Abstract
The transcription factor GATA-3 is indispensable for the development of all innate lymphoid cells (ILCs) that express the interleukin 7 receptor α-chain (IL-7Rα). However, the function of low GATA-3 expression in committed group 3 ILCs (ILC3 cells) has not been identified. We found that GATA-3 regulated the homeostasis of ILC3 cells by controlling IL-7Rα expression. In addition, GATA-3 served a critical function in the development of the NKp46+ ILC3 subset by regulating the balance between the transcription factors T-bet and RORγt. Among NKp46+ ILC3 cells, although GATA-3 positively regulated genes specific to the NKp46+ ILC3 subset, it negatively regulated genes specific to lymphoid tissue–inducer (LTi) or LTi-like ILC3 cells. Furthermore, GATA-3 was required for IL-22 production in both ILC3 subsets. Thus, despite its low expression, GATA-3 was critical for the homeostasis, development and function of ILC3 subsets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Change history
08 December 2015
In the version of this article initially published online, in the third and fourth paragraphs of the Discussion section, the mice are incorrectly identified as Rorcgfb/+. The correct genotype is Rorcgfp/+. The error has been corrected for the print, PDF and HTML versions of this article.
References
Spits, H. et al. Innate lymphoid cells–a proposal for uniform nomenclature. Nat. Rev. Immunol. 13, 145–149 (2013).
Hoyler, T. et al. The transcription factor GATA-3 controls cell fate and maintenance of type 2 innate lymphoid cells. Immunity 37, 634–648 (2012).
Yagi, R. et al. The transcription factor GATA-3 is critical for the development of all IL-7Ralpha-expressing innate lymphoid cells. Immunity 40, 378–388 (2014).
Klein Wolterink, R.G. et al. Essential, dose-dependent role for the transcription factor Gata3 in the development of IL-5+ and IL-13+ type 2 innate lymphoid cells. Proc. Natl. Acad. Sci. USA 110, 10240–10245 (2013).
Mjösberg, J. et al. The transcription factor GATA-3 is essential for the function of human type 2 innate lymphoid cells. Immunity 37, 649–659 (2012).
Moro, K. et al. Innate production of TH2 cytokines by adipose tissue-associated c-Kit+Sca-1+ lymphoid cells. Nature 463, 540–544 (2010).
Neill, D.R. et al. Nuocytes represent a new innate effector leukocyte that mediates type-2 immunity. Nature 464, 1367–1370 (2010).
Price, A.E. et al. Systemically dispersed innate IL-13-expressing cells in type 2 immunity. Proc. Natl. Acad. Sci. USA 107, 11489–11494 (2010).
Satoh-Takayama, N. et al. Microbial flora drives interleukin 22 production in intestinal NKp46+ cells that provide innate mucosal immune defense. Immunity 29, 958–970 (2008).
Buonocore, S. et al. Innate lymphoid cells drive interleukin-23-dependent innate intestinal pathology. Nature 464, 1371–1375 (2010).
Sonnenberg, G.F., Monticelli, L.A., Elloso, M.M., Fouser, L.A. & Artis, D. CD4+ lymphoid tissue-inducer cells promote innate immunity in the gut. Immunity 34, 122–134 (2011).
Klose, C.S. et al. Differentiation of type 1 ILCs from a common progenitor to all helper-like innate lymphoid cell lineages. Cell 157, 340–356 (2014).
Fuchs, A. et al. Intraepithelial type 1 innate lymphoid cells are a unique subset of IL-12- and IL-15-responsive IFN-γ-producing cells. Immunity 38, 769–781 (2013).
Halim, T.Y. et al. Group 2 innate lymphoid cells are critical for the initiation of adaptive T helper 2 cell-mediated allergic lung inflammation. Immunity 40, 425–435 (2014).
Basu, R. et al. Th22 cells are an important source of IL-22 for host protection against enteropathogenic bacteria. Immunity 37, 1061–1075 (2012).
Qiu, J. et al. Group 3 innate lymphoid cells inhibit T-cell-mediated intestinal inflammation through aryl hydrocarbon receptor signaling and regulation of microflora. Immunity 39, 386–399 (2013).
Hepworth, M.R. et al. Innate lymphoid cells regulate CD4+ T-cell responses to intestinal commensal bacteria. Nature 498, 113–117 (2013).
Oliphant, C.J. et al. MHCII-mediated dialog between group 2 innate lymphoid cells and CD4+ T cells potentiates type 2 immunity and promotes parasitic helminth expulsion. Immunity 41, 283–295 (2014).
Gasteiger, G. & Rudensky, A.Y. Interactions between innate and adaptive lymphocytes. Nat. Rev. Immunol. 14, 631–639 (2014).
Klose, C.S. et al. A T-bet gradient controls the fate and function of CCR6-RORγt+ innate lymphoid cells. Nature 494, 261–265 (2013).
