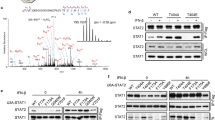
Abstract
Enhancing the response to interferon could offer an immunological advantage to the host. In support of this concept, we used a modified form of the transcription factor STAT1 to achieve hyper-responsiveness to interferon without toxicity and markedly improve antiviral function in transgenic mice and transduced human cells. We found that the improvement depended on expression of a PARP9-DTX3L complex with distinct domains for interaction with STAT1 and for activity as an E3 ubiquitin ligase that acted on host histone H2BJ to promote interferon-stimulated gene expression and on viral 3C proteases to degrade these proteases via the immunoproteasome. Thus, PARP9-DTX3L acted on host and pathogen to achieve a double layer of immunity within a safe reserve in the interferon signaling pathway.
Similar content being viewed by others
Main
The interferon signaling pathway is considered a mainstay of the immune system1,2, a concept that has been proven for the most part by loss-of-function approaches. Thus, deficiency in interferon production or signal transduction leads to defects in immune system–mediated control of microbial pathogens in experimental animal models and in humans3,4. The biological consequences of increased activity of the interferon signaling pathway are less certain, but chronic activation of the production of type I interferons has been linked to autoimmune disease in humans5. Moreover, the administration of exogenous interferons is accompanied by toxicity that limits their pharmacological use6. Such outcomes suggest that regulation of the interferon signaling pathway is already optimized under native conditions and that enhancing this pathway might not provide a therapeutic benefit without substantial toxicity.
However, previous approaches to enhancing the effects of interferon have generally relied on excessive expression or the administration of unregulated interferon ligand. An alternative strategy might be to enhance the efficiency of endogenous interferon to activate downstream signal transduction. While targeting the responsiveness of the interferon signal-transduction pathway might be desirable, it is made difficult by the complexity of the interferon signaling pathway and its functional activities. The protective actions of interferons rely on signaling through three types of interferon receptors (for type I, II and III interferons) and the JAK-STAT pathway that includes receptor-associated members of the JAK family of kinases and STAT family of transcription factors, as well as downstream modulators, transcription factors, enhancers and coactivators7. In the case of viral infection, the overall process leads to the expression of hundreds of distinct products of interferon-stimulated genes (ISGs) that provide functional effector activities but also feed back into interferon signaling2,8. Moreover, interferon signaling may also interact with other cytokine signaling pathways (for example, a pathway that includes the cytokine TNF) with additional immunological consequences9.
Despite such complexity, STAT1 stands out as a critical functional component common to each of the interferon-receptor signaling pathways10. Moreover, a mutant STAT1 with substitution of cysteine for alanine at positions 656 and 658 (STAT1-CC) enhances signal transduction via type I and II interferons in human cells, at least in vitro11. This enhancement is due to the enhanced capacity of STAT1-CC for transcriptosome assembly at the promoters of ISGs, which is manifested by prolonged activation (indicated by phosphorylation of Tyr701 of STAT1-CC), localization to the nucleus (indicated by binding of DNA by STAT1-CC) and recruitment of co-activators (indicated by binding of the histone acetyltransferase p300 (CBP) to STAT1-CC). Notably, STAT1-CC preserves the requirement for ligand-dependent activation. Expression of the gene encoding STAT1-CC thus offers a distinct gain-of-function approach for understanding how an improvement in interferon signaling might affect immunological function in vivo.
We initiated these experiments to define the consequences of a hyper-responsive interferon signaling pathway for host defense and, if successful, to establish new mechanisms for improving the outcome of infection. We therefore developed gain-of-function models in transgenic mice and transduced human cells for controlled expression of STAT1-CC. These models showed an increase in the efficiency of interferon signaling and a consequent benefit for host defense against a broad range of viruses. Whole-genome analysis of these models showed that the poly(ADP-ribose) polymerase PARP9 (also known as BAL1 or ARTD9)12 and the E3 ubiquitin ligase DTX3L were induced in concert with the interferon hyper-responsiveness. PARP9 was originally identified as being overexpressed in chemotherapy-resistant diffuse large B-cell lymphomas13 and was later reported to bind to DTX3L, which in turn was found to function as an E3 ubiquitin ligase that selectively ubiquitinates histone H4 and protects cells against DNA-damaging agents14,15,16. That function was consistent with the conventional role of members of the PARP family in the DNA-repair response17. However, PARP9 and DTX3L have also been found in a head-to-head orientation regulated by the same bidirectional interferon-responsive promoter, and overexpression of PARP9 in a malignant B cell lymphoma cell line has been shown to cause widespread induction of ISG expression, which would suggest an alternative role for PARP9 in the interferon signaling pathway18. Here we found that PARP9 and DTX3L were STAT1-associated components for interferon signal transduction that were rate limiting and were thus able to enhance interferon efficacy and consequent host defense when expressed in larger amounts. We also identified PARP9-DTX3L as an E3 ubiquitin ligase complex that targeted both host histone H2BJ to promote ISG expression and viral 3C protease to disrupt viral assembly. These observations offer proof of principle of how a single molecular complex can act on host and pathogen to provide a double benefit for interferon-dependent host defense and thereby afford some guidance for the development of therapeutic agents for interferon-sensitive conditions.
Results
STAT1-CC and interferon hyper-responsiveness in vivo
To achieve interferon hyper-responsiveness in vivo, we generated mice with a cytomegalovirus–β-actin–β-globin (CAG) gene promoter to drive widespread expression of a transgene encoding human STAT1-CC, to improve ISG expression (CAG-STAT1-CC mice)11. We also similarly generated transgenic mice expressing wild-type STAT1 instead as a control (CAG-STAT1 mice) (Supplementary Fig. 1a). CAG-STAT1 and CAG-STAT1-CC mice showed similar levels of transgene expression in heart, pancreas, brain and lungs (Supplementary Fig. 1b) and exhibited normal birth frequency, development and longevity without histological evidence of autoimmunity or other disease at 2 years of age (Supplementary Fig. 1c). STAT1-CC exhibited prolonged activation (as monitored by accumulation in the nucleus in heart tissue) compared with that of endogenous STAT1 or the transgene-derived wild-type STAT1 in response to administration of interferon-γ (IFN-γ) in vivo (Supplementary Fig. 1d,e). Similarly, CAG-STAT1-CC mice showed significantly higher ISG expression (monitored as expression of mRNA from the ISGs Irf1 and Gbp3 in the pancreas) at baseline and after IFN-β administration than did CAG-STAT1 or wild-type mice (Fig. 1a).
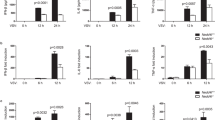
(a) Expression of Irf1 and Gbp3 mRNA in pancreas from wild-type (WT), CAG-STAT1 and CAG-STAT1-CC mice left untreated (− IFN-β) or treated (+ IFN-β) for 1 d with IFN-β (2 × 105 U). (b) Survival of wild-type, CAG-STAT1 and CAG-STAT1-CC mice (n = 15–27 per group) 0–12 d after infection with various amounts (above plots) of EMCV. (c) Expression of RNA encoding EMCV RNA polymerase 3D (EMCV 3Dpol) in the heart, brain and pancreas of wild-type, CAG-STAT1 and CAG-STAT1-CC mice (n = 5–8 per group) 1 or 3 d after infection with EMCV (100 PFU). (d) Immunostaining of EMCV in tissue sections of pancreas from mice 0 or 4 d after inoculation with EMCV (100 PFU). (e) Hematoxylin-and-eosin staining of tissue sections of heart, brain and pancreas from mice 0 or 4 d after infection with EMCV (100 PFU). Scale bars (d,e), 40 μm. *P < 0.01, versus wild-type mice (unpaired t-test (a,c) or Kaplan-Meier analysis (b)). Data are representative of three independent experiments (mean and s.e.m. in a,c).
We next investigated whether CAG-STAT1-CC mice were protected from viral infection. After being infected with encephalomyocarditis virus (EMCV) at an inoculum dose of 100 plaque-forming units (PFU), 100% of the wild-type and CAG-STAT1 mice died, whereas 97% of CAG-STAT1-CC mice survived at this inoculum, and 82% of them survived at an inoculum dose 100-fold higher (Fig. 1b). At lower viral inoculum dose (3 PFU), the survival of CAG-STAT1-CC mice was 100%, while only 25–28% of wild-type or CAG-STAT1 mice survived (Fig. 1b). Improved survival was associated with a markedly lower viral load at the main sites of infection (heart, brain and pancreas) in CAG-STAT1-CC mice than in wild-type or CAG-STAT1 mice (Fig. 1c). Similarly, by immunostaining of the pancreas, we found lower EMCV loads in CAG-STAT1-CC mice than in wild-type or CAG-STAT1 mice and concomitantly less tissue inflammation and injury in the heart, brain and pancreas of CAG-STAT1-CC mice (Fig. 1d,e). CAG-STAT1-CC mice were also protected against infection with influenza A virus (IAV; strain A/WS/33 or A/Vietnam/1203/04) and Venezuelan equine encephalitis virus, although to a lesser degree than the degree of their protection against infection with EMCV (Supplementary Fig. 2). Together these findings indicated that expression of STAT1-CC allowed better control of viral loads and virus-induced tissue damage across a broad range of viruses and tissue sites.
Interferon hyper-responsiveness and PARP9-DTX3L expression
To define the basis for the benefit afforded by STAT1-CC during viral infection, we analyzed whole-genome expression arrays of pancreas from CAG-STAT1-CC and CAG-STAT1 mice (Fig. 2a and Supplementary Table 1). We also analyzed gene expression in STAT1-deficient U3A human fibrosarcoma cells transduced with retrovirus for the expression of STAT1 (U3A-STAT1) or STAT1-CC (U3A-STAT1-CC) 11. U3A-STAT1-CC cells showed more translocation of STAT1-CC to the nucleus and higher ISG expression and better control of virus following infection with EMCV, IAV (strain A/WS/33) or Sindbis virus (SINV) than did U3A or U3A-STAT1 cells, with EMCV showing the greatest sensitivity to the actions of STAT1-CC (Supplementary Fig. 3). Similar to our in vivo approach, we used whole-genome expression arrays to identify genes that were expressed differentially in U3A-STAT1-CC cells than in U3A-STAT1 at baseline or after treatment with a low dose of IFN-β that mimics physiological conditions in vivo (Fig. 2b,c and Supplementary Tables 2 and 3). By comparing the sets of genes expressed differentially in three conditions—U3A-STAT1 cells versus U3A-STAT1-CC cells at baseline, U3A-STAT1 cells versus U3A-STAT1-CC cells with IFN-β, and CAG-STAT1 pancreatic tissues versus CAG-STAT1-CC pancreatic tissues—we identified an overlapping set of genes as candidates for encoding products that mediated the observed benefit of STAT1-CC in interferon signaling function (Fig. 2b,c and Supplementary Table 4). Within this set, we identified genes whose products were not previously assigned to any direct or indirect antiviral function as candidates for further study. Among those, the gene encoding PARP9 showed the greatest difference in expression in IFN-β-treated U3A-STAT1-CC cells relative to its expression in IFN-β-treated U3A-STAT1 cells (Fig. 2c and Supplementary Table 3). Consistent with the proposal of shared transcriptional regulation18, PARP9 mRNA and DTX3L mRNA and their corresponding proteins were regulated together at baseline and after interferon treatment in U3A-STAT1 and U3A-STAT1-CC cells and in 2fTGH cells, the parental U3A cell line that expresses native STAT1, as well as in pancreatic tissues from CAG-STAT1 and CAG-STAT1-CC mice, and demonstrated greater induction in STAT1-CC-expressing conditions than in STAT1-expressing conditions (Supplementary Table 1 and Supplementary Fig. 4). Together these results indicated that PARP9 and DTX3L were products of ISGs that were upregulated during the expression of STAT1 or STAT1-CC in vitro (in U3A-STAT1 and U3A-STAT1-CC cells) and in vivo (in CAG-STAT1 and CAG-STAT1-CC mice) and were therefore candidates for mediating STAT1- and STAT1-CC-dependent signal transduction.
(a) Expression (log2 normalized) of mRNA from pancreatic tissue of CAG-STAT1 and CAG-STAT1-CC mice at baseline, from a whole-genome array. Each symbol represents an individual gene: magenta, white and blue indicate genes expressed differentially in CAG-STAT1 mice versus CAG-STAT1-CC mice (antiviral function, key); gray indicates genes not expressed differentially; labels indicate genes (n = 25) with the greatest increase in expression. (b) Expression of mRNA from U3A-STAT1 and U3A-STAT1-CC cells at baseline (presented as in a); red indicates genes expressed differentially in both transgenic tissue and U3A cell lines (overlap, bottom right). Top left, retroviral vectors for U3A cell lines. (c) Expression (as in a,b) of mRNA from U3A-STAT1 and U3A-STAT1-CC cells treated for 1 d with IFN-β (10 U/ml), presented relative to expression at baseline. TG, transgenic tissue. (d) Expression of RNA encoding EMCV 3D (assessed 1 d after infection with EMCV (MOI, 1)), the IAV polymerase P3 (assessed 2 d after infection with IAV (MOI, 1)) or the SINV RNA-dependent RNA polymerase Nsp4 (assessed 1 d after infection with SINV (MOI, 10)) in U3A, U3A-STAT1, and U3A-STAT1-CC cells stably transduced with lentivirus encoding control shRNA or shRNA specific for PARP9 or DTX3L (two shRNAs for each (shRNA-1 and shRNA-2); horizontal axes). *P < 0.05, versus U3A cells, and **P < 0.05, versus U3A-STAT1-CC cells transduced with control shRNA (unpaired t-test). Data are from one experiment with pooling of three mice or samples (a–c) or are representative of three independent experiments (d; mean and s.e.m.).
