Abstract
Sumoylation regulates many cellular processes, but its role in signaling via the T cell antigen receptor (TCR) remains unknown. We found that the kinase PKC-θ was sumoylated upon costimulation with antigen or via the TCR plus the coreceptor CD28, with Lys325 and Lys506 being the main sumoylation sites. We identified the SUMO E3 ligase PIASxβ as a ligase for PKC-θ. Analysis of primary mouse and human T cells revealed that sumoylation of PKC-θ was essential for T cell activation. Desumoylation did not affect the catalytic activity of PKC-θ but inhibited the association of CD28 with PKC-θ and filamin A and impaired the assembly of a mature immunological synapse and central co-accumulation of PKC-θ and CD28. Our findings demonstrate that sumoylation controls TCR-proximal signaling and that sumoylation of PKC-θ is essential for the formation of a mature immunological synapse and T cell activation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Johnson, E.S. Protein modification by SUMO. Annu. Rev. Biochem. 73, 355–382 (2004).
Geiss-Friedlander, R. & Melchior, F. Concepts in sumoylation: a decade on. Nat. Rev. Mol. Cell Biol. 8, 947–956 (2007).
Flotho, A. & Melchior, F. Sumoylation: a regulatory protein modification in health and disease. Annu. Rev. Biochem. 82, 357–385 (2013).
Liu, B. & Shuai, K. Summon SUMO to wrestle with inflammation. Mol. Cell 35, 731–732 (2009).
Zhang, E.Y., Kong, K.F. & Altman, A. The yin and yang of protein kinase C-θ (PKCθ): a novel drug target for selective immunosuppression. Adv. Pharmacol. 66, 267–312 (2013).
Baier-Bitterlich, G. et al. Protein kinase C-θ isoenzyme selective stimulation of the transcription factor complex AP-1 in T lymphocytes. Mol. Cell. Biol. 16, 1842–1850 (1996).
Coudronniere, N., Villalba, M., Englund, N. & Altman, A. NF-κB activation induced by T cell receptor/CD28 costimulation is mediated by protein kinase C-θ. Proc. Natl. Acad. Sci. USA 97, 3394–3399 (2000).
Lin, X., O'Mahony, A., Mu, Y., Geleziunas, R. & Greene, W.C. Protein kinase C-θ participates in NF-κB activation induced by CD3–CD28 costimulation through selective activation of IκB kinase β. Mol. Cell. Biol. 20, 2933–2940 (2000).
Pfeifhofer, C. et al. Protein kinase C θ affects Ca2+ mobilization and NFAT cell activation in primary mouse T cells. J. Exp. Med. 197, 1525–1535 (2003).
Sun, Z. et al. PKC-θ is required for TCR-induced NF-κB activation in mature but not immature T lymphocytes. Nature 404, 402–407 (2000).
Zanin-Zhorov, A. et al. Protein kinase C-θ mediates negative feedback on regulatory T cell function. Science 328, 372–376 (2010).
Monks, C.R., Freiberg, B.A., Kupfer, H., Sciaky, N. & Kupfer, A. Three-dimensional segregation of supramolecular activation clusters in T cells. Nature 395, 82–86 (1998).
Fooksman, D.R. et al. Functional anatomy of T cell activation and synapse formation. Annu. Rev. Immunol. 28, 79–105 (2010).
Freiberg, B.A. et al. Staging and resetting T cell activation in SMACs. Nat. Immunol. 3, 911–917 (2002).
Yokosuka, T. et al. Spatiotemporal regulation of T cell costimulation by TCR-CD28 microclusters and protein kinase C θ translocation. Immunity 29, 589–601 (2008).
Hashimoto-Tane, A. et al. Dynein-driven transport of T cell receptor microclusters regulates immune synapse formation and T cell activation. Immunity 34, 919–931 (2011).
Bi, K. et al. Antigen-induced translocation of PKC-θ to membrane rafts is required for T cell activation. Nat. Immunol. 2, 556–563 (2001).
