Abstract
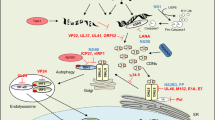
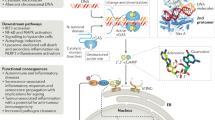
Cytosolic detection of pathogen-derived nucleic acids is critical for the initiation of innate immune defense against diverse bacterial, viral and eukaryotic pathogens. Conversely, inappropriate responses to cytosolic nucleic acids can produce severe autoimmune pathology. The host protein STING has been identified as a central signaling molecule in the innate immune response to cytosolic nucleic acids. STING seems to be especially critical for responses to cytosolic DNA and the unique bacterial nucleic acids called 'cyclic dinucleotides'. Here we discuss advances in the understanding of STING and highlight the many unresolved issues in the field.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Barbalat, R., Ewald, S.E., Mouchess, M.L. & Barton, G.M. Nucleic acid recognition by the innate immune system. Annu. Rev. Immunol. 29, 185–214 (2011).
Marshak-Rothstein, A. Toll-like receptors in systemic autoimmune disease. Nat. Rev. Immunol. 6, 823–835 (2006).
Jin, L. et al. MPYS, a novel membrane tetraspanner, is associated with major histocompatibility complex class II and mediates transduction of apoptotic signals. Mol. Cell Biol. 28, 5014–5026 (2008).
Ishikawa, H. & Barber, G.N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008).
Zhong, B. et al. The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity 29, 538–550 (2008).
Sun, W. et al. ERIS, an endoplasmic reticulum IFN stimulator, activates innate immune signaling through dimerization. Proc. Natl. Acad. Sci. USA 106, 8653–8658 (2009). References 4–6 first identified STING as a signaling molecule involved in cytosolic responses to nucleic acids.
Ishikawa, H., Ma, Z. & Barber, G.N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009).
Sauer, J.D. et al. The N-ethyl-N-nitrosourea-induced Goldenticket mouse mutant reveals an essential function of Sting in the in vivo interferon response to Listeria monocytogenes and cyclic dinucleotides. Infect. Immun. 79, 688–694 (2011).
Jin, L. et al. MPYS is required for IFN response factor 3 activation and type I IFN production in the response of cultured phagocytes to bacterial second messengers cyclic-di-AMP and cyclic-di-GMP. J. Immunol. 187, 2595–2601 (2011).
Chen, H. et al. Activation of STAT6 by STING is critical for antiviral innate immunity. Cell 147, 436–446 (2011). This paper describes activation of STAT6 as a previously unknown signaling pathway downstream of STING.
Rasmussen, S.B. et al. Activation of autophagy by α-herpesviruses in myeloid cells is mediated by cytoplasmic viral DNA through a mechanism dependent on stimulator of IFN genes. J. Immunol. 187, 5268–5276 (2011).
Jones, J.W. et al. Absent in melanoma 2 is required for innate immune recognition of Francisella tularensis. Proc. Natl. Acad. Sci. USA 107, 9771–9776 (2010).
Manzanillo, P.S., Shiloh, M.U., Portnoy, D.A. & Cox, J.S. Mycobacterium tuberculosis activates the DNA-dependent cytosolic surveillance pathway within macrophages. Cell Host Microbe 11, 469–480 (2012).
Watson, R.O., Manzanillo, P.S. & Cox, J.S. Extracellular M. tuberculosis DNA Targets Bacteria for Autophagy by Activating the Host DNA-Sensing Pathway. Cell 150, 803–815 (2012). References 13 and 14 demonstrate a critical role for DNA sensing and STING-dependent signaling in the activation of autophagy and innate immune defense against infection with Mycobacterium tuberculosis.
Parker, D. et al. Streptococcus pneumoniae DNA initiates type I interferon signaling in the respiratory tract. mBio 2, e00016–00011 (2011).
de Almeida, L.A. et al. MyD88 and STING signaling pathways are required for IRF3-mediated IFN-beta induction in response to Brucella abortus infection. PLoS ONE 6, e23135 (2011).
Gratz, N. et al. Type I interferon production induced by Streptococcus pyogenes-derived nucleic acids is required for host protection. PLoS Pathog. 7, e1001345 (2011).
Koppe, U. et al. Streptococcus pneumoniae stimulates a STING- and IFN regulatory factor 3-dependent type I IFN production in macrophages, which regulates RANTES production in macrophages, cocultured alveolar epithelial cells, and mouse lungs. J. Immunol. 188, 811–817 (2012).
Prantner, D., Darville, T. & Nagarajan, U.M. Stimulator of IFN gene is critical for induction of IFN-β during Chlamydia muridarum infection. J. Immunol. 184, 2551–2560 (2010).
Lippmann, J. et al. Dissection of a type I interferon pathway in controlling bacterial intracellular infection in mice. Cell Microbiol. 13, 1668–1682 (2011).
Holm, C.K. et al. Virus-cell fusion as a trigger of innate immunity dependent on the adaptor STING. Nat. Immunol. 13, 737–743 (2012).
DeFilippis, V.R., Alvarado, D., Sali, T., Rothenburg, S. & Fruh, K. Human cytomegalovirus induces the interferon response via the DNA sensor ZBP1. J. Virol. 84, 585–598 (2010).
Sun, L. et al. Coronavirus papain-like proteases negatively regulate antiviral innate immune response through disruption of STING-mediated signaling. PLoS ONE 7, e30802 (2012).
Nazmi, A., Mukhopadhyay, R., Dutta, K. & Basu, A. STING mediates neuronal innate immune response following Japanese encephalitis virus infection. Sci. Rep. 2, 347 (2012).
Stein, S.C. & Falck-Pedersen, E. Sensing adenovirus infection: activation of interferon regulatory factor 3 in RAW 264.7 cells. J. Virol. 86, 4527–4537 (2012).
Yan, N., Regalado-Magdos, A.D., Stiggelbout, B., Lee-Kirsch, M.A. & Lieberman, J. The cytosolic exonuclease