PARP9-DTX3L and STAT1-dependent ISG expression
To analyze the function of PARP9 and DTX3L, we used lentiviral transduction for stable expression of short hairpin RNA (shRNA) targeting PARP9 mRNA or DTX3L mRNA in U3A, U3A-STAT1 and U3A-STAT1-CC cells. Specific knockdown of either PARP9 mRNA or DTX3L mRNA resulted in knockdown of both PARP9 and DTX3L protein without influencing the phosphorylation or activation of STAT1 or STAT1-CC (Supplementary Fig. 5a–e). Knockdown PARP9 mRNA decreased the expression of DTX3L protein without a decrease in DTX3L mRNA (Supplementary Fig. 5a–e), which suggested that PARP9 might regulate the expression of DTXL protein at a post-translational level. In contrast, knockdown of DTX3L mRNA caused a decrease in the expression of PARP9 protein that reflected the downregulation of PARP9 mRNA (Supplementary Fig. 5a–e), which suggested that DTX3L might regulate expression of the PARP9 gene at a transcriptional level. Knockdown of either PARP9 protein or DTX3L protein had no effect on the level of STAT1 or STAT1-CC (Supplementary Fig. 5c–e) but still resulted in loss of STAT1-CC-mediated control of infection with EMCV, IAV (strain A/WS/33) or SINV in U3A cells (Fig. 2d). Thus, PARP9 and DTX3L were necessary for the beneficial effect of STAT1-CC in controlling viral replication. In addition, restoring PARP9 expression to U3A cells in which PARP9 had been knocked down reversed the downregulation of DTX3L expression and reconstituted the antiviral effect of STAT1-CC on EMCV (Supplementary Fig. 5f,g), which again supported the proposal of a role for PARP9 in regulating the expression of DTX3L protein. We also found knockdown of PARP9 mRNA was overcome by treatment with interferon in U3A-STAT1 and U3A-STAT1-CC cells (Supplementary Fig. 5h), which indicated the difficulty in analyzing PARP9-DTX3L loss of function under interferon-stimulation conditions.
We next investigated whether PARP9-DTX3L was able to enhance the antiviral function of wild-type STAT1 in U3A cells overexpressing PARP9 (U3A-STAT1-PARP9 cells) or DTX3L (U3A-STAT1-DTX3L cells) or both PARP9 and DTX3L (U3A-STAT1-PARP9-DTX3L cells). We found that control of the EMCV, IAV (strain A/WS/33) or SINV viral load was improved in U3A-STAT1-PARP9-DTX3L cells but not U3A-STAT1-PARP9 or U3A-STAT1-DTX3L cells, and the beneficial effect was increased by combination with interferon treatment in U3A-STAT1-PARP9-DTX3L cells (Fig. 3). These results raised the possibility that PARP9 and DTX3L might work together with STAT1 to enhance interferon signaling. Co-immunoprecipitation assays showed that PARP9 and DTX3L interacted separately or together with STAT1 with or without interferon stimulation in U3A-STAT1 cells (which expressed FLAG-tagged STAT1, to facilitate these assays), and immunoblot analysis and immunostaining showed that PARP9 and DTX3L accumulated with STAT1 in the nucleus following stimulation of these cells with interferon (Fig. 4a–c). In addition, we found more binding of STAT1 to the promoter of the gene encoding the interferon-response factor (and transcription factor) IRF1 (as assessed by crosslinking chromatin immunoprecipitation (ChIP) assay), more STAT1-driven activity of promoters of an interferon-stimulated response element (ISRE) and an IFN-γ-activated site (GAS) (based on luciferase reporter–transactivation assays) and more localization of STAT1 to the nucleus (based on cell-imaging assays) in U3A-STAT1-PARP9-DTX3L cells than in U3A-STAT1 cells (Fig. 4d–f). However, the interactions of PARP9 and DTX3L with STAT1 in U3A-STAT1 cells were unchanged by stimulation with interferon (Fig. 4a) or substitution of STAT1 (Y701F) that eliminated the interferon-stimulated tyrosine-phosphorylation activation site (U3A-STAT1-Y701F cells) (Supplementary Fig. 6a). Moreover, we found no difference between U3A-STAT1-PARP9-DTX3L cells and U3A-STAT1 cells in the interferon-driven phosphorylation of STAT1 or in the recruitment of p300 to STAT1-containing transcriptional complexes (Supplementary Fig. 6b,c). Those results were in contrast to the STAT1-driven increases and STAT1-CC-driven further increases in the phosphorylation of STAT1 and recruitment of p300 to the STAT1 transcriptional complex (Supplementary Fig. 6c) to increase ISG expression and antiviral function in U3A-STAT1-CC cells11. Together these findings indicated that PARP9 and DTX3L associated with STAT1 at baseline and under interferon-stimulated conditions to modulate the localization of STAT1 to the nucleus and its transcriptional activity; however, the precise mechanism by which PARP9-DTX3L enhanced the STAT1-dependent expression of ISGs and antiviral function still needed to be defined.
Viral RNA (as in Fig. 2d) in U3A-STAT1, U3A-STAT1-PARP9, U3A-STAT1-DTX3L and U3A-STAT1-PARP9-DTX3L cells (key) with (− IFN-β) or without (+ IFN-β) pretreatment for 6 h with various doses IFN-β (horizontal axes), assessed 1 d after infection with EMCV, 2 d after infection with IAV (strain A/WS/33) or 1 d after infection with SINV. *P < 0.05, versus U3A-STAT1 cells (unpaired t-test). Data are representative of three independent experiments (mean and s.e.m.).
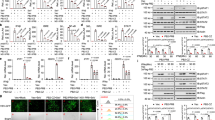
(a) Immunoblot analysis of the co-immunoprecipitation of FLAG-tagged STAT1 (left and middle) or STAT1-CC (right) with PARP9 or DTX3L (left margin) in parental U3A cells (left) or U3A cells expressing FLAG-tagged STAT1 (middle) or STAT1-CC (right), with (γ or β) or without (−) treatment for 24 h with IFN-γ (100 U/ml) or IFN-β (1,000 U/ml) (above lanes). (b) Immunoblot analysis of tyrosine-phosphorylated STAT1 (pY-STAT1), serine-phosphorylated STAT1 (pS-STAT1), or total STAT1, PARP9 or DTX3L (left margin) in cytosolic (C) and nuclear (N) fractions of U3A, U3A-STAT1 or U3A-STAT1-CC cells with or without treatment for 0.3 h with IFN-γ (as in a). Dashed outlines indicate tyrosine-phosphorylated STAT1, PARP9 and DTX3L in the nuclear fraction. (c) Immunostaining of U3A-STAT1-PARP9-DTX3L cells with (+ LMB) or without treatment for 1 h with leptomycin B (10 ng/ml), then with (+ IFN-β) or without (− IFN-β) treatment for 0.3 h with IFN-β (1,000 U/ml) plus leptomycin B. Scale bar, 10 μm. (d) Crosslinking ChIP assay of the binding of STAT1 to the IRF1 promoter in U3A-STAT1 and U3A-STAT1-PARP9-DTX3L cells left untreated (− IFN-β) or treated (+ IFN-β) for 0.3 h with IFN-β; after incubation of lysates with antibody to STAT1, the immunoprecipitated DNA was analyzed by real-time PCR with primers for IRF1 promoter, then results were normalized to those of input DNA. Results obtained with control antibody (mouse immunoglobulin G (IgG)) were similar to those of cells transfected with empty vector only (data not shown). (e) Transactivation of ISRE and GAS in U3A-STAT1 and U3A-STAT1-PARP9-DTX3L cells transfected for 12 h with a luciferase reporter vector for ISRE (left) or GAS (right) and left untreated (−) or treated (+) with IFN-β (left) or IFN-γ (right); results are presented as luminescence units (LU) relative to the activity of renilla luciferase. (f) Translocation of STAT1 to the nucleus in U3A-STAT1 and U3A-STAT1-PARP9-DTX3L cells before (−) and after (+) treatment for 0.3 h with IFN-β (1,000 U/ml) (left) or IFN-γ (100 U/ml) (right), presented as fluorescence units (FU) (nuclear fluorescence − cytoplasmic fluorescence); initial negative values (for −IFN-β) indicate that STAT1 was mainly cytoplasmic. *P < 0.01, versus U3A-STAT1 cells (unpaired t-test). Data are representative of three independent experiments (mean and s.e.m. (d,e) or mean and s.e.m. of three wells with 500 cells per well (f)).
PARP9-DTX3L and ISG expression with histone modification
We recognized from published work on the DNA-repair response that PARP9 has amino-terminal Macro1 and Macro2 domains that recognize poly(ADP)-ribose (PAR) for recruitment to sites of DNA damage and that DTX3L has a carboxy-terminal E3 ubiquitin ligase RING domain to ubiquitinate histone H4 at those sites14,15,16,19,20. We therefore investigated whether the ability of the PARP9-DTX3L complex to detect and modify histone H4 chromatin sites during the DNA-repair response might also apply to the response to interferon stimulation. However, we did not detect ubiquitination of histone H4 at baseline or under interferon-stimulated conditions in U3A-STAT1-PARP9-DTX3L cells (Supplementary Fig. 6d). To define an alternative mechanism by which PARP9-DTX3L enhanced ISG expression and antiviral control in these cells, we made structural predictions for targeted substitutions of conserved glycine residues in the PARP9 macro domains that are needed for the binding of PAR21,22 and in the cysteine residues in the DTX3L RING domain that are needed for catalytic binding of zinc23 (Fig. 5a and Supplementary Fig. 7a). The mutant PARP9 protein (PARP9M) was unable to bind to PAR, while the mutant DTX3L protein (DTX3LM) lacked E3 ligase activity (Supplementary Fig. 7b–d). We found that the interaction of PARP9M-DTX3LM with STAT1 (Supplementary Fig. 7e), the interaction between PARP9M and DTX3LM (Supplementary Fig. 7f) and the phosphorylation of STAT1 in response to interferon (Supplementary Fig. 7g) were all preserved in U3A-STAT1-PARP9M-DTX3LM cells relative to those interactions and phosphorylation in U3A-STAT1-PARP9-DTX3L cells. These results indicated that the PAR-binding activity of PARP9 and the E3 ubiquitin ligase activity of DTX3L were not required for binding to or activation of STAT1 or formation of the PARP9-DTX3L complex.
(a) Domains of PARP and DTX3L (top; amino acid positions above and right), and structure (below) of the two macro domains (Macro1 and Macro2) in PARP9, for binding of ADP-ribose (left), and the RING domain in DTX3L, for E3 ligase activity (right), based on homology models, identifying glycine residues (left) or cysteine zinc-chelating residues (right) targeted for loss-of-function substitution. Models based on the crystal structure of histone macroH2A1.1 (Protein Data Bank accession code, 1ZR3) with bound ADP-ribose (left) or the structure of the equine herpesvirus-1 RING domain (Protein Data Bank accession code, 1CHC) (right). (b) Gene expression (log2 normalized) in U3A-STAT1-PARP9-DTX3L cells versus U3A-STAT1-PARP9M-DTX3LM cells, from whole-genome array, presented relative to expression in U3A-STAT1 cells; red indicates genes expressed differentially here whose expression was increased by treatment with IFN-β (as reported in Fig. 2c). (c) ISG expression in U3A-STAT1, U3A-STAT1-PARP9-DTX3L, U3A-STAT1-PARP9M-DTX3LM, U3A-STAT1-PARP9-DTX3LM and U3A-STAT1-PARP9M-DTX3L cells (key) left untreated (− IFN-β) or treated (+ IFN-β) for 12 h with IFN-β (10 U/ml). (d) CHART PCR assays of cells as in c. (e) Viral RNA (as in Fig. 2d) in U3A-STAT1 cell lines given no pretreatment (− IFN-β) or pretreated with various concentrations (horizontal axes) of IFN-β (+ IFN-β) and then inoculated with EMCV (left), IAV (middle) or SINV (right). (f) Significance of histone ubiquitination (P value) versus ubiquitination of various histones (key) with DTX3L relative to their ubiquitination without DTX3L (horizontal axis), assessed by ubiquitin proteome array with DTX3L as the E3 ligase; dashed lines indicate a P value of 0.05 and a change in ubiquitination of '0-fold', so results in the top right quadrant are both statistically significant (P < 0.05) and biologically relevant (change in ubiquitination of >0-fold). (g) Native ChIP assay of the binding of ubiquitinated histone H2B (H2B-Ub), histone H3 trimethylated at Lys4 (H3K4me3) or H2B to the IFIT1 promoter in various U3A cell lines (as in c) left untreated (− IFN-β) or treated (+ IFN-β) for 16 h with IFN-β (1 U/ml), followed by incubation of nucleosomes from the cells with antibody to ubiquitinated H2B, trimethylated H3K4 or H2B, then real-time PCR analysis of the IFIT1 promoter sequence in the immunoprecipitated DNA; results were normalized to those of the corresponding input DNA and are presented relative to those of U3A-STAT1 cells, set as 1. Results obtained with control antibody (mouse IgG) were similar to those of cells transfected with empty vector only (data not shown). *P < 0.05, versus U3A-STAT1 cells (unpaired t-test). Data from one experiment with pooling of three cell samples (b,f) or are representative of three independent experiments (c–e,g; mean and s.e.m.).