Hayashi, K. & Altman, A. Filamin A is required for T cell activation mediated by protein kinase C-θ. J. Immunol. 177, 1721–1728 (2006).
Huang, J. et al. CD28 plays a critical role in the segregation of PKC θ within the immunologic synapse. Proc. Natl. Acad. Sci. USA 99, 9369–9373 (2002).
Thuille, N. et al. Critical role of novel Thr-219 autophosphorylation for the cellular function of PKCθ in T lymphocytes. EMBO J. 24, 3869–3880 (2005).
Kong, K.F. et al. A motif in the V3 domain of the kinase PKC-θ determines its localization in the immunological synapse and functions in T cells via association with CD28. Nat. Immunol. 12, 1105–1112 (2011).
Yeh, E.T. SUMOylation and De-SUMOylation: wrestling with life’s processes. J. Biol. Chem. 284, 8223–8227 (2009).
Wertz, I.E. et al. De-ubiquitination and ubiquitin ligase domains of A20 downregulate NF-κB signalling. Nature 430, 694–699 (2004).
Knuesel, M., Cheung, H., Hamady, M., Barthel, K.K. & Liu, X. A method of mapping protein sumoylation sites by mass spectrometry using a modified small ubiquitin-like modifier 1 (SUMO-1) and a computational program. Mol. Cell. Proteomics 4, 1626–1636 (2005).
Seeler, J.S. & Dejean, A. Nuclear and unclear functions of SUMO. Nat. Rev. Mol. Cell Biol. 4, 690–699 (2003).
Marsland, B.J., Soos, T.J., Spath, G., Littman, D.R. & Kopf, M. Protein kinase C θ is critical for the development of in vivo T helper (Th)2 cell but not Th1 cell responses. J. Exp. Med. 200, 181–189 (2004).
Salek-Ardakani, S., So, T., Halteman, B.S., Altman, A. & Croft, M. Differential Regulation of Th2 and Th1 Lung Inflammatory Responses by Protein Kinase C. J. Immunol. 173, 6440–6447 (2004).
Roose, J.P., Mollenauer, M., Gupta, V.A., Stone, J. & Weiss, A. A diacylglycerol-protein kinase C-RasGRP1 pathway directs Ras activation upon antigen receptor stimulation of T cells. Mol. Cell. Biol. 25, 4426–4441 (2005).
Shuai, K. & Liu, B. Regulation of gene-activation pathways by PIAS proteins in the immune system. Nat. Rev. Immunol. 5, 593–605 (2005).
Grakoui, A. et al. The immunological synapse: a molecular machine controlling T cell activation. Science 285, 221–227 (1999).
Cartwright, N.G., Kashyap, A.K. & Schaefer, B.C. An active kinase domain is required for retention of PKCθ at the T cell immunological synapse. Mol. Biol. Cell 22, 3491–3497 (2011).
Kim, H. & McCulloch, C.A. Filamin A mediates interactions between cytoskeletal proteins that control cell adhesion. FEBS Lett. 585, 18–22 (2011).
Tavano, R. et al. CD28 interaction with filamin-A controls lipid raft accumulation at the T-cell immunological synapse. Nat. Cell Biol. 8, 1270–1276 (2006).
Bunnell, S.C., Kapoor, V., Trible, R.P., Zhang, W. & Samelson, L.E. Dynamic actin polymerization drives T cell receptor-induced spreading: a role for the signal transduction adaptor LAT. Immunity 14, 315–329 (2001).
Liu, Y.C., Penninger, J. & Karin, M. Immunity by ubiquitylation: a reversible process of modification. Nat. Rev. Immunol. 5, 941–952 (2005).
Zhang, W., Trible, R.P. & Samelson, L.E. LAT palmitoylation: its essential role in membrane microdomain targeting and tyrosine phosphorylation during T cell activation. Immunity 9, 239–246 (1998).
Mowen, K.A. & David, M. Unconventional post-translational modifications in immunological signaling. Nat. Immunol. 15, 512–520 (2014).
