of C. rodentium on day 0.
(a) Pathological features of the colons of the indicated mice, on day 8 after infection. Representative H&E staining of formalin-fixed paraffin-embedded sections (micrographs) of colons are shown. The scale bar represents 1000 µm and the insets are x 3.4 magnifications. (b) Colon length of Mcl-1fl/flNcr1-iCre and Ncr1-iCre control mice 8 days after infection. Data were pooled from three independent experiments (n=8-9 mice per group in total) and are shown as mean ± s.d. Statistical significance was determined in unpaired Student’s t-test. n.s., not statistically significant (P > 0.05). (c) C. rodentium bacterial loads (CFU) in the spleen (left histogram) and liver (right histogram) 8 days after infection. Data shown are representative of three independent experiments (n=3 mice per group in each).
Supplementary Figure 6 The intestinal bacterial microbiota is unaffected by the absence of NCR+ ILC3 cells.
(a) Left panel: bar plots depicting the microbiota composition, at family level, in R26DTA/+Ncr1-iCre mice (n=13) and Ncr1-iCre control littermates (n=12). Right panel: principal coordinates analysis based on weighted unifrac distances describing the similarity between mice microbiota. (b) Relative amounts of SFB DNA in the feces of R26DTA/+Ncr1-iCre and Ncr1-iCre control mice at the ages of 10 and 15 weeks. (c) Quantification of IL-17A-producing CD4+ T cells from the small intestinal lamina propria from R26DTA/+Ncr1-iCre (n=24) and Ncr1-iCre control (n=18) mice, with or without stimulation with PMA/ionomycin. Data were pooled from three independent experiments.
Supplementary Figure 7 Regional distribution and phenotype of gut ILCs.
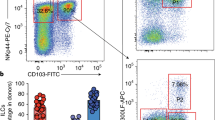
(a) NK1.1 expression on NCR+RORγt−, NCR+ ILC3 and NCR− ILC3 in different regions of the gastrointestinal tract. Representative flow cytometry plots show RORγt and surface NKp46 expression within Lin−CD45+ cells isolated from the lamina propria of the small intestine (S.I.), cecum and colon of naïve wild-type mice. The histogram shows the expression of NK1.1 and the frequency of NK1.1-positive cells within the indicated population. Data shown are representative of two experiments (n=6 per group in each). (b) Depletion of NK1.1-positive cells in the gastrointestinal tract. Wild-type mice were injected with 50 µg of anti-NK1.1 antibody or control PBS and they were then killed 5 days later. Bars represent the percentage of remaining NCR+ RORγt− cells, NCR+ ILC3 and NCR− ILC3 after treatment, in small intestine, cecum and colon, relative to that in control PBS-injected animals. Data shown are representative of two independent experiments (n=3 mice per group in each) and are showed as means ± s.d. Statistical significance was determined in unpaired Student’s t-test. *P < 0.05, ***P < 0.001 and ****P < 0.0001.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–3 (PDF 4046 kb)
Rights and permissions
About this article
Cite this article
Rankin, L., Girard-Madoux, M., Seillet, C. et al. Complementarity and redundancy of IL-22-producing innate lymphoid cells. Nat Immunol 17, 179–186 (2016). https://doi.org/10.1038/ni.3332
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3332
This article is cited by
-
PLZF restricts intestinal ILC3 function in gut defense
Cellular & Molecular Immunology (2023)
-
Gut microbiota and acute kidney injury: immunological crosstalk link
International Urology and Nephrology (2023)
-
Inulin diet uncovers complex diet-microbiota-immune cell interactions remodeling the gut epithelium
Microbiome (2023)
-
RIPK3 controls MAIT cell accumulation during development but not during infection
Cell Death & Disease (2023)
-
Interferon regulatory factor 1 (IRF-1) promotes intestinal group 3 innate lymphoid responses during Citrobacter rodentium infection
Nature Communications (2022)