To next identify ISGs that might be regulated in response to the activity of the PARP9 PAR-binding domain and DTX3L RING domain, we analyzed whole-genome expression arrays of U3A-STAT1-PARP9-DTX3L cells and U3A-STAT1-PARP9M-DTX3LM cells and compared each set to results obtained for U3A-STAT1 cells. We identified a set of ISGs (for example, IFIT1, IFIT3 and OASL) whose expression was enhanced in U3A-STAT1-PARP9-DTX3L cells relative to that in U3A-STAT1 cells, but not in U3A-STAT1-PARP9M-DTX3LM cells relative to that in U3A-STAT1 cells, and that overlapped with genes with increased expression (ISGs) from the array (Fig. 2c) of U3A-STAT1 cells treated with IFN-β versus those not treated with IFN-β (Fig. 5b and Supplementary Table 5). Further analysis with gene-specific PCR assays showed that baseline and IFN-β-stimulated expression of these ISGs was enhanced in U3A-STAT1-PARP9-DTX3L cells compared with that in U3A-STAT1 cells, and this increase was blocked in U3A-STAT1-PARP9M-DTX3LM and U3A-STAT1-PARP9-DTX3LM cells but was preserved U3A-STAT1-PARP9M-DTX3L cells (Fig. 5c). We next used a CHART PCR assay ('chromatin accessibility by real-time PCR') to determine whether the pattern for the expression of these ISGs corresponded to an increase in chromatin accessibility (Supplementary Fig. 7h). Indeed, we found an IFN-β-stimulated increase in chromatin accessibility in U3A-STAT1-PARP9-DTX3L cells compared with that in U3A-STAT1 cells, and this increase was blocked in U3A-STAT1-PARP9M-DTX3LM and U3A-STAT1-PARP9-DTX3LM cells but was preserved in U3A-STAT1-PARP9M-DTX3L cells (Fig. 5d and Supplementary Fig. 7i). Similarly, we found that antiviral function against EMCV, IAV (strain A/WS/33) and SINV was enhanced in U3A-STAT1-PARP9-DTX3L compared with that in U3A-STAT1 cells and that this enhancement was blocked in U3A-STAT1-PARP9M-DTX3LM and U3A-STAT1-PARP9-DTX3LM cells but was preserved U3A-STAT1-PARP9M-DTX3L cells (Fig. 5e). Together these results indicated that PARP9-DTX3L enhanced ISG expression and viral control by a mechanism that did not depend on the PAR-binding function of the PARP9 macro domains but might have required the ubiquitination function of the DTX3L-RING domain.
We next aimed to better define the ubiquitination function of DTX3L, particularly in relation to histone ubiquitination that might be linked to chromatin accessibility for ISG expression. Given the uncertainty over which histone(s) might be (a) DTX3L substrate(s), we used ubiquitin-proteome microarrays that included 29 different histones. We found that application of recombinant DTX3L to the microarray resulted in significant ubiquitination of histones H2BJ and H2AFV (isoform 3) (Fig. 5f and Supplementary Table 6), which indicated a high degree of selectivity for the E3 ubiquitin ligase activity of DTX3L. Native ChIP assays showed greater ubiquitination of H2B and concomitant methylation of histone H3 at Lys4 at the IFIT1 promoter in U3A-STAT1-PARP9-DTX3L than in U3A-STAT1 cells and that this increase was blocked in U3A-STAT1-PARP9M-DTX3LM and U3A-STAT1-PARP9-DTX3LM cells but was preserved U3A-STAT1-PARP9M-DTX3L cells (Fig. 5g). Moreover, co-immunoprecipitation assays demonstrated binding of histone H2BJ to the PARP9-DTX3L-STAT1 complex in HEK293T human embryonic kidney cells transfected to express H2BJ, PARP9 and DTX3L (Supplementary Fig. 7j). We also observed more mono-ubiquitination of endogenous H2B (but not of H2A) in U3A-STAT1-PARP9-DTX3L cells than in U3A-STAT1 cells at 6 h after infection with EMCV, and this effect was blocked in U3A-STAT1-PARP9M-DTX3LM cells (Supplementary Fig. 7k). Together these findings demonstrated that PARP9-DTX3L selectively ubiquitinated a subset of histones (notably H2BJ) and thereby increased histone methylation as a mechanism for chromatin remodeling and gene expression in at least a subset of ISGs (notably IFIT1), to confer enhanced antiviral function.
PARP9-DTX3L domains for interaction and function
To further address the role of PARP9 and STAT1 in the actions of PARP9-DTX3L, we identified domain-based regions of PARP9 and DTX3L proteins (Fig. 6a) for expression in HEK293T cells and used co-immunoprecipitation assays to map the regions of PARP9 and DTX3L required for interactions with each other and with STAT1. At baseline (without interferon stimulation), the interaction of DTX3L with STAT1 was confined to the amino-terminal portion of DTX3L, particularly the D3 domain (Fig. 6b); the interaction of PARP9 with STAT1 was mediated through both amino- and carboxy-terminal portions of PARP9, particularly the Macro1-Macro2 and PARP domains (Fig. 6c). Similarly, binding of PARP9 to DTX3L was localized to the D3 domain of DTX3L and the PARP domain of PARP9 (Fig. 6d,e). In addition, the interaction of PARP9-DTX3L domains with STAT1 were mostly unaffected by deletion of the STAT1 transactivation domain (Fig. 6b,c), the site of interferon-inducible binding of p300 (ref. 24). Moreover, the amino-terminal domain of PARP9 interacted with STAT1 even in the absence of binding of DTX3L (Fig. 6b,c).
(a) PARP9, DTX3L and STAT1 constructs (left half of each side) used for the identification of interacting domains, with tagging of epitopes with FLAG, c-Myc or hemagglutinin (HA) (key) and inclusion or deletion of various domains (construct designations, left margins); fold and secondary structure prediction were used for the identification of reasonable boundaries for domains D1, D2 and D3 of DTX3L. R, RING domain; CC, coiled-coil domain; SH2, Scr-homology 2 domain; TA, transactivation domain; del, deletion. Right half (of each side), domain interactions. (b–e) Immunoblot analysis (IB) of protein complexes immunoprecipitated (IP) together in lysates of HEK293T cells co-expressing various combinations of the constructs in a (above lanes), identifying the following binding (dashed outlines): STAT1-HA with c-Myc–DTX3L domains (b), STAT1-HA with FLAG-PARP9 domains (c), c-Myc–DTX3L with FLAG-PARP9 domains (d), and FLAG-PARP9 with c-Myc–DTX3L domains (e); arrowheads immunoprecipitated protein. Below, control immunoblot analysis with the same antibody used for immunoprecipitation and immunoblot. (f) Constructs for PARP9-M12 and DTX3L-delD3 (top), and immunoprecipitation and immunoblot analysis (below) of U3A-STAT1 cells transfected with vector alone (far left lanes) or vector expressing various constructs (above lanes), as well as a control immunoblot for protein input (far right); β-actin serves as a loading control throughout. (g) Construct for DTX3L-RDTX (top), and immunoblot analysis (below) of U3A-STAT1 cell lines transduced with vector alone (far left lane) or vector expressing various constructs (above lanes), assessed 6 h after infection with EMCV. (h) Expression of mRNA from the ISGs IFIT1, IFIT3 and OASL in U3A-STAT1 cell lines transduced with various constructs (key), then left untreated or treated for 12 h with IFN-β (10 U/ml). Data are representative of three independent experiments (mean and s.e.m. in h).
Given such interactions (Supplementary Fig. 8), we generated a mutant PARP9 lacking the PARP domain (PARP9-M12), mutant DTX3L lacking the D3 domain (DTX3L-delD3), and mutant DTX3L with only the RING and DTXH domains (DTX3L-RDTXH) (Fig. 6f,g), for lentiviral transduction and stable expression in U3A-STAT1 cells. We found that DTX3L-delD3, which did not interact with PARP9 (Fig. 6f) but maintained ubiquitination activity (discussed below), did not control EMCV levels (Fig. 6g) or induce ISG expression (Fig. 6h) in U3A-STAT1-PARP9-DTX3L-delD3 cells, whereas wild-type DTX3L did control EMCV levels and induce ISG expression in U3A-STAT1-PARP9-DTX3L cells. Similarly, PARP9-M12, which did not interact with DTX3L (Fig. 6f), did not induce ISG expression (Fig. 6h) in U3A-STAT1-PARP9-M12-DTX3L cells, whereas wild-type PARP9 did induce ISG expression in U3A-STAT1-PARP9-DTX3L cells. Of note, both PARP9-M12, which did not interact with DTX3L, and DTX3L-delD3, which did not interact with PARP9, still interacted with STAT1 (Fig. 6g), which indicated a redundant role in recruiting STAT1 to the complex. Together these results defined a carefully ordered set of domain requirements for the interactions of PARP9 and DTX3L with each other as well as of PARP9-DTX3L with STAT1 in the ISG-dependent response for the control of viral infection.
PARP9-DTX3L and degradation of viral 3C proteases
Since the ubiquitin-proteasome system can degrade the EMCV 3C protease25, we investigated whether PARP9-DTX3L might be an E3 ubiquitin ligase that targets this protease. We found lower expression of EMCV 3C in U3A-STAT1-PARP9-DTX3L cells than in U3A-STAT1 cells at 6 h after infection with EMCV (Fig. 7a). We did not detect a decrease in EMCV 3C in infected U3A cells that had not undergone complementation for STAT1 expression (Fig. 7a), consistent with a role for the products of STAT1-dependent ISGs (for example, PSMB, PSME and PMSF) in proteasome activity26 (Fig. 2a and Supplementary Tables 1–4). Moreover, the decrease in EMCV 3C persisted in U3A-STAT1-PARP9M-DTX3L cells, was lost in U3A-STAT1-PARP9-DTX3LM cells and was preserved in U3A-STAT1-DTX3L cells but not in U3A-STAT1-PARP9 cells (Fig. 7a). The observations that DTX3L alone was able to decrease the amount of EMCV 3C but was unable to fully control viral loads (Fig. 3a) suggested that PARP9 must have also contributed to the antiviral effect under conditions of interferon stimulation or viral infection, as well as at baseline. Furthermore, an inhibitor of the immunoproteasome component, PSMB8, effectively attenuated the degradation of EMCV 3C at 6 h after infection in U3A-STAT1-PARP9-DTX3L cells but not in U3A-STAT1-PARP9M-DTX3LM cells (Fig. 7b). The effect of DTX3L was specific to the degradation of EMCV 3C rather than an overall decrease in viral load, because the effect was independent of the amount of viral RNA at 6 h after infection (Fig. 7c). We also found a smaller amount of EMCV 3C in U3A-STAT1-PARP9-DTX3L cells than in U3A-STAT1 cells at 18 h after infection with or without treatment with IFN-β (Fig. 7d). Expression of PARP9-DTX3L or DTX3L via transient transfection in HEK293T cells also caused a decrease in EMCV 3C expression, and this effect was sensitive to blockade with the proteasome inhibitor MG-132 and was similar to the decrease in the amount of EMCV 3C found with expression of the E3 ubiquitin ligase TRIM22 (Fig. 7e), which has been linked to the ubiquitination of EMCV 3C27. Treatment with MG-132 also increased the amount of EMCV 3C without influencing the expression of EMCV RNA during infection of primary cultures of human airway epithelial cells (Fig. 7f,g). These cells also showed induction of the expression of endogenous PARP9-DTX3L after treatment with IFN-β or IFN-γ and interaction of endogenous PARP9-DTX3L with STAT1 (Fig. 7h).
(a) Immunoblot analysis of EMCV 3C in various U3A (left) or U3A-STAT1 (right) cell lines (above lanes) at 6 h after infection with EMCV (MOI, 1). (b) Immunoblot analysis of EMCV 3C and PSMB8 in various U3A cell lines (above lanes) left untreated (−) or treated (+) for 1 or 2 h (below blot) with the PSMB8-specific inhibitor ONX-0914 (0.1 μM) and then infected for 6 h with EMCV (MOI, 1). (c) EMCV RNA in cells as in a. (d) Immunoblot analysis of various proteins (left margin) in various U3A-STAT1 cell lines (above lanes) treated for 6 h with IFN-β (10 U/ml) or not (below blot) and then infected for 18 h with EMCV (MOI, 1). (e) Immunoblot analysis of EMCV 3C and other proteins (left margin) in HEK293T cells transfected for 36 h with various vectors (above lanes) alone (−) or together with (+) vectors encoding histidine- and V5-tagged EMCV 3C (His-V5-EMCV-3C) plus histidine-tagged ubiquitin (His-Ub) and then treated for 14 h with MG-132 (20 μM) (+MG-132) or not (−MG-132) (below blots). (f) Immunoblot analysis of EMCV 3C in human tracheal epithelial cells infected with EMCV at an MOI of 1 or 10 (top) and then, at 32 h later, treated for 12 h with MG-132 (20 μM) (+) or not (−). (g) EMCV 3Dpol RNA in cells as in f. (h) Immunoblot analysis of primary cultures of human tracheal epithelial cells with or without treatment for 24 h with interferon (above lanes), showing the co-immunoprecipitation of endogenous STAT1 with PARP9-DTX3L (top), and input controls (below). Data are representative of three independent experiments (mean and s.e.m. in c,g).
We also found that DTX3L localized with EMCV 3C and STAT1 at nuclear and cytosolic sites in EMCV-infected U3A cells, as assessed by immunostaining of U3A-STAT1-PARP9-DTX3L cells at 4–6 h after infection (Fig. 8a), consistent with localization of EMCV 3C reported before25,28,29. In addition, we found that DTX3L interacted with EMCV 3C and STAT1 as well as with PARP9, as assessed by co-immunoprecipitation from U3A-STAT1-PARP9-DTX3L cells with or without treatment with IFN-β (Fig. 8b), consistent with preserved formation of the PARP9-DTX3L-STAT1 complex despite any additional (potentially competing) interaction with EMCV 3C. Similarly, we detected binding of DTX3L to the IFIT1 promoter in U3A-STAT1-PARP9-DTX3L cells, and this binding was enhanced by infection with EMCV (Fig. 8c), consistent with EMCV 3C–assisted trafficking of PARP9-DTX3L to the nucleus and the loss of nuclear integrity during picornavirus infection. The binding of DTX3L to this promoter was inhibited in U3A-STAT1-PARP9M-DTX3LM cells (Fig. 8c), consistent with a decrease in chromatin accessibility (Fig. 5d).