Zhou, R.W. et al. N-glycosylation bidirectionally extends the boundaries of thymocyte positive selection by decoupling Lck from Ca signaling. Nat. Immunol. 15, 1038–1045 (2014).
Smith-Garvin, J.E., Koretzky, G.A. & Jordan, M.S. T cell activation. Annu. Rev. Immunol. 27, 591–619 (2009).
Bunnell, S. in Immunological Synapse, Vol. 340 (eds. Saito, T. & Batista, F.D.) 123–154 (Springer Berlin Heidelberg, 2010).
Alcover, A. & Thoulouze, M.-I. in Immunological Synapse, Vol. 340 (eds. Saito, T. & Batista, F.D.) 191–207 (Springer Berlin Heidelberg, 2010).
Kumari, S., Curado, S., Mayya, V. & Dustin, M.L. T cell antigen receptor activation and actin cytoskeleton remodeling. Biochim. Biophys. Acta 1838, 546–556 (2014).
Liang, Y. et al. The lymphoid lineage-specific actin-uncapping protein Rltpr is essential for costimulation via CD28 and the development of regulatory T cells. Nat. Immunol. 14, 858–866 (2013).
Ulrich, H.D. The fast-growing business of SUMO chains. Mol. Cell 32, 301–305 (2008).
Vardhana, S., Choudhuri, K., Varma, R. & Dustin, M.L. Essential role of ubiquitin and TSG101 protein in formation and function of the central supramolecular activation cluster. Immunity 32, 531–540 (2010).
Ulrich, H.D. Mutual interactions between the SUMO and ubiquitin systems: a plea of no contest. Trends Cell Biol. 15, 525–532 (2005).
Castillo-Lluva, S. et al. SUMOylation of the GTPase Rac1 is required for optimal cell migration. Nat. Cell Biol. 12, 1078–1085 (2010).
Huang, J. et al. SUMO1 modification of PTEN regulates tumorigenesis by controlling its association with the plasma membrane. Nat. Commun. 3, 911 (2012).
Dustin, M.L. Hunter to gatherer and back: immunological synapses and kinapses as variations on the theme of amoeboid locomotion. Annu. Rev. Cell Dev. Biol. 24, 577–596 (2008).
Suzuki, A. et al. T cell-specific loss of Pten leads to defects in central and peripheral tolerance. Immunity 14, 523–534 (2001).
Wu, X. et al. Investigation of receptor interacting protein (RIP3)-dependent protein phosphorylation by quantitative phosphoproteomics. Mol. Cell. Proteomics 11, 1640–1651 (2012).
Acknowledgements
We thank W.-L. Hu for assistance with the preparation of primary human CD4+ T cells; A. Balancio for animal assistance; C.-y. Wu for sharing experience with experiments with peripheral blood mononuclear cells; and former and current laboratory members, K. Wang, S. Li, X. Feng, C. Zhao and D. Feng, for additional assistance. Supported by the Ministry of Science and Technology of China (2013CB835300 and 2009CB522202), the National Natural Science Foundation of China (31170846), the open research fund of State Key Laboratory of Cellular Stress Biology at Xiamen University, and the US National Institutes of Health (CA035299).
Author information
Authors and Affiliations
Contributions
X.-D.W., Y.G., Z.-L.C. and B.-N.G. did experiments and analyzed data; X.-D.W. and Y.G. prepared materials; J.-J.X. did experiments with Prkcq−/− T cells; C.-Q.Z. performed MS experiments; Q.-L.W. helped with experiments with human primary T cells; L.-H.D. helped with experiments with confocal imaging; A.X. contributed reagents and suggestions; J.H. and A.A. contributed reagents, helped with some experimental design, analyzed data and edited the manuscript; and Y.L. conceived of and designed the experiments, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of PKC-θ sumoylation.