Cella, M. et al. A human natural killer cell subset provides an innate source of IL-22 for mucosal immunity. Nature 457, 722–725 (2009).
Constantinides, M.G., McDonald, B.D., Verhoef, P.A. & Bendelac, A. A committed precursor to innate lymphoid cells. Nature 508, 397–401 (2014).
Sawa, S. et al. Lineage relationship analysis of RORγt+ innate lymphoid cells. Science 330, 665–669 (2010).
Wang, L. et al. Distinct functions for the transcription factors GATA-3 and ThPOK during intrathymic differentiation of CD4+ T cells. Nat. Immunol. 9, 1122–1130 (2008).
Serafini, N. et al. Gata3 drives development of RORγt+ group 3 innate lymphoid cells. J. Exp. Med. 211, 199–208 (2014).
Zhu, J. et al. Conditional deletion of Gata3 shows its essential function in TH1-TH2 responses. Nat. Immunol. 5, 1157–1165 (2004).
Eberl, G. & Littman, D.R. Thymic origin of intestinal αβ T cells revealed by fate mapping of RORγt+ cells. Science 305, 248–251 (2004).
Sciumé, G. et al. Distinct requirements for T-bet in gut innate lymphoid cells. J. Exp. Med. 209, 2331–2338 (2012).
Rankin, L.C. et al. The transcription factor T-bet is essential for the development of NKp46+ innate lymphocytes via the Notch pathway. Nat. Immunol. 14, 389–395 (2013).
Wei, G. et al. Genome-wide analyses of transcription factor GATA-3-mediated gene regulation in distinct T cell types. Immunity 35, 299–311 (2011).
Zhu, J. et al. The transcription factor T-bet is induced by multiple pathways and prevents an endogenous Th2 cell program during Th1 cell responses. Immunity 37, 660–673 (2012).
Robinette, M.L. et al. Transcriptional programs define molecular characteristics of innate lymphoid cell classes and subsets. Nat. Immunol. 16, 306–317 (2015).
Qiu, J. et al. The aryl hydrocarbon receptor regulates gut immunity through modulation of innate lymphoid cells. Immunity 36, 92–104 (2012).
Yu, Q., Erman, B., Park, J.H., Feigenbaum, L. & Singer, A. IL-7 receptor signals inhibit expression of transcription factors TCF-1, LEF-1, and RORγt: impact on thymocyte development. J. Exp. Med. 200, 797–803 (2004).
Acknowledgements
We thank D.R. Littman (New York University) for Rorc-Cre mice on the C57BL/6 background; A. Singer (National Cancer Institute) for Tg-hCD2-Il7ra mice on the C57BL/6 background; C. Dulac (Harvard) for Tbx21-Cre mice; D. Artis (University of Pennsylvania) for C. rodentium strain DBS100; A. Bhandoola, R. Germain, J. O'Shea, P. Schwartzberg and A. Sher for critical reading of our manuscript; L. Guo for suggestions and technical assistance in analyzing ILCs; K. Weng for assistance in cell sorting; L. Feigenbaum for assistance in generating RORγt–E2-Crimson mice; and the DNA Sequencing and Computational Biology Core of the National Heart, Lung, and Blood Institute for sequencing the RNA-Seq and ChIP-Seq libraries. Supported by the Intramural Research Program of the US National Institutes of Health, the National Institute of Allergy and Infectious Diseases and the National Heart, Lung, and Blood Institute.
Author information
Authors and Affiliations
Contributions
C.Z. performed all experiments; K.C. helped with constructing libraries for RNA-Seq and ChIP-Seq; C.W. helped with C. rodentium–infection experiments; G.H. performed initial analyses of the RNA-Seq and ChIP-Seq data; K.M. performed imaging experiments; K.C., C.W., G.H., K.M., Y.B. and K.Z. offered suggestions on the project and edited the paper; and C.Z. and J.Z. conceived of the project, designed the experiments, analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Deletion of GATA3 in ILC3 cells.
(a) Gating strategy for ILC3s from small intestine lamina propria (siLP). (b) Flow cytometry analysis of GATA3 in ILC2s and ILC3s of Gata3fl/fl and Gata3ΔILC3 mice. Data are representative of more than three independent experiments.
Supplementary Figure 2 Characterization of Gata3-deficient ILC3 cells.
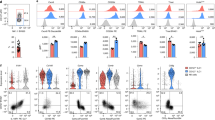
(a) Expression of the indicated cell-surface markers by CCR6+ and CCR6- siLP ILC3 subsets after Gata3 deletion. Data are representative of at least two independent experiments. (b) Heatmap of the expression of selected genes in ILC3s affected by Gata3 deletion, normalized by expression within each row. Live lineage-CD127+Thy-1highKLRG-1- ILC3s were sorted from siLP of Gata3fl/fl and Gata3ΔILC3 mice and subjected to RNA-Seq analysis. Signature genes in ILC3s affected after Gata3 deletion were selectively shown. Data are from two individual mice (n = 2 replicates) in each group.