(a) Immunostaining of EMCV 3C, DTX3L, and STAT1 in U3A-STAT1-PARP9M-DTX3LM cells at 5 h after infection with EMCV (MOI, 1). Scale bar, 10 μm. (b) Immunoblot analysis of the co-immunoprecipitation of EMCV 3C with PARP9-DTX3L in lysates of various U3A and U3A-STAT1 cell lines (above lanes) incubated for 5 min at 25 °C with recombinant histidine- and V5-tagged EMCV 3C (His-V5-EMCV-3C), followed by immunoprecipitation with antibody to (anti-) V5, probed with anti-c-Myc or anti-FLAG. (c) Crosslinking ChIP assay of various U3A-STAT1 cell lines without infection (−ECMV) or 6 h after infection with EMCV (MOI, 1) (+ ECMV) (key); after incubation of lysates with antibody to c-Myc–DTX3L, the immunoprecipitated DNA was analyzed by real-time PCR assay for IFIT1 promoter sequence, and results were normalized to input DNA and those of U3A-STAT1 cells. Results obtained with control antibody (mouse IgG) were similar to those of U3A-STAT1 cells (data not shown). (d) Immunoblot analysis of the ubiquitination of EMCV 3C in reaction mixtures containing various combinations (above lanes) of histidine- and V5-tagged EMCV (0.5 μg) as substrate, plus hemagglutinin-tagged ubiquitin (HA-Ub), E1 (ubiquitin-activating enzyme), the E2 ubiquitin-conjugating enzyme HbcH5a and DTX3L, detected with antibody to V5-tagged EMCV 3C and anti-hemagglutinin. The smear of bands with higher molecular weight indicates ubiquitinated proteins; native (non-ubiquitinated) histidine- and V5-tagged EMCV 3C monomer migrates as a 22-kDa protein. (e) Immunoblot analysis of the ubiquitination of EMCV 3C as in c, but with the addition of various amounts (above lanes) of histidine-tagged PARP9 and DTX3L to the reaction mixtures. (f) Immunoblot analysis of the ubiquitination of recombinant histidine- and V5-tagged tagged EMCV 3C after incubation with E1, E2 and DTX3L, with (+) or without (−) hemagglutinin-tagged ubiquitin, analyzed directly (−Ni-NTA) or after purification by nickel–nitrilotriacetic acid chromatography (+Ni-NTA), probed with anti-hemagglutinin and anti-V5. (g) Immunoblot analysis of the ubiquitination of HRV 3C in an assay as in f. (h) Immunoblot analysis of the K48-linked ubiquitination of EMCV 3C in an assay as in f, but with incubation with K48-linked ubiquitin (K48-Ub), and probed with antibody to K48-linked ubiquitin. (i) Immunoblot analysis of the ubiquitination of EMCV 3C and respiratory syncytial virus NS1 (RSV NS1) in the assay conditions in f. (j) Ubiquitination of EMCV 3C by DTX3L, DTX3L-delD3 and DTX3L-RDTXH in the assay conditions in f. Arrowheads (f–h,j) indicate ubiquitinated forms. (k) Binding of DTX3L-RING-DTXH to EMCV 3C (top) or HRV 3C (bottom), with DTX3L-RING-DTXH concentrations of 10, 5, 2.5, 1.25 and 0.625 μM (for EMCV 3C) or 8.5, 4.25, 2.13, 1.06 and 0.53 μM (for HRV 3C), assessed by real-time biolayer interferometry and presented as sensorgrams (left); solid lines indicate reference-corrected data. Right, orientation of biosensor pin. (l) Ubiquitination of EMCV 3C in various HEK293T cells transfected with various constructs (above lanes), assessed by immunoprecipitation of lysates with anti-V5 beads and then with anti-V5 in solution, followed by immunoblot analysis with antibody to hemagglutinin-tagged ubiquitin and anti-V5. *P < 0.05, versus uninfected cells (unpaired t-test). Data are representative of at least three independent experiments (mean and s.e.m. in c).
The findings reported above raised the possibility that DTX3L might contain a distinct domain for direct targeting of viral 3C proteases for ubiquitination. Indeed, by using recombinant proteins in an in vitro ubiquitination assay, we found that DTX3L ubiquitinated EMCV 3C (Fig. 8d). The ubiquitination of EMCV 3C by DTX3L was not affected by titration of various amounts of recombinant PARP9 into the in vitro ubiquitination assay (Fig. 8e), which ruled against the possibility of competition between the two proteins for binding to their substrate. We confirmed specificity for the ubiquitination of EMCV 3C by purification of the histidine- and V5-tagged ubiquitinated EMCV 3C reaction product with nickel–nitrilotriacetic acid columns (Fig. 8f). DTX3L also ubiquitinated the related human rhinovirus (HRV) 3C protease (Fig. 8g) that exhibits a pattern of early localization to the nucleus and late localization to the cytosol during infection similar to that of the EMCV 3C protease30. In vitro ubiquitination assays further showed that DTX3L did not require PARP9 for activity against EMCV 3C or HRV 3C (Fig. 8d–g) and that DTX3L was able to mediate Lys48 (K48)-linked ubiquitination of EMCV 3C (Fig. 8h). Recombinant DTX3L did not ubiquitinate the respiratory syncytial virus nonstructural protein NS1 (Fig. 8i), which is linked to ubiquitination-dependent degradation of host proteins31; this indicated that ubiquitination by DTX3L was at least somewhat specific to viral 3C proteases. DTX3L-delD3 and even DTX3L-RDTXH alone maintained ubiquitin ligase activity in vitro (Fig. 8j), which suggested that the RING and DTXH domains of DTX3L mediated ubiquitination and substrate recognition. Through the use of biolayer interferometry (a system for measuring protein interactions in real time), we were able to detect concentration-dependent interactions between recombinant DTX3L-RDTXH and both EMCV 3C and HRV 3C (Fig. 8k). Thus, the RING-DTXH domains of DTX3L were sufficient for the recognition of EMCV 3C and HRV 3C proteases and complemented the ability of the D1–D3 domains of DTX3L to bind PARP9-STAT1. In addition, using co-immunoprecipitation assays to detect V5-labeled EMCV 3C and hemagglutinin-labeled ubiquitin, we found that the ubiquitination of EMCV 3C was lower in HEK293T cells expressing PARP9M-DTX3LM than in those expressing PARP9-DTX3L (Fig. 8l). Together these results indicated that PARP9-DTX3L interacted with viral 3C proteases to target them for degradation and thus attacked the pathogen directly, in concert with interaction at ISG promoters to enhance the host response to viruses.
Discussion
Our study has shown that a single molecular complex, PARP9-DTX3L, mediated ubiquitination of both host histones and viral proteases to enhance interferon-dependent immunity and thereby control viral infection. We identified PARP9 and DTX3L on the basis of their increased expression in a gain-of-function system wherein modification of STAT1 to STAT1-CC (containing two cysteine substitutions in the Src homology 2 domain) enhanced the response to type I and II interferons and thereby broadly and safely provided protection against otherwise lethal viral infections in vitro and in vivo. Our analysis of this enhanced interferon responsiveness showed that PARP9 interacted with DTX3L and STAT1 and functioned as a chaperone to enhance levels of the PARP9-DTX3L protein complex and STAT1-mediated ISG expression. In addition, DTX3L interacted with a subset of host histones (notably histone H2BJ) and viral proteases (specifically viral 3C proteases) to function as an E3 ubiquitin ligase for both of these substrates. By targeting these substrates, DTX3L tailored chromatin accessibility for the expression of a specific subset of ISGs and promoted immunoproteasome-dependent blockade of viral replication. Each of these endpoints contributed separately to the control of viral infection. These findings place PARP9-DTX3L in a new context and establish interferon-driven ubiquitination as a key immunomodulatory step that might be targeted to enhance interferon signaling and control infection.
The multifunctional nature of PARP9-DTX3L was based on the use of distinct protein domains for interactions among PARP9, DTX3L, STAT1 and specific ubiquitination substrates (histones and viral 3C proteases). Thus, the dual function of DTX3L was mediated by binding of DTX3L to PARP9 via the D3 domain of DTX3L (to increase the level of DTX3L and its consequent function) and to ubiquitnation substrates via the RDTXH domain of DTX3L (to enable degradation of viral 3C and histone modification). In addition, the D1–D3 domains of DTX3L bound weakly to STAT1, and the macro and PARP domains of PARP9 bound strongly to STAT1, consistent with evidence that PARP9 also interacts with STAT1 via the macro domains of PARP9 in B cell lymphoma cell lines32. However, we found that the binding of PARP9-STAT1 was independent of conventional binding of the PARP macro domain to ADP-ribosylated histones described for the DNA-repair response16,33 and thus seemed to be specially 'tuned' to the interferon system. Since both STAT1 and the EMCV 3C protease traffic to nuclear and cytosolic sites25,28,29, binding of PARP9-DTX3L to STAT1 and EMCV 3C would allow at least two aids for the PARP9-DTX3L complex to also traffic to these two sites. This fits with detection of the PARP9–DTX3L–STAT1–EMCV 3C complex at both sites during infection and the ability of the PARP9-DTX3L complex to control viral and host gene expression in the nucleus and viral assembly in the cytoplasm. Since we also found that the PARP-DTX3L-STAT1 interaction occurred at baseline, the complex seemed to be ready in advance of infection. Moreover, PARP9 enhanced the level of DTX3L protein, and PARP9-DTX3L in turn promoted the expression of PARP9 and DTX3L, so that formation of the PARP9-DTX3L complex was also able to auto-amplify in response to the interferon production that developed after infection.
The properties of PARP9-DTX3L we have reported here appear to be distinct from those of other members of the PARP family in terms of the antiviral or interferon response. For example, PARP1 directly 'poly(ADP-ribosyl)ates' viral proteins to control viral replication34,35. In contrast, PARP9 completely lacks PARP enzymatic activity. PARP13 (ZAP) interacts directly with viral proteins but recruits the RNA exosome to mediate degradation without a ubiquitination mechanism36, and a shorter isoform of PARP13 (ZAPS) associates with the RNA helicase RIG-I to increase expression of interferons and other cytokines as an antiviral mechanism37. PARP12L, PARP7 and PARP10 might also have some antiviral activity, but the basis for this activity has been linked to the translation of viral and cellular RNA38. PARP14 regulates the activity of STAT6, but in contrast to results obtained for PARP9-DTX3L, the enzymatic activity of PARP was again required, no E3 ligase partner was identified, and DNA binding was undetectable39. In addition, we found no decrease in the level or function of STAT1 after overexpression of PARP9-DTX3L. Therefore, the action of PARP9-DTX3L was also distinct from that of E3 ubiquitin ligases that target STAT1 to downregulate interferon signaling40.
The identification of the actions of PARP9-DTX3L could provide valuable lessons for harnessing a safe reserve in the interferon signaling pathway for therapeutic benefit. In particular, we learned that PARP9-DTX3L contained modules for interaction with each other and with STAT1, specific cellular histones and viral 3C proteases to control viral infection at two levels (expression of host ISGs and degradation of viral 3C proteases) and two cellular sites (nucleus and cytoplasm). It therefore seems to be critical to understand and retain the multifunctional nature of PARP9-DTX3L to safely and effectively enhance interferon signal performance. We also learned that PARP9-DTX3L was especially suited to control viral 3C proteases, which, along with 3C-like proteases, are required for the wide range of common picornaviruses and other important pathogens in the picornavirus-like supercluster that includes certain coronaviruses and caliciviruses41. Thus, PARP9-DTX3L provides a particular 'road map' to guide the translation of these results toward a broad-acting next-generation therapeutic for this class of viruses. Indeed, small-molecule interferon signal–enhancer compounds have been identified in a cell-based screen for ISRE-driven luciferase reporter activity and have been shown to increase host ISG expression and control the level of EMCV and HRV in 2fTGH cells and primary cultures of human airway epithelial cells42. Whether these compounds also promote the degradation of viral 3C proteases and depend on PARP9-DTX3L expression for antiviral activity still needs to be determined. Nonetheless, small-molecule interferon signal–enhancer–type agents that mimic the antiviral mechanism of STAT1-CC and PARP9-DTX3L should help to fill the unmet need for protection against the acute illness and chronic inflammatory disease that develop with picornavirus infection43 and perhaps the need for the treatment of other interferon-sensitive conditions.
Methods
Generation of transgenic mice.
C57BL/6J mice were from Jackson Laboratory. CAG-STAT1 and CAG-STAT1-CC transgenic mice were generated using the pCAGGS vector that carries the cytomegalovirus early enhancer element; the promoter, first exon, and first intron of the chicken β-actin gene; and the splice acceptor of the rabbit β-globin gene44 to generate CAG-STAT1-CC-FLAG and CAG-STAT1-FLAG plasmids. A SalI- and PvuI-digested cDNA encoding CAG-STAT1-CC-FLAG or CAG-STAT1-FLAG was purified and microinjected into C57BL/6J zygotes. Transgenic founders were screened by PCR and then were bred to generate male mice for experiments. To assess interferon responsiveness, mice were treated with recombinant mouse IFN-γ or IFN-β (PBL Biomedical Laboratories) given by intraperitoneal injection of 2 × 104 or 2 × 105 U, respectively. All mouse strains were maintained on the C57BL6/J background. All mouse studies were performed under an IACUC-sanctioned protocol approved by the Animal Studies Committee of Washington University.
Immunoblot analysis.