(a-e) Immunoblot analysis of Flag-PKC-θ sumoylation in HEK293T cells transfected with the indicated combinations of expression vectors (a), in Jurkat E6.1 cells stimulated with SEE-pulsed Raji B cells (b), in Jurkat E6.1 cells transfected with empty vector (EV) YFP-EV or YFP-SENP1 and stimulated with SEE-pulsed Raji B cells (c), in Jurkat E6.1 cells stimulated with 10 μg/mL anti-CD3 (α-CD3) or 10 μg/mL anti-CD3 plus 2 μg/mL anti-CD28 (α-CD3 + α-CD28) (d), or in HEK293T cells were transfected with the indicated combinations of expression vectors (e). PKC-θ immunoprecipitates (IPs) or whole cell lysates (WCL) were immunobloted with the indicated Abs. Empty circles and arrowheads indicate sumoylated PKC-θ recognized by both anti-Myc and anti-PKC-θ Abs. Solid arrowheads indicate sumoylated PKC-θ recognized both by anti-SUMO1 and by anti-PKC-θ Abs. The asterisk denotes nonspecific bands. (f) Samples preparation for LC-MS/MS analysis. Flag-tagged PKC-θ together with Myc-tagged SUMO1-T95R were overexpressed in Jurkat TAg cells. 48 h post-transfection, cells were lysed in buffer containing 20 mM NEM. After clearing by centrifugation, 1% SDS (v/v) was added to the supernatants and proteins were dissociated by heating at 90°C for 10 min. Samples were diluted 1:10 with lysis buffer, and Flag-tagged PKC-θ were immunoprecipitated with anti-Flag M2 Ab at 4°C overnight. Immunoprecipitates were extensively washed 5x with lysis buffer and subjected to SDS-PAGE. Then the gel was cut into two parts, one for commassie blue staining and the other for immunoblotting with anti-SUMO1 and anti-PKC-θ Abs. After aligning the two parts, the gel band corresponding to nonsumoylated PKC-θ and the gel panel containing upper-shifted bands recognized by SUMO1 antibody were excised from the Commassie blue stained gel and analyzed by LC-MS/MS individually. (g) MS identification of sumoylated PKC-θ peptides. (h) Sequence alignment of ψKXD/E sumoylation motifs surrounding K325 or K506 in PKC-θ from the indicated species. The sequences correspond to amino acid residues 322-330 and 504-512, respectively, of human PKC-θ. The conserved lysine, glutamic acid, and aspartic acid residues in the ψKXD/E motif are underlined and bolded. (i) In vivo sumoylation assay of overexpressed Amphioxus (B. belcheri) PKC-δ (bbPKC-δ, NCBI accession number: JQ684102, an ancestor of both PKC-δ and PKC-θ) immunoprecipitated from HEK293T cells transfected with the indicated expression vectors. (j) Immunoblot analysis of endogenous PKC-θ protein expression in primary human CD4+ T cells transfected with siNC (a scrambled sequence as negative control) or siPKC-θ (a PKC-θ-specific siRNA targeting the 3’-UTR of PKC-θ mRNA). Anti-actin immunoblotting served as a loading control. Data are representative of at least three experiments in (a -e, i-j).
Supplementary Figure 2 Desumoylation of PKC-θ inhibits T cell activation.