Supplementary Figure 3 GATA3 affects the homeostasis of ILC3 cells in a cell-intrinsic manner.
(a) Schematic diagram of the mixed bone marrow (BM) chimera experiments. BM cells from Gata3fl/fl or Gata3ΔILC3 mice were mixed at 1:1 ratio with BM cells from wild type mice with a congenic CD45.1 marker, and transferred into irradiated Rag2-/-Il2rg-/- recipient mice. Repopulation of the transferred BM cell derived lymphocytes in recipient mice were analyzed 4-6 weeks later. (b) Flow cytometry analysis of repopulated lLC3s in the siLP and cLP, and B cells in the spleens of recipient mice. (c) Analysis of lLC3 and B cell repopulation from Gata3fl/fl and Gata3ΔILC3 BM, relative to the internal control cells derived from CD45.1+ BM. Data are representative of three independent experiments with 3-5 mice in each group. ***, p value < 0.001. NS, non-significant.
Supplementary Figure 4 Deletion of GATA3 affects the development of ‘ex-ILC3’ cells.
(a) Gating strategy and flow cytometry analysis of exILC3s from wild type and Gata3ΔILC3 mice. Gata3ΔILC3 mice were crossed with the RORγt fate mapping reporter mice (RORγtfm: Rosa26TdTomatoRorc-Cre) to generate Gata3ΔILC3RORγtfm. Gata3+/+RORγtfm mice were used as a wild type control. (b) Analysis of ILC1 and exILC3 cell number in Gata3+/+RORγtfm and Gata3ΔILC3RORγtfm mice. Three mice per each group were analyzed. **, p value < 0.01. NS, non-significant. Data are from one experiment.
Supplementary Figure 5 Deletion of Gata3 affects the NKp46+ ILC3 subset.
Gata3fl/fl mice and Gata3fl/flCre-ERT2 mice were treated with tamoxifen every other day for three times. Three month later, mice were sacrificed and NKp46+ ILC3s from siLP were analyzed. Data are representative of two independent experiments with more than three mice per group in each experiment.
Supplementary Figure 6 GATA3 suppresses RORγt expression in all ILC3 subsets independently of the regulation of IL-7R.
(a) Flow cytometry analysis of RORγt level in CCR6+ and CCR6- ILC3 subsets from Gata3fllfl and Gata3ΔILC3 mice. (b) Gating strategy of small intestine LTi cells from Gata3fllfl and Gata3ΔILC3 embryos at E15.5 days and analysis of the RORγt expression. (c) GATA3 negatively regulates RORγt expression by ILC3s in a cell intrinsic manner. Gata3fllfl and Gata3ΔILC3 BM were mixed with wild type congenic CD45.1 BM and transferred into irradiated Rag2-/-Il2rg-/- recipient mice. RORγt expression in repopulated ILC3s from different chimeras were analyzed by flow cytometry 4-6 weeks after transfer. (d) RORγt expression in siLP ILC3s from indicated mice were detected by flow cytometry. All data are representative of at least two independent experiments with more than three mice per group in each experiment.
Supplementary Figure 7 Expression of IL-7R and RORγt, as well as the development of NKp46+ ILC3 cells, are dysregulated in Rag1−/− Gata3ΔILC3 mice.
Flow cytometry analysis of CD127 and RORγt expression in siLP ILC3s (a) and of siLP ILC3 subsets (b) in the Rag1-/-Gata3fl/fl (n=4) and Rag1-/-Gata3ΔILC3 (n=6) mice. **, p value < 0.01; ***, p value < 0.001. Data are representative of two independent experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 771 kb)
Supplementary Table 1
Gene lists (XLSX 80 kb)
Rights and permissions
About this article
Cite this article
Zhong, C., Cui, K., Wilhelm, C. et al. Group 3 innate lymphoid cells continuously require the transcription factor GATA-3 after commitment. Nat Immunol 17, 169–178 (2016). https://doi.org/10.1038/ni.3318
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3318
This article is cited by
-
Proline uptake promotes activation of lymphoid tissue inducer cells to maintain gut homeostasis
Nature Metabolism (2023)
-
PD-1 signaling facilitates activation of lymphoid tissue inducer cells by restraining fatty acid oxidation
Nature Metabolism (2022)
-
Requirements for the differentiation of innate T-bethigh memory-phenotype CD4+ T lymphocytes under steady state
Nature Communications (2020)
-
Innate lymphoid cells control signaling circuits to regulate tissue-specific immunity
Cell Research (2020)
-
Orchestration between ILC2s and Th2 cells in shaping type 2 immune responses
Cellular & Molecular Immunology (2019)