Cells were lysed and tissues were homogenized in 1% Nonidet P-40, 0.05M Tris, pH 8.0, 250 mM NaCl, and 1 mM EDTA containing Halt Protease and Phosphatase Inhibitor (Thermo Fisher Scientific). Cell and tissue extracts were subjected to SDS-PAGE, and proteins were blotted onto PVDF membrane (Millipore) and incubated with anti-FLAG (M2; Sigma), rabbit antibody to human STAT1 phosphorylated at Tyr701 (9171; Cell Signaling Technology), rabbit antibody to human STAT1 phosphorylated at Ser727 (9177; Cell Signaling Technology), rabbit antibody to human PARP9 (40-4400; Thermo Fisher Scientific), rabbit antibody to human DTX3L (A300-834A; Bethyl Laboratories), mouse monoclonal antibody (mAb) to c-Myc (9E10; Santa Cruz), mouse antibody to human p300 (NM11; BD Biosciences), rabbit antibody to human STAT2 (sc-476; Santa Cruz), mouse antibody to human STAT1 (1; BD Biosciences), mAb to V5 (R960-25; Thermo Fisher Scientific), anti-hexahistidine (190-114a; Bethyl Laboratories), anti-RIG-I (ALX-804-960; Enzo), mouse mAb to EMCV 3D (from A. C. Palmenberg, University of Wisconsin), or polyclonal antibody to respiratory syncytial virus NS1 (J141; East Coast Biologics). Antibody to EMCV 3C was generated as described45. Primary antibodies were detected with horseradish peroxidase–conjugated goat antibody to rabbit IgG (7074; Cell Signaling) or horse antibody to mouse IgG (7076; Cell Signaling), by enhanced chemiluminescence.
For histone analysis, acid-extracted nuclear proteins were isolated46 and analyzed by immunoblot with rabbit antibody to histone H4 (2592; Cell Signaling Technology), mouse mAb to ubiquitinated histone H2B (56; Millipore), rabbit mAb to ubiquitinated histone H2A (D603A; Cell Signaling Technology), rabbit antibody to histone H2B (07-371; Millipore) and rabbit antibody to histone H3 (9715; Cell Signaling Technology), detected with enhanced chemiluminescence. In addition, gels were transferred to PVDF membranes (Li-COR, Biosciences) at 4 °C, and then the membranes were incubated for 1 h at 25 °C with Odyssey blocking buffer before incubation overnight at 4 °C with mouse mAb to ubiquitinated histone H2B, together with either rabbit mAb to ubiquitinated histone H2A or rabbit antibody to histone H3 (all identified above). The secondary antibody IRDye 680RD–conjugated goat anti-mouse (926-68070; Li-COR Biosciences) or IRDye 800CW–conjugated goat anti-rabbit (926-32211; Li-COR Biosciences) was applied for 1.5 h at 25 °C. Imaging and quantification was performed with a LI-COR Odyssey CLx scanner.
Immunohistochemistry.
Immunostaining for phosphorylated STAT1 and FLAG reporter for immunofluorescence microscopy was performed on frozen heart tissue that was embedded in OCT compound (Sakura Finetechnical) and cut into 6-μm sections, followed by blockade of nonspecific binding by incubation with 5% (wt/vol) normal donkey serum, and then incubation with rabbit antibody to phosphorylated STAT1 (9171; Cell Signaling Technology) and mouse mAb to FLAG (M2; Sigma) followed by Cy3-conjugated donkey antibody to rabbit IgG (711-165-152; Jackson ImmunoResearch) and fluorescein isothiocyanate (FITC)-conjugated donkey anti-mouse IgG (715-225-150; Jackson ImmunoResearch).
Immunostaining of the FLAG reporter for light microscopy was performed on paraffin-embedded heart tissue that was cut into 6-μm sections, followed by blockade of nonspecific binding by incubation with 5% normal goat serum, and incubation with rabbit antibody to FLAG (M2; Sigma). Primary antibodies were detected with biotinylated goat antibody to rabbit and the VECTASTAIN ABC-AP kit (PK-4001; Vector Laboratories). Sections were stained with an alkaline phosphatase red substrate and counterstained with hematoxylin.
Immunostaining for EMCV was performed on tissue that was frozen, cut into 6-μm-thick sections and fixed in cold acetone, followed by blockade of nonspecific binding by incubation with 10% nonimmune goat serum, then by incubation with mouse mAb to EMCV 3D (from A.C. Palmenberg), followed by incubation with peroxidase-conjugated goat antibody to mouse IgG (03117723001; Roche) and exposure to 3-amino-9-ethylcarbazole (06696-1G; Sigma).
For histopathological studies, mouse organs were fixed in 10% formalin, embedded in paraffin, and stained with hematoxylin and eosin. For staining of IAV (strain A/Vietnam/1203/04), nonspecific binding in paraffin-embedded lung and brain tissues was blocked by incubation with 5% (wt/vol) normal rabbit serum, followed by incubation with goat antibody to IAV (H1N1) (20-IG23; Fitzgerald Industries), followed by Cy3-conjugated donkey antibody to goat IgG (711-165-152; Jackson ImmunoResearch). Sections were mounted on slides with Vectashield medium containing DAPI (4,6-diamidino-2-phenylindole; Vector Laboratories).
RNA analysis.
RNA was isolated using the RNeasy Plus Mini Kit (QIAGEN) and was reverse-transcribed with a High Capacity cDNA Transcription Kit (Life Technologies). Real-time PCR was performed using fluorogenic probe-primer combinations (Supplementary Table 7) specific for the target gene and TaqMan FAST Universal master mix (Life Technologies). All mRNA levels were normalized to levels of the control gene mouse Gapdh mRNA, determined with the TaqMan Rodent GAPDH Control Kit or human GAPDH mRNA with primers for GAPDH nucleotides 37–910 or OAZ1 mRNA with the IDT PrimeTime Pre-designed assay Hs.PT.42.328511.g from Integrated DNA Technologies47.
Oligonucleotide microarray analysis was performed as described48. Gene-expression analysis was performed using Illumina Mouse-WG6 v1.1 BeadChips for mouse samples, and Illumina HumanHT-12 v3 BeadChips for human samples (Illumina). Isolated total RNA was amplified and biotinylated with an Ambion Illumina TotalPrep Kit. Hybridization and scanning of BeadChip arrays was performed according to the manufacturer's instructions, with BeadStudio 3.0 software (Illumina). Treatment and conditions were randomized across BeadChip slides to avoid confounding. For bead-level analysis, raw data were exported from BeadStudio.
Viral inoculation and monitoring.
Mouse EMCV (strain VR-129B) was titered by a viral plaque-forming assay49. IAV (strains A/WS/33 and A/Vietnam/1203/04) and Venezuelan equine encephalitis virus (strain ZPC738) were cultured as described50,51,52. SINV stocks were generated by electroporation of BHK-21 baby hamster kidney cells with RNA transcribed from plasmid TOTO 1101 (ref. 53). Mice were inoculated by intraperitoneal injection of EMCV (3, 1 × 102, or 1 × 104 PFU in 100 μl PBS), intranasal delivery of IAV strain A/WS/33 (25 PFU in 30 μl PBS) or strain A/Vietnam/1203/04 (1 × 105 TCID50 in 30 μl PBS), or subcutaneous injection of Venezuelan equine encephalitis virus (16 PFU in 100 μl PBS). Infected tissues were either fixed in 10% formalin for histology or homogenized for plaque-forming or TCID50 assay. Cell cultures were inoculated as described31. Cell supernatants were used for plaque-forming or TCID50 analysis and real-time quantitative PCR assays as described50,51,52,54. Cell lysates were used for isolation of RNA with an RNeasy kit (Qiagen) or for immunoblot analysis of EMCV proteins. For inhibition of PSMB8, cells were treated for 24 or 48 h with ONX-0914 (PR-957; Xcess Biosciences) at 0.1 μM, followed by infection for 6 h with EMCV (MOI 1), then were lysed for immunoblot analysis with antibody to EMCV 3C45. Real-time quantitative PCR for viral RNA was performed as described above with specific forward and reverse primers (Supplementary Table 7).
Cell line and primary-culture cell generation.
The U3A cell line was obtained from G. Stark, and STAT1 deficiency was confirmed as described11. U3A cells were transduced with retroviral vectors Mx-GFP, Mx-STAT1-IRES-GFP, Mx-STAT1-FLAG-IRES-GFP, Mx-STAT1-CC-IRES-GFP, or Mx-STAT1-CC-FLAG-IRES-GFP as described11. For expression of PARP9 and DTX3L, cDNA encoding FLAG-tagged human PARP9 or PARP9-M12 or c-Myc- tagged DTX3L or DTX3LM (Invitrogen) was inserted into lentiviral vector pLvx-IRES-puro (Clonetech) or pRRLsin.hCMV.IRES.EGFP to generate pLvx-FLAG-PARP9-IRES-puro, pLvx-c-Myc-DTX3L-IRES-Puro and pRRL-PARP9-IRES-GFP. Sequence encoding green fluorescent protein (GFP) was replaced with sequence encoding red fluorescent protein (RFP) (Evrogen) to generate pRRL-PARP9-IRES-RFP. For mutation of PARP9 (to generate mutant PARP9 (PARP9M)), human PARP9 cDNA encoding a Gly112-to-Glu112 substitution in Macro1 and a Gly311-to-Glu311 substitution in Macro2 were generated with the following primers in the QuikChange site-directed mutagenesis kit (Stratagene): 5′- GAAGATCTTCTGCATGGGGAGGGCCTGGCCCTGGCCCTG-3′ as forward and 5′- CAGGGCCAGGGCCAGGCCCTCCCCATGCAGAAGATCTTC-3′as reverse, and 5′- CCCACATGATGTTACAGTTGAGCCTGTGGCAAAGTCAATTC-3′ as forward and 5′- GAATTGACTTTGCCACAGGCTCAACTGTAATATCATGTGGG-3′ as reverse (where underlining and italics indicate the mutated nucleotides, here and below). For mutation of DTX3L (to create mutant DTX3L (DTX3LM), DTX3L cDNA encoding the Cys561-to-Ser561 and Cys564-to-Ser564 substitutions was generated with 5′-GAACTGGACAAGAAGGAAAAGGGCATCAGC GTCATCAGCATGGACACCATTAGTAACAAAAAAGTG-3′ as a forward primer and 5′- CACTTTTTTGTTACTAATGGTGTCCATGCTGATGACGCTGATGCCCTTTTCCT-TCTTGTCCAGTTC-3′ as a reverse primer. The mutants PARP9-M12, DTX3L with deletion of D3 (DTX3L-del-D3) and DTX3L-RING-DTXH (DTX3L-RDTXH) were also generated by PCR. These vectors were used for transduction of U3A-GFP, U3A-STAT1-IRES-GFP, and U3A-STAT1-CC-IRES-GFP cells. Primary cultures of human tracheobronchial epithelial cells were isolated from lung explants and cultured under 'submerged' conditions as described55.
STAT nuclear-translocation assay.
The high-content STAT1 nuclear-translocation assay was performed as described42. Cells (7 × 103 per well) were seeded into 96-well black tissue culture–treated plates and then were cultured overnight. After treatment with IFN-β (1,000 U/ml), cells were washed with Dulbecco's PBS, fixed with 4% formaldehyde and permeabilized, and then immunostained for STAT1 with mouse antibody to human STAT1 (1; BD Biosciences) and for STAT2 with rabbit antibody to human STAT2 (sc-476; Santa Cruz), followed by Alexa Fluor 660–conjugated goat antibody to mouse IgG (A-21054; Thermo Fisher Scientific) for STAT1, and Alexa Fluor 555–conjugated donkey antibody to rabbit IgG (A-31572; Thermo Fisher Scientific) for STAT2, and then nuclear staining with Hoechst 33342 (62249; Thermo Fisher Scientific). The plates were then imaged with the ArrayScan VTI High Content Imaging system (Cellomics-Thermo Fisher Scientific) with a 10× objective. The resultant images were processed with ArrayScan analysis software. Up to 15 images were obtained per well for analysis of up to 500 cells per well.
Gene knockdown.
Stable shRNA-expressing cell lines were generated with MISSION shRNA lentiviruses (Sigma). The sequences of shRNA for targeting human PARP9 were CCGGGCAGGTTCTAAAGGTGGAGAACTCGAGTTCTCCACCTTTAGAACCTGCTTTTTG (PARP9-1) and CCGGGCAAAGTCAATTCTACAACAACTCGAGTTGTTGTAGAATTGACTTTG-CTTTTTG (PARP9-2), and for DTX3L, they were CCGATGGACATTGATAGCGATCTCGAGATCGCTATCAA-TGTCCATCGGTTTTTG (DTX3L-1) and TTAGAGGTGGGTCCGAAATAACTCGAGTTATTTC-GGACCCACCTCTAATTTTTG (DTX3L-2). The Mission non-target shRNA vector was used as a control (Sigma). At 48 h after lentiviral inoculation, cells were selected with puromycin (1 μg/ml) to establish stable cell lines. Gene knockdown was assessed by real-time PCR assay and Immunoblot analysis.
Co-immunoprecipitation and crosslinking ChIP assays.