(a) Flow cytometry analysis of GFP expression in unstimulated (US) or stimulated Prkcq−/− primary CD4+ T cells transduced with empty GFP vector, PKC-θ-WT, or PKC-θ-2KR (K325R-K506R). The mean fluorescence intensity (MFI) of GFP-expression is shown. Data in (a) are representatives of at least two independent experiments. (b) Flow cytometry analysis of activation markers expression on YFP+ gated naive (CCR7+CD45RO-) CD4+ cells from primary human CD4+ T cells transfected with siPKC-θ together with YFP–PKC-θ-WT or YFP–PKC-θ-2KR, cultured for 24 h, and stimulated with SEE-pulsed human APCs at a ratio of 1:1 for 24 h at 37°C. Left plots, one out of 4 representative experiments. Right panels, the frequency of gated CD25+ (left) and CD69+ (right) cells in 4 independent experiments. *P < 0.05, **P < 0.01 (two-way ANOVA with Bonferroni’s post-hoc test). (c) Immunoblot analyzing the proteins expression in WCL of Jurkat E6.1 cells transfected with siNC or siPKC-θ, and stimulated with anti-CD3 plus -CD28. (d) Jurkat E6.1 cells, in which endogenous PKC-θ was knocked down, were cotransfected with an empty vector or the indicated Flag-PKC-θ vectors plus NF-κB, AP-1, or NFAT reporter genes and an internal control Renilla luciferase reporter. Eighteen h later, the cells were stimulated with or without anti-CD3 plus -CD28 for the final 6 h, and luciferase activity in cell lysates was measured. Results are expressed as relative luciferase units (RLU) normalized to Renilla luciferase activity. Flag-PKC-θ (immunoblotted with anti-PKC-θ) and actin (control for equal protein loading) expression in WCL are shown to the left of the bar graphs. *P < 0.05, **P < 0.01, ***P < 0.001 (two tailed, unpaired Student’s t-test) (e,f) Immunoblot analysis of the phosphorylation signals of indicated kinases in WCL of PKC-θ-knockdown Jurkat E6.1 cells transfected with empty vectors or the indicated Flag-PKC-θ vectors, stimulated as in (d),. TCR-induced phospho- and total IKKβ, ERK, p38, and JNK in cells lysates were determined by immunoblotting with the indicated Abs. Actin served as a control for equal protein loading. (g) The relative strength of the phosphorylated kinase signal in each sample was calculated and normalized to the signal in unstimulated, empty vector-transfected cells (=1). *P < 0.05, **P < 0.01 (two tailed, unpaired Student’s t-test), PKC-θ WT or 2KR group vs. vector group at each time point. Data are representative of four (d) or three (e,f) and from three (g) independent experiments (mean and s.e.m.).
Supplementary Figure 3 Identification of SUMO E3 ligase for PKC-θ and in vitro PKC-θ kinase assay.
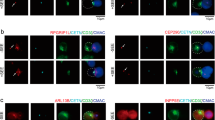
(a) In vivo sumoylation of PKC-θ immunoprecipitated from HEK293T cells cotransfected with empty vector or the indicated YFP- or Flag-tagged PIAS vectors together with Flag-tagged PKC-θ and Myc-tagged SUMO1. (b) Immunoblot analysis of the association between endogenous PKC-θ and PIASxβ in PKC-θ IPs from Jurkat E6.1 T cells stimulated with anti-CD3 plus -CD28 for the indicated times. (c) Representative confocal analysis of Jurkat E6.1 T cells stimulated with SEE-pulsed (+SEE) or unpulsed CMAC-labeled Raji B cells (blue) for the indicated times. Fixed conjugates were stained with anti-PIASxβ plus Alexa Fluor 488-coupled (green) and anti-PKC-θ plus Alexa Fluor 555-coupled (red) secondary Abs. Magnified images of the areas enclosed by yellow rectangles are shown to the right of the merged images. Original magnification, x63. Scale bar = 5 μm. DIC, differential interference contrast. Data are representative of ~100 cells in each group imaged in 5 independent experiments. (d,e) In vivo sumoylation assay of endogenous PKC-θ immunoprecipitated from Jurkat-TAg cells that were transfected with empty vector or Myc-tagged PIASxβ (d), or from Jurkat E6.1 cells transfected with control (siNC; scrambled siRNA) or PIASxβ-specific siRNA-2 (siPIASxβ-2) (e). The cells were stimulated for the indicated times with anti-CD3 plus -CD28 48 h post-transfection. Cells in (d) were also transfected with a CD28 expression vector. Right panels in (d,e) show the signal strength of sumoylated PKC-θ in stimulated cells normalized to the signal in empty vector (d) or siNC (e) -expressing, unstimulated cells (=1). *P<0.05 (two tailed, unpaired Student’s t-test), PIASxβ-1 group vs. vector group (d), siPIASxβ-2 group vs. siNC group (e). Data are pooled from three independent experiments (mean ± s.e.m.). (f) In vitro kinase assay of the catalytic activities for Myc-PKC-θ-WT, -2KR or –K409R immunoprecipitates (IPs) from HEK293T cells transfected with empty vector or the indicated Myc-tagged PKC-θ plasmids, MBP was used as a substrate in the presence of PtdSer/PMA cofactors (upper panel). Aliquots of the IPs were immunoblotted with anti-Myc to confirm similar expression levels of PKC-θ (bottom panel). A representative data of at least three experiments is shown. (g) Jurkat E6.1 cells were transfected with YFP–EV or YFP–SENP1 and were left unstimulated (0) or stimulated with SEE-pulsed Raji B cells for 5 min. Endogenous PKC-θ was immunoprecipitated, and the in vitro PKC-θ kinase assay was performed in the absence of PtdSer/PMA (upper panel) or in the presence of PtdSer/PMA (bottom panel). Aliquots of the IPs were immunoblotted with anti-PKC-θ to confirm similar protein levels of PKC-θ (middle panel). Data are representative of at least two independent experiments.
Supplementary Figure 4 Confocal imaging analysis of the localization of wild-type PKC-θ and lysine-to-arginine PKC-θ mutants in T cell–B cell conjugates.
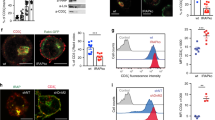
(a,b) Jurkat E6.1 T cells transfected with the indicated Myc-tagged PKC-θ vectors were stimulated with unpulsed or SEE-pulsed CMAC-labeled Raji B cells (blue) for the indicated times. Fixed conjugates were stained with anti-Myc plus Alexa Fluor 488-coupled (green) and anti-CD28 plus Alexa Fluor 594-coupled (red) secondary Abs, and imaged by confocal microcopy. Original magnification, x63. Magnified images of the areas enclosed by yellow rectangles in (a) as well as z-stacks of the x-y sections are shown in (b). Scale bar = 5 μm (a) or 2 μm (b). (c) The accumulation of transfected PKC-θ proteins within the IS after 2 or 30 min of stimulation was quantitated by the length ratio (length of PKC-θ/length of the IS at the middle Z-section of the T-B interface). Each dot represents an individual T cell. Horizontal lines represent the population mean and s.e.m. *P < 0.001; n.s., not significant (one-way ANOVA with Bonferroni’s post-hoc test). A representative of three independent experiments is shown. (d) Confocal analysis of Jurkat E6.1 T cells co-transfected with siPKC-θ and Myc–PKC-θ–K409R, stimulated with SEE-pulsed Raji B cells for 30 min and stained as in (a). Scale bar = 2 μm. (e) Quantification of the localization of PKC-θ aggregated in the immunological synapse (IS aggregation) or not (No IS aggregation) in T-B conjugates as described in a (n = ~50), mean, *P < 0.001 (one-way ANOVA with Bonferroni’s post-hoc test). Data are from 2 independent experiments. (f) Confocal analysis of Jurkat E6.1 T cells stably expressing shNC or shPIASxβ-2 and stimulated as in (a,b) for 30 min. Fixed conjugates were stained with anti-CD28 plus Alexa Fluor 594-coupled (red) and anti-PKC-θ plus Alexa Fluor 647-coupled (magenta) secondary Abs. Scale bar = 5 μm. (g) Scatter plots showing correlation between fluorescent intensity of PKC-θ per unit length (Y axis; fluorescence intensity of PKC-θ in IS/fluorescence intensity of PKC-θ in whole cell body/length of PKC-θ signal in IS) and accumulation of PKC-θ (X axis; length of PKC-θ signal/length of the IS at middle Z-section of the T-B interface). At 30 min, Y is negatively correlated to X (R=-0.4369, P<0.01; Two-tailed Pearson correlation coefficient). The differences of PKC-θ accumulation and its fluorescence intensity per unit length between shNC and shPIASxβ-2 groups were significant (P < 0.0001 and P<0.05, respectively; two tailed, unpaired Student’s t-test). Approximately 25-35 cells were analyzed in each group. Data are representatives of at least two experiments. (h) Jurkat E6.1 T cells transfected with YFP– or YFP–SENP1 (green) were stimulated with SEE-pulsed Raji B cells for 5 min. Fixed conjugates were stained with anti-PKC-θ plus Alexa Fluor 647-coupled (magenta) and anti-CD28 plus Alexa Fluor 594-coupled (red) secondary Abs, and then imaged by confocal microcopy. Scale bar = 10 μm. (i) Accumulation of PKC-θ within the IS after 5 min of stimulation was calculated as in (g), * P < 0.001(two tailed, unpaired Student’s t-test). Data are representative of at least two independent experiments.