For STAT1-FLAG co-immunoprecipitation, cell lysates were incubated with agarose beads conjugated to anti-FLAG (M2; Sigma). Cross-linking ChIP (X-ChIP) promoter assays were performed as described11. Immunoprecipitated DNA was extracted with phenol-chloroform and ethanol precipitation, and an aliquot of 10% was analyzed by real time PCR with primers for the human IRF1 promoter (5′-CTTCGCCGCTAGCTCTAC-3′ and 5′-CCCATTGGCCGGCTGCGT-3′ as the forward and reverse primer pair, and 5′-CAGCCTGATTTCCCCGAAATGACG-3′ as the probe). For X-ChIP protein interaction assays, cell lysates were sheared by a 29-gauge syringe (instead of by sonication) and were incubated with agarose beads conjugated to antibody to p300 (sc-584; Sigma), then were washed and analyzed by immunoblot with anti-STAT1, anti-PARP9, anti-DTX3L or anti-p300 (all identified above). For co-immunoprecipitation of histone H2BJ with PARP9-DTX3L-STAT1, HEK293T cells were transfected for 48 h to express hemagglutinin (HA)-tagged H2BJ (Life Technologies), FLAG-tagged PARP9 and c-Myc-tagged DTX3L, and cell lysates were immunoprecipitated with anti-HA (HA-7; Sigma) or c-Myc (E6654; Sigma) and were analyzed by immunoblot with antibody to HA-H2BJ (H6908; Sigma), antibody to FLAG-PARP (M2; Sigma), antibody to c-Myc–DTX3L (9E10; Santa Cruz) and antibody to STAT1 (1; BD Biosciences). For X-ChIP assay of the binding of DTX3L to IFIT1 promoter, cells were infected with EMCV (MOI, 1) for 6 h and then were treated with formaldehyde. Cell lysates were incubated with anti-c-Myc (E6654, Sigma), and immunoprecipitated DNA was analyzed by real-time PCR assay with primers for the human IFIT1 promoter sequence (5′-CAGCAGGAATTCCGCTAGC-3′ and 5′-GCCAATGGTGTAAGCTGTGG-3′ as the forward and reverse primer pair, and 5′-GTGTGTCCTTGCAAGTTGG-3′ as the probe).
Immunocytochemistry.
For analysis of nuclear localization, U3A-STAT1-PARP9-DTX3L cells were treated with leptomycin B56 and IFN-β, then were fixed for 8 min at −20 °C with methanol and then incubated with mouse antibody to STAT1 (1:100 dilution; 1; BD Biosciences) and Alexa Fluor 488–labeled donkey anti-mouse (A-21202; Thermo Fisher Scientific), rabbit anti-PARP9 (40-4400; Thermo Fisher Scientific) and Alexa Fluor 647–labeled goat anti-rabbit (A-21246; Thermo Fisher Scientific), or rabbit anti-DTX3L (A300-834A; Bethyl) labeled with Alexa Fluor 555–labeled donkey anti-rabbit (A-31572; Thermo Fisher Scientific) and then with DAPI. For experiments on the co-localization of EMCV 3C and DTX3L, U3A-STAT1-PARP9M-DTX3LM cells were fixed and then were incubated with rabbit antibody to EMCV 3C (ref. 45) and Alexa Fluor 555–labeled donkey anti-rabbit (A-31572; Thermo Fisher Scientific) and with goat anti-DTX3L (sc-102496; Santa Cruz) and Alexa Fluor 633–labeled donkey anti-goat (A-21082; Thermo Fisher Scientific), and with mouse anti-STAT1 (1; BD Biosciences) and Alexa Fluor 488–labeled donkey anti-mouse (identified above), and then with DAPI. Cells were imaged using a Zeiss LSM 750 confocal microscope.
Gene-transactivation assays.
For luciferase reporter gene-transactivation assays, cells were co-transfected with the plasmid pGAS-Luciferase or pISRE-Luciferase (0.5 μg; Stratagene) and pRL-SV40-Renilla luciferase plasmid (0.03 μg, Promega) using X-tremeGene 9 (Roche Applied Science). After 24 h, transfected cells were treated for 12 h with IFN-γ (100 U/ml) or IFN-β (1,000 U/ml) and then split into aliquots for analysis of luciferase activity. The level of renilla luciferase activity was used to correct for transfection efficiency.
Protein domain mapping.
cDNA encoding STAT1, PARP9 or DTX3L was subcloned into pcDNA3.1 vectors with primers encoding epitope tag sequences to generate STAT1-HA, FLAG-PARP9 and c-Myc–DTX3L constructs. These plasmids (15 μg) were mixed with polyethylenimine (15 μg/ml) for transfection of HEK293T cells. After 48–72 h, transfected cells were lysed, and protein complexes were immunoprecipitated in 150 mM NaCl with anti-HA (HA-7; Sigma), anti-FLAG (M2; Sigma) or anti-c-Myc affinity gel (E6654; Sigma). Complexes were then eluted with FLAG, c-Myc or HA peptides and were subjected to immunoblot analysis with anti-FLAG (M2; Sigma), c-Myc (9E10; Santa Cruz) or anti-HA (HA-7; Sigma). The same co-immunoprecipitation conditions were used for U3A-STAT1-PARP9M12-DTX3L and U3A-STAT1-PARP9-DTX3l-delD3 cells.
Poly(ADP)-ribose (PAR)-binding assay.
In the first set of experiments, Nunc Maxisorp ELISA Plates (Thermo Fisher Scientific) were coated overnight at 4 °C with anti-PAR (4335-MC-01K-AC; Trevigen) in PBS, then washed, and nonspecific binding was blocked by incubation for 1 h with 2% BSA, followed by incubation for 2 h at 25 °C with PAR polymer (5 μl in 50 μl PBS) (4336-100-01; Trevigen) and washing again. Cell lysate (25 μg) containing FLAG-PARP9 or FLAG-PARP9M was then applied, followed by incubation for 3 h at 25 °C. After washes with PBS containing 0.05% Tween-20, mAb to FLAG (M2; Sigma) was applied, followed by incubation for 1 h at 25 °C and then additional washes with PBS containing 0.05% Tween-20. Horseradish peroxidase–conjugated anti-mouse (7076; Cell Signaling) was added, followed by incubation for 1 h at 25 °C, and binding was detected with a TMB 2 component Substrate Kit (KPL). The reaction was stopped by the addition of 2 N sulfuric acid, and absorbance at 450 nm was determined with a SpectraMax Plus microplate reader (Molecular Devices).
In the second set of experiments, binding of the PARP macro domain was assayed as described21,22. Cell lysates containing 1 mg protein in 1 ml of a solution of 50 mM Tris, pH 8.0, 150 mM NaCl, 1 mM EDTA, 0.5% NP40, and HALT inhibitor cocktail were added to 25 μl EZView Red FLAG M2 affinity gel (Sigma), followed by incubation overnight at 4 °C. Beads were then washed, followed by incubation overnight at 4 °C with 225 pmole (3 μl) of poly-ADP-ribosylated PARP1 (PARP-PAR; Trevigen) in 500 μl of the same buffer used for lysate binding. The complex was eluted with 200 μg/ml FLAG peptide in TBS with HALT inhibitor cocktail. Samples were then cleared by centrifugation and subjected to immunoblot analysis.
CHART PCR assay.
CHART PCR assays were performed as described57. Cells were treated for 0.5, 4 or 12 h with IFN-β (1000 U/ml). The cell pellet was resuspended in 1 ml ice-cold Nonidet P-40 lysis buffer (10 mM Tris (pH 7.4), 10 mM NaCl, 3 mM MgCl2, 0.5% Nonidet P-40, 0.15 mM spermine, and 0.5 mM spermidine), followed by incubation on ice for 5 min. The suspension was centrifuged at 3,000 r.p.m. for 5 min to pellet the nuclei. Nuclei were then suspended in the recommended New England BioLabs buffer (total volume of 50 μl) and 8 units of AvrII was added to initiate the digestion reaction at 37 °C for 15 min. The digested genomic DNA was purified with a QIAamp blood kit (Qiagen). CHART PCR assays for the human IFIT1 promoter were performed with the following forward and reverse primers: 5′-CAGCAGGAATTCCGCTAGC-3′ and 5′-GCCAATGGTGTAAGCTGTGG-3′ and the probe: 5′-TGTGTCCTTGCAAGTTGGA to cover ISRE sequence. Normalization of AvrII-cutting DNA samples was performed by real-time PCR against 2 kilobases of upstream genomic sequence of ISRE site lacking the AvrII restriction site. The following primers and probe were used for this PCR: 5′-CATAACTGGTCAGTCCCTTGC-3′ as a forward primer and 5′-TCAGCACTAGGTTGTGAGTTCG-3′ as a reverse primers, and 5′-AGCCCAGCTTAGACTCCT-3′ as the probe58. The relative accessible chromatin DNA was calculated as follows: value for AvrII-cutting sample stimulated with IFN-β (1000 U/ml) divided by value for AvrII-cutting samples not stimulated with IFN-β.
Ubiquitin proteome microarray.
Possible substrates of DTX3L were identified with a 17,000-protein microarray (E3 Substrate ID; MA102) from CDI Laboratories. Microarrays were incubated with recombinant E1 ubiquitin-activating enzyme, E2 (UBE2D1) ubiquitin-conjugating enzyme, ubiquitin, and ATP with or without recombinant DTX3L, and ubiquitinated substrates were detected with tandem ubiquitin-binding entities (TUBEs)59 with a system from LifeSensors. Comparison of conditions with and without DTX3L were analyzed by published methods55 and by methods described below for analysis of gene-expression microarrays. For data analysis, data files were generated by GenePix software containing signal intensities from the test sample (with DTX3L added) and background control sample (without DTX3L added). The median intensity values were processed with R software package limma for the following: background correction with the saddle model; normalization by positive-control spots (for example, IgG555/647); and additional normalization by array printer tip and location. By this normalization process, an M value (log2 test − log2 control value) and an A value [(log2 test + log2 control value)/2] were generated for each spot on the array. All proteins with an M value of <0, indicative of a lower signal in the test sample than in the control sample, were removed without further analysis. For the remaining proteins, the M values of duplicate spots for a protein on the array were subjected to a paired t-test to determine statistical significance.
Native ChIP assays.
To detect histone post-translational modifications, native ChIP was used as described with minor modifications60. Cells were treated for 16 h with IFN-β (1 U/ml). The cell pellet was suspended in 2 ml N-ChIP buffer I (0.3 M sucrose, 60 mM KCl, 15 mM NaCl, 5 mM MgCl2, 0.1 mM EGTA, 15 mM Tris-HCl, pH 7.5, 0.5 mM DTT, and protease inhibitors), then 2 ml of N-ChIP buffer with 0.4% NP-40, and then 8 ml of N-ChIP buffer with 1.2 M sucrose instead of 0.3 M sucrose. The mixture was centrifuged at 3,000 r.p.m. for 30 min, and the nuclei-containing pellet was resuspended in 1 ml of digestion buffer and then micrococcal nuclease (100 units in 1 μl; Thermo Fisher Scientific) was added, followed by incubation for 7 min at 37 °C. Nucleosomes were pelleted by centrifugation, resuspended in 1 ml of N-ChIP dialysis buffer (1 mM Tris-HCl, pH 7.5, 0.2 mM EDTA), and then dialyzed overnight at 4 °C. The dialyzed sample was centrifuged, and the supernatant was separated by electrophoresis through a 2.0% agarose gel and analyzed by immunoprecipitation by the same procedure as for X-ChIP but with mouse mAb to histone H2B-Ub (56; Millipore), antibody to histone H3 trimethylated at Lys4 (ab1012; Abcam) or rabbit antibody to histone H2B (07-371; Millipore).
Ubiquitination assays in HEK293T cells.
For confirmation of DTX3LM expression, HEK293T cells were transfected for 36 h with expression vectors for FLAG-PARP9-c-Myc–DTX3L or FLAG-PARP9M-c-Myc–DTX3LM, then were incubated for 14 h with 20 μM MG-132 (Enzo Life Sciences) and then subjected to lysis and Immunoblot analysis with anti-c-Myc (9E10; Santa Cruz). For analysis of the degradation of EMCV 3C, cDNA encoding EMCV 3C was amplified from EMCV virus stock, its sequence was confirmed by sequencing, then it was tagged with sequence encoding V5 and inserted into the expression vector pcDNA3.1. The plasmid pCI-His-Ub (31815; Addgene) was a gift from A. Winoto (University of California, Berkeley), and plasmid pHA-Ubiquitin (18712; Addgene) was a gift from E. Yeh (University of Texas, Houston). The plasmid encoding TRIM22 was from GeneCopoeia. Cells were transfected with vectors encoding V5-EMCV 3C and His-Ub using X-tremeGene 9 (Roche) and, 36 h later, were treated for 14 h with 20 μM MG-132 (or not) before lysis and Immunoblot analysis. For analysis of the ubiquitination of EMCV 3C, cell lysates were immunoprecipitated with mAb to V5 conjugated to agarose (V5-10; Sigma), followed by elution with 1% SDS, subsequent dilution to 0.1% SDS, reprecipitation with mAb to V5 (identified above), and immunoblot analysis with mAb to HA-tagged ubiquitin (HA-7; Sigma) followed by anti-V5 (identified above).
In vitro ubiquitination assay with recombinant proteins.
Sequence encoding V5-EMCV 3C, V5-HRV-A16 3C (from HRV-A16 obtained from J. Gern, University of Wisconsin), c-Myc–DTX3L-delD3 or c-Myc–DTX3L-RDTXH was cloned into plasmid pET28a as an amino-terminal hexahistidine-tagged construct, followed by transformation into Rosetta2 (DE3) cells (Novagen), and expression as soluble proteins that were purified by nickel–nitrilotriacetic acid chromatography followed by gel-filtration chromatography. The purified protein was concentrated to 10–20 mg/ml and was stored in a buffer consisting of 20 mM HEPES, pH 7.5, and 150 mM NaCl. Ubiquitination reactions were carried out in a 50-μl reaction mixture consisting of 50 mM Tris, pH 7.4, containing 5 mM MgCl2, 2 mM ATP, 2 mM DTT, 0.5 μg His-V5-EMCV 3C or His-V5-HRV 3C, 1 μg HA-Ub (Boston Biochem), 0.3 μg E1 ubiquitin-activating enzyme (Uba1; Boston Biochem), 0.6 μg E2 ubiquitin-conjugating enzyme (UbcH5a; Boston Biochem) and 0.5 μg DTX3L (Origene). The mixture was incubated for 30 min at 37 °C and was separated by SDS-PAGE14. Antibody to HA-tagged ubiquitin (3933; Cell Signaling Technology) and antibody to K48-linkage-specific polyubiquitin (4289; Cell Signaling Technology) were used for the detection of ubiquitinated protein. For experiments with combinations of PARP9 and DTX3L, various amounts of glutathione S-transferase (GST)-His-PARP9 (Abcam) and DTX3L were added to the reaction mixture. Respiratory syncytial virus NS1 protein (obtained from G. Amarasinghe, Washington University) was tested in the same way.
Biolayer interferometry assay.
Sequence encoding DTX3L-RING-DTXH (amino acids 555–740) was cloned into plasmid pET28a as an amino-terminal hexahistidine-tagged construct, and its sequence was confirmed by sequencing. The protein was purified by the procedure used for EMCV 3C and HRV-3C, described above. The binding of DTX3L-RING-DTXH to EMCV 3C and HRV 3C was assessed using biolayer interferometry with an Octet (ForteBio). Streptavidin-coated biosensors from ForteBio were used for capture of biotinylated proteins onto the surface of the sensor. After baseline was reached, sensors were moved to association step for 300 s and then dissociation for 300 s. Curves were corrected by subtraction of signal from a reference pin. The running buffer consisted of 10 mM HEPES, pH 7.5, 150 mM NaCl, 0.05% Tween, and 1% BSA.
Statistical analysis.
A two-tailed unpaired Student's t-test was used to assess statistical significance between means. Mouse survival was assessed by Kaplan-Meier analysis.
Accession codes.
GEO: whole-genome expression data, GSE61411, GSE61413, GSE61414 and GSE61421.
References
Stark, G.R. & Darnell, J.E. Jr. The JAK-STAT pathway at twenty. Immunity 36, 503–514 (2012).
Schneider, W.M., Chevillotte, M.D. & Rice, C.M. Interferon-stimulated genes: a complex web of host defenses. Annu. Rev. Immunol. 32, 513–545 (2014).
Sancho-Shimizu, V., Perez de Diego, R., Jouanguy, E., Zhang, S.-Y. & Casanova, J.-L. Inborn errors of anti-viral interferon immunity in humans. Curr. Opin. Virol. 1, 487–496 (2011).
Casanova, J.-L., Holland, S.M. & Notaragelo, L.D. Inborn errors of human JAKS and STATs. Immunity 36, 515–528 (2012).
Pascual, V. & Banchereau, J. Tracking interferon in autoimmunity. Immunity 36, 7–9 (2012).
George, P.M., Badiger, R., Alazawi, W., Foster, G.R. & Mitchell, J.A. Pharmacology and therapeutic potential of interferons. Pharmacol. Ther. 135, 44–53 (2012).
Schindler, C., Levy, D.E. & Decker, T. JAK-STAT signaling: from interferons to cytokines. J. Biol. Chem. 282, 20059–20063 (2007).
Schoggins, J.W. et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472, 481–485 (2011).
Shornick, L.P. et al. Airway epithelial versus immune cell Stat1 function for innate defense against respiratory viral infection. J. Immunol. 180, 3319–3328 (2008).
Villarino, A.V., Kanno, Y., Ferdinand, J.R. & O'Shea, J.J. Mechanisms of Jak/STAT signaling in immunity and disease. J. Immunol. 194, 21–27 (2015).
Zhang, Y. et al. Modification of the Stat1 SH2 domain broadly improves interferon efficacy in proportion to p300/CREB-binding protein coactivator recruitment. J. Biol. Chem. 280, 34306–34315 (2005).
Hottiger, M.O., Hassa, P.O., Luscher, B., Schuler, H. & Koch-Nolte, F. Toward a unified nomenclature for mammalian ADP-ribosyltransferases. Trends Biochem. Sci. 35, 208–219 (2010).
Aguiar, R. et al. BAL is a novel risk-related gene in diffuse large B-cell lymphomas which enhances cellular migration. Blood 96, 4328–4334 (2000).
Takeyama, K. et al. The BAL-binding protein BBAP and related deltex family members exhibit ubiquitin-protein isopeptide ligase activity. J. Biol. Chem. 278, 21930–21937 (2003).
Yan, Q. et al. BBAP monoubiquitylates histone H4 at lysine 91 and selectively modulates the DNA damage response. Mol. Cell 36, 110–120 (2009).
Yan, Q. et al. BAL1 and its partner E3 ligase, BBAP, link poly(ADP-ribose) activation, ubiquitylation, and double-strand DNA repair independent of ATM, MDC1, and RNF8. Mol. Cell. Biol. 33, 845–857 (2013).
Ciccia, A. & Elledge, S.J. The DNA damage response: making it safe to play with knives. Mol. Cell 40, 179–204 (2010).
Juszczynski, P. et al. BAL1 and BBAP are regulated by a γ interferon-responsive bidirectional promoter and are overexpressed in diffuse large B-cell lymphomas with a prominent inflammatory infiltrate. Mol. Cell. Biol. 26, 5348–5359 (2006).
Timinszky, G. et al. A macrodomain-containing histone rearranges chromatin upon sensing PARP1 activation. Nat. Struct. Mol. Biol. 16, 923–929 (2009).
Beneke, S. Regulation of chromatin structure by poly(ADP)-ribosylation. Front. Genet. 3, 169 (2012).
Neuvonen, M. & Ahola, T. Differential activities of cellular and viral macro domain proteins in binding of ADP-ribose metabolites. J. Mol. Biol. 385, 212–225 (2009).
Karras, G.I. et al. The macro domain is an ADP-ribose binding module. EMBO J. 24, 1911–1920 (2005).
Ozkan, E., Yu, H. & Deisenhofer, J. Mechanistic insight into the allosteric activation of a ubiquitin-conjugating enzyme by RING-type ubiquitin ligases. Proc. Natl. Acad. Sci. (USA) 102, 18890–18895 (2005).
Wojciak, J.M., Martinez-Yamout, M.A., Dyson, H.J. & Wright, P.E. Structural basis for recruitment of CBP/p300 coactivators by STAT1 and STAT2 transactivation domains. EMBO J. 28, 948–958 (2009).
Schlax, P.E., Zhang, J., Lewis, E., Planchart, A. & Lawson, T.G. Degradation of the encephalomyocarditis virus and hepatitis A virus 3C proteases by the ubiquitin/26S proteasome system in vivo. Virology 360, 350–363 (2007).
Tanahashi, N. et al. Hybrid proteasomes: induction by interferon-γ and contribution to ATP-dependent proteolysis. J. Biol. Chem. 275, 14336–14345 (2000).
Eldin, P. et al. TRIM22 E3 ubiquitin ligase activity is required to mediate antiviral activity against encephalomyocarditis virus. J. Gen. Virol. 90, 536–545 (2009).
Aminev, A.G., Amineva, S.P. & Palmenberg, A.C. Encephalomyocarditis virus (EMCV) proteins 2A and 3BCD localize to nuclei and inhibit cellular mRNA transcription but not rRNA transcription. Virus Res. 95, 59–73 (2003).
Lidsky, P.V. et al. Nucleocytoplasmic traffic disorder induced by cardioviruses. J. Virol. 80, 2705–2717 (2006).
Amineva, S.P., Aminev, A.G., Palmenberg, A.C. & Gern, J.E. Rhinovirus 3C protease precursors 3CD and 3CD' localize to nuclei of infected cells. J. Gen. Virol. 85, 2969–2979 (2004).
Lo, M.S., Brazas, R.M. & Holtzman, M.J. Respiratory syncytial virus nonstructural proteins NS1 and NS2 mediate inhibition of Stat2 expression and type I interferon responsiveness. J. Virol. 79, 9315–9319 (2005).
Camicia, R. et al. BAL1/ARTD9 represses the anti-proliferative and pro-apoptotic IFNg-STAT1–IRF1-p53 axis in diffuse large B-cell lymphoma. J. Cell Sci. 126, 1969–1980 (2013).
Aguiar, R.C., Takeyama, K., He, C., Kreinbrink, K. & Shipp, M.A. B-aggressive lymphoma family proteins have unique domains that modulate transcription and exhibit poly(ADP-ribose) polymerase activity. J. Biol. Chem. 280, 33756–33765 (2005).
Ohsaki, E. et al. Poly(ADP-ribose) polymerase 1 binds to Kaposi's sarcoma-associated herpesvirus (KSHV) terminal repeat sequence and modulates KSHV replication in latency. J. Virol. 78, 9936–9946 (2004).
Tempera, I. et al. Regulation of Epstein-Barr virus OriP replicaiton by poly(ADP-ribose) polymerase 1. J. Virol. 84, 4988–4997 (2010).
Chen, G., Guo, X., Lv, F., Xu, Y. & Gao, G. p72 DEAD box RNA helicase is required for optimal function of the zinc-finger antiviral protein. Proc. Natl. Acad. Sci. USA 105, 4352–4357 (2008).
Hayakawa, S. et al. ZAPS is a potent stimulator of signaling mediated by the RNA helicase RIG-I during antiviral responses. Nat. Immunol. 12, 37–44 (2011).
Atasheva, S., Frolova, E.I. & Frolov, I. Interferon-stimulated poly(ADP-ribose) polymerases are potent inhibitors of cellular translation and virus replication. J. Virol. 88, 2116–2130 (2014).
Goenka, S., Cho, S.H. & Boothby, M. Collaborator of Stat1 (CoaSt6)-associated poly(ADP-ribose) polymerase activity modulates Stat6-dependent gene transcription. J. Biol. Chem. 282, 18732–18739 (2007).
Yuan, C., Qi, J., Zhao, X. & Gao, C. Smurf1 protein negatively regulates interferon-γ signaling through promoting STAT1 protein ubiquitination and degradation. J. Biol. Chem. 287, 17006–17015 (2012).
Kim, Y. et al. Broad-spectrum antivirals against 3C or 3C-like proteases of picornaviruses, noroviruses, and coronaviruses. J. Virol. 86, 11754–11762 (2012).
Patel, D.A. et al. High-throughput screening normalized to biological response: application to antiviral drug discovery. J. Biomol. Screen. 19, 119–130 (2014).
Holtzman, M.J., Byers, D.E., Alexander-Brett, J. & Wang, X. The role of airway epithelial cells and innate immunce cells in chronic respiratory disease. Nat. Rev. Immunol. 14, 686–698 (2014).
Niwa, H., Yamamura, K. & Miyazaki, J. Efficient selection for high-expression transfectants with a novel eukaryotic vector. Gene 108, 193–199 (1991).
Lawson, T.G. et al. The encephalomyocarditis virus 3C protease is a substrate for the ubiquitin-mediated proteolytic system. J. Biol. Chem. 269, 28429–28435 (1994).
Fonseca, G.J. et al. Adenovirus evasion of interferon-mediated innate immunity by direct antagonism of a cellular histone posttranslational modification. Cell Host Microbe 11, 597–606 (2012).
Patel, D.A., Patel, A.C., Nolan, W.C., Zhang, Y. & Holtzman, M.J. High throughput screening for small molecule enhancers of the interferon signaling pathway to drive next-generation antiviral drug discovery. PLoS ONE 7, e36594 (2012).
Wu, K. et al. TREM-2 promotes macrophage survival and lung disease after respiratory viral infection. J. Exp. Med. 212, 681–697 (2015).
Binder, J.J., Hoffman, M.A. & Palmenberg, A.C. Genetic stability of attenuated mengovirus vectors with duplicate primary cleavage sequences. Virology 312, 481–494 (2003).
Choi, H.J., Song, J.H., Park, K.S. & Kwon, D.H. Inhibitory effects of quercetin 3-rhamnoside on influenza A virus replication. Eur. J. Pharm. Sci. 37, 329–333 (2009).
Song, L. et al. Superior efficacy of a recombinant flagellin:H5N1 HA globular head vaccine is determined by the placement of the globular head within flagellin. Vaccine 27, 5875–5884 (2009).
Yun, N.E. et al. CD4+ T cells provide protection against acute lethal encephalitis caused by Venezuelan equine encephalitis virus. Vaccine 27, 4064–4073 (2009).
Rice, C.M., Levis, J.H., Strauss, J.H. & Huang, H. Production of infectious RNA transcripts from Sindbis virus cDNA clones: mapping of lethal mutations, rescue of a temperature-sensitive marker, and in vitro mutagenesis to generate defined mutants. J. Virol. 61, 3809–3819 (1987).
Chen, Z. et al. Generation of live attenuated novel influenza virus A/California/7/09 (H1N1) vaccines with high yield in embryonated chicken eggs. J. Virol. 84, 44–51 (2010).
Byers, D.E. et al. Long-term IL-33-producing epithelial progenitor cells in chronic obstructive lung disease. J. Clin. Invest. 123, 3967–3982 (2013).
Begitt, A., Meyer, T., van Rossum, M. & Vinkemeier, U. Nucleocytoplasmic translocation of Stat1 is regulated by a leucine-rich export signal in the coiled-coil domain. Proc. Natl. Acad. Sci. (USA) 97, 10418–10423 (2000).
Rao, S., Procko, E. & Shannon, M.F. Chromatin remodeling, measured by a novel real-time polymerase chain reaction assay, across the proximal promoter region of the IL-2 gene. J. Immunol. 167, 4494–4503 (2001).
Wathelet, M.G., Clauss, I.M., Nols, C.B., Content, J. & Huez, G.A. New inducers revealed by the promoter sequence analysis of two interferon-activated human genes. Eur. J. Biochem. 169, 313–321 (1987).
Hjerpe, R. et al. Efficient protection and isolation of ubiquitylated proteins using tandem ubiquitin-binding entities. EMBO Rep. 10, 1250–1258 (2009).
Umlauf, D., Goto, Y. & Feil, R. Site-specific analysis of histone methylation and acetylation. Methods Mol. Biol. 287, 99–120 (2004).