Supplementary Figure 5 Effects of desumoylation on the association of PKC-θ, CD28 and filamin A.
Jurkat E6.1 cells transfected with siPKC-θ together with the indicated Flag-tagged PKC-θ vectors (a,c), or transfected with empty vector or tagged SENP1 (b,d), were stimulated for 5 min with SEE-pulsed Raji B cells at a 1:1 ratio (a-c) or with anti-CD3 plus -CD28 mAbs (d). Cells were lysed 48 h post-transfection in buffers containing 1% digitonin (a,b) or 1% NP-40 (c,d). CD28 and PKC-θ IPs, or WCL were immunoblotted with the indicated Abs. Data are representatives of at least three experiments. (e-g) Jurkat E6.1 cells transfected with the indicated Myc-tagged PKC-θ vectors were stimulated for 30 min with SEE-pulsed Raji B cells. Fixed conjugates were stained with anti-Myc and anti-FLNa Abs followed by Alexa Fluor 488-coupled (green; Myc–PKC-θ) or Alexa Fluor 594-coupled (red; FLNa) secondary Abs, and then imaged by confocal microcopy. Representative x-y (en face), Z-stack and DIC images of ~30 conjugates analyzed in each group are shown (e). The Z-stacks are enlarged x2 relative to the x-y images. Scale bar = 5 μm. Line intensity graphs representing the average pixel intensity along a 5 pixel-wide line drawn from points M to N in (e), and bar graphs depicting the percentage of synapses with pSMAC localization of FLNa are shown in (f) and (g), respectively. Data shown are representative of three independent experiments. ~Eighty cells were analyzed in each group. (h-j) Similar confocal analysis of Jurkat E6.1 T cell lines stably expressing shNC or shPIASxβ-2. Localization of FLNa and endogenous PKC-θ was analyzed as in (e-g). One of three independent experiments is shown. ~Sixty cells were analyzed in each group. n.s., not significant; *P < 0.001 (two-way ANOVA with Bonferroni’s post-hoc test), mean and s.d.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Table 1 (PDF 3819 kb)
Rights and permissions
About this article
Cite this article
Wang, XD., Gong, Y., Chen, ZL. et al. TCR-induced sumoylation of the kinase PKC-θ controls T cell synapse organization and T cell activation. Nat Immunol 16, 1195–1203 (2015). https://doi.org/10.1038/ni.3259
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3259
This article is cited by
-
The emerging roles of SUMOylation in the tumor microenvironment and therapeutic implications
Experimental Hematology & Oncology (2023)
-
SUMOylation of PDPK1 Is required to maintain glycolysis-dependent CD4 T-cell homeostasis
Cell Death & Disease (2022)
-
SUMOylation activates large tumour suppressor 1 to maintain the tissue homeostasis during Hippo signalling
Oncogene (2021)
-
Regulatory mechanisms in T cell receptor signalling
Nature Reviews Immunology (2018)
-
SENP3 maintains the stability and function of regulatory T cells via BACH2 deSUMOylation
Nature Communications (2018)