Acknowledgements
We thank A.C. Palmenberg (University of Wisconsin) for mouse monoclonal antibody to EMCV 3D; G. Stark (Cleveland Clinic) for the U3A cell line; A. Winoto (University of California, Berkeley) for plasmid pCI-His-Ub; E. Yeh (University of Texas, Houston) for plasmid pHA-Ubiquitin; J. Gern (University of Wisconsin) for HRV A16; and G. Amarasinghe (Washington University) for respiratory syncytial virus NS1. Supported by the US National Institutes of Health (National Institute of Allergy and Infectious Diseases Asthma and Allergic Diseases Cooperative Research Center U19-AI070489 to M.J.H.; U54-AI05160 to M.J.H.; RO1-AI111605 to M.J.H.; and R15-AI099134 to T.G.L.) and the Martin Schaeffer Fund (M.J.H.).
Author information
Authors and Affiliations
Contributions
Y.Z. organized and performed cell and mouse experiments; D.M. performed cell and mouse experiments; W.T.R. performed immunoblot analyses; X.J. performed confocal microscopy; A.C.P. analyzed gene-expression microarray data; D.A.P. performed nuclear imaging and domain mapping; E.A. prepared viruses; Z.W. performed protein purification; R.M.T. performed knockout mouse breeding; J.J.A. performed mouse experiments; G.H. performed transcription-activity experiments; R.M. performed cell injections and implantations; J.Y. analyzed ubiquitination array data; N.E.Y. and S.P. performed Biosafety Level 3+ experiments; T.G.L. generated antibodies to 3C; N.S.O. performed biolayer interferometry assays; T.J.B. designed mutant PARP-DTX3L; M.J.H. designed experimental plans, analyzed data and wrote the manuscript; and all authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 CAG-STAT1 and CAG-STAT1-CC mice exhibit widespread transgene expression and activation without disease.
(a) Scheme for the expression cassette for CAG-STAT1 and CAG-STAT1-CC transgenic mice using CAG promoter and human STAT1 and STAT1-CC genes. (b) Immunoblots of tissue homogenates from wild-type (WT) mice and CAG-STAT1 and CAG-STAT1-CC transgenic mice using anti-FLAG or anti-β-actin Ab. (c) Hematoxylin-eosin staining of tissue sections from CAG-STAT1-CC transgenic mice at 2 years of age. Bar = 500 µm. (d) Immunostaining for STAT1 and STAT1-CC in myocardial tissue from WT mice and CAG-STAT1 and CAG-STAT1-CC transgenic mice at indicated times after treatment with IFN-γ (20,000 U intraperitoneally). Sections were stained using anti-FLAG Ab and an alkaline phosphatase system and then counterstained with hematoxylin. Control staining with non-immune IgG gave no signal above background (data not shown). Scale bar, 40 µm. Analysis of pancreas tissue gave similar results (data not shown). (e) For treatment conditions in (c), immunostaining of myocardial tissue with anti-phospho-STAT1 or anti-FLAG Ab and then secondary Ab conjugated to cyanine or FITC, respectively. Scale bar, 40 µm. Data are representative of two independent experiments (b,c,d,e).
Supplementary Figure 2 CAG-STAT1-CC mice show protection against infection with IAV and VEEV.
(a) Survival rates of WT and CAG-STAT1-CC transgenic mice inoculated with IAV-A/WS/33 at 25 PFU (n=15 mice per group). * P < 0.01. (b) Corresponding lung levels of IAV polymerase 3 (P3) RNA (n = 5-8 mice per group). (c) Corresponding hematoxyline-eosin staining of lung tissue. Scale bar, 40 µm. (d) Survival rates of WT and CAG-STAT1-CC transgenic mice inoculated with IAV-Vietnam/1203/04 at 1 x 105 TCID50 (n = 16 mice per group). * P < 0.01. (e) Corresponding IAV levels based on plaque-forming assay of samples from lung and brain at 6 days after infection. Values were normalized per gm of tissue. * P < 0.01. (f) Corresponding immunostaining for IAV with DAPI counterstaining of lung and brain tissue at 6 days after infection. Bar = 40 µm. (g) Survival rates of CAG-STAT1 and CAG-STAT1-CC transgenic mice infected with VEEV-ZPC738 (16 PFU given subcutaneously) (n = 11 mice per CAG-STAT1 and 16 mice per CAG-STAT1-CC group). * P < 0.01. (h) Corresponding viral levels based on plaque-forming assay of samples from lung 5 days after infection. * P < 0.01. Significance was determined with unpaired t-test (a,d,e,g,h).
Supplementary Figure 3 STAT1-CC expression enhances ISG expression, translocation of STAT and viral control in U3A cells.
(a) Immunostaining for STAT2 from high-throughput nuclear translocation assay in U3A-STAT1 and U3A-STAT1-CC cells at indicated times after treatment with IFN-β (1000 U/ml). Bar = 40 µm. (b) For the assay in (a), quantification of nuclear translocation of STAT1 and STAT2 after treatment with IFN-β (1000 U/ml) or IFN-γ (100 U/ml) in U3A-STAT1 and U3A-STAT1-CC cells for the indicated times. Values represent the difference in fluorescence intensity between nucleus and cytoplasm (mean ± SEM for 3 wells, 500 cells/well). Initial negative values indicate that STAT1 is predominantly cytoplasmic. * P < 0.01. (c) Target mRNA levels in indicated U3A cell lines with and without IFN-β (1000 U/ml) for 1 day. * P < 0.01. (d) Viral titers in indicated U3A cell lines at 1 day after infection with EMCV or 2 days after infection with IAV-A/WS/33 or SINV. * P < 0.01. (e) Viral titers in indicated U3A cell lines at 1 day after infection with EMCV (MOI 1), 2 days after infection with IAV-A/WS/33 (MOI 1) or SINV (MOI 10), or 1 day after infection with IAV-Vietnam/1203/04 (MOI 1) with or without pretreatment with the indicated concentrations of IFN-γ or IFN-β for 6 hours. * P < 0.01. Significance was determined with unpaired t-test (b,c,d,e). Data are representative of two independent experiments (a-e).
Supplementary Figure 4 STAT1-CC enhances PARP9-DTX3L expression in tissue and cells.
(a) Levels of PARP9 and DTX3L mRNA at baseline and after IFN-γ (1 U/ml) or IFN-β (10 U/ml) for 1 day in indicated U3A cell lines. (b) Levels of PARP9 and DTX3L mRNA at baseline and after IFN-γ (100 U/ml) or IFN-β (1000 U/ml) treatment for 1 day in indicated U3A cell lines. (c) For conditions in (a), corresponding levels of PARP9, DTX3L, STAT1, and β-actin protein in U3A cell lines and 2fTGH cells. (d) Parp9 and Dtx3l mRNA levels in pancreas tissue from WT mice and CAG-STAT1 and CAG-STAT1-CC transgenic mice treated with or without IFN-β (200,000 U i.p.) for 1 day. * P < 0.01. Analysis of lung tissue from these mice showed similar results (data not shown). Significance was determined using unpaired t-test (a,b,d). Data are representative of three independent experiments (a-e).
Supplementary Figure 5 Knockdown of PARP9 and DTX3L regulates expression and activation of PARP9-DTX3L but not of STAT1.
(a) Levels of PARP9 and DTX3L mRNA in indicated U3A cell lines stably transduced with lentivirus encoding PARP9 or control shRNA. (b) Corresponding analysis for lentivirus encoding DTX3L shRNA. For (a,b), * P < 0.01 for a significant decrease from corresponding control shRNA level. (c) Levels of PARP9 and DTX3L in indicated U3A cell lines stably transduced with lentivirus encoding PARP9 or control shRNA. (d) Levels of PARP9 and DTX3L for conditions in (c) but using DTX3L shRNA. (e) Immunoblot for the indicated proteins in U3A-STAT1 and U3A-STAT1-CC cells that were subjected to DTX3L gene knockdown and then treated with IFN-γ (100 U/ml) or IFN-β (1000 U/ml) for the indicated times with subsequent treatment with staurosporine to block JAK activity. (f) Immunoblot for the indicated proteins in indicated U3A cell lines that were subjected to PARP9 gene knockdown and reconstitution and infected with EMCV (MOI 1 for 6 hours). (g) Corresponding levels of EMCV-3D RNA for conditions in (f). (h) Levels of PARP9 mRNA in indicated U3A cell lines stably transduced with lentivirus encoding PARP9 or control shRNA as in (a) but including IFN treatment for 1 day. Significance was determined using unpaired t-test (a,b). Data are representative of three independent experiments (a-h).
Supplementary Figure 6 PARP9-DTX3L interacts with STAT1 without Tyr701 phosphorylation and does not enhance STAT1 phosphorylation, p300 recruitment, or histone H4 ubiquitination.
(a) Co-IP of STAT1 with PARP9 using U3A cells that express WT or mutant (Tyr701 to Phe701) STAT1 and PARP9 and were treated with IFN as indicated. (b) Immunoblot of STAT1 phosphorylation in U3A-STAT1 and U3A-STAT1-PARP9-DTX3L cells that were treated with IFN-β (1000 U/ml) or IFN-γ (100 U/ml) plus staurosporine at indicated times. (c) Co-IP of STAT1 with p300 in U3A-STAT1 and U3A-STAT1-PARP9-DTX3L versus STAT1-CC in U3A-STAT1-CC cells with or without IFN-β (1000 U/ml) or IFN-γ (100 U/ml) for 0.5 hours. (d) Immunoblot with anti-histone H4 Ab and anti-phospho-STAT1 Ab in indicated U3A cell lines with and without IFN-β (1000 U/ml) treatment. Predicted size of undetectable histone H4-Ub is also indicated. Data are representative of three independent experiments (a-d).
Supplementary Figure 7 Structure-based mutations inactivate PARP9-DTX3L function.
(a) Diagram of lentiviral vector constructs for WT and mutant PARP9 and DTX3L expression with sequence for double-glycine to glutamic acid mutations in the PARP9 macrodomains and double-cysteine to serine mutations in the DTX3L RING domain. (b) Levels of poly(ADP-ribose) (PAR) binding to PARP9 or PARP9 macrodomain mutant (PARP9M) in indicated U3A-STAT1 cell lines determined with a PARP9-PAR binding assay captured by anti-PAR Ab. (c) PAR binding assay for the indicated U3A-STAT1 cell lysates incubated with anti-FLAG M2 affinity gel for 6 h at 4 °C and then PAR-PARP1 (PAR-PARP) for 16 hours at 4 °C. The PAR-binding complex was eluted using FLAG peptide competition and immunoblotted with anti-PAR Ab. (d) Levels of ubiquitination of DTX3L or DTX3L ring domain mutant (DTX3LM) in HEK293T cells that were transiently transfected with the indicated vectors and then treated with MG-132 (20 µM) for 14 hours before immunoblotting with anti-c-Myc Ab. (e) Co-IP of DTX3L with STAT1 from indicated U3A-STAT1 cell lines using anti-c-Myc Ab and then immunoblot for the indicated proteins. (f) Co-IP of PARP9 with DTX3L from indicated cell lines using anti-FLAG or anti-cMyc Ab. (g) Immunoblot for STAT1 phosphorylation in U3A-STAT1 cell lines that were treated with or without IFN as indicated. (h) Diagram of the chromatin accessibility by real-time PCR (CHART) assay using Avr II endonuclease digestion of the human IFIT1 promoter region. Accessibility was assessed using IFIT1 promoter-specific PCR primers directed to nucleotides -135 to -50 relative to the transcription start site (TSS). (i) CHART assays for U3A-STAT1, U3A-STAT1-PARP9-DTX3L, and U3A-STAT1-PARP9M-DTX3LM cells under the indicated IFN-β treatment conditions. Values are expressed as the percentage of the accessibility observed in unstimulated digested DNA sample, i.e., % Accessibility = (1- (+)IFN/(-)IFN) x 100). (j) Co-IP of histone HA-H2BJ, FLAG-PARP9, c-Myc–DTX3L, and STAT1 in indicated HEK293T cell transfections. (k) Immunoblot levels of H2B-Ub and H2A-Ub detected and quantified with an infrared imaging system in indicated U3A-STAT1 cell lines (V, vectors alone; WT, PARP9-DTX3L; M, PARP9M-DTX3LM) with and without EMCV infection (MOI 1, 6 h). Significance was determined using unpaired t-test (b). Data are representative of three independent experiments (a-k).
Supplementary Figure 8 Formation of the PARP9-DTX3L-STAT1 complex depends on specific domain-mediated interactions.
Scheme for PARP9-DTX3L-STAT1 domain interactions based on co-ip assays of HEK293T cells transfected with individual domain components. Thin dashed lines indicate weak interactions, and thick lines indicate strong interactions in this assay system.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–7 (PDF 9222 kb)
Rights and permissions
About this article
Cite this article
Zhang, Y., Mao, D., Roswit, W. et al. PARP9-DTX3L ubiquitin ligase targets host histone H2BJ and viral 3C protease to enhance interferon signaling and control viral infection. Nat Immunol 16, 1215–1227 (2015). https://doi.org/10.1038/ni.3279
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3279
This article is cited by
-
SARS-CoV-2 and innate immunity: the good, the bad, and the “goldilocks”
Cellular & Molecular Immunology (2023)
-
Identification of Poly(ADP-ribose) Polymerase 9 (PARP9) as a Potent Suppressor for Mycobacterium tuberculosis Infection
Phenomics (2023)
-
PARP14 inhibition restores PD-1 immune checkpoint inhibitor response following IFNγ-driven acquired resistance in preclinical cancer models
Nature Communications (2023)
-
Exploring key molecular signatures of immune responses and pathways associated with tuberculosis in comorbid diabetes mellitus: a systems biology approach
Beni-Suef University Journal of Basic and Applied Sciences (2022)
-
BRCA1 mutations in high-grade serous ovarian cancer are associated with proteomic changes in DNA repair, splicing, transcription regulation and signaling
Scientific Reports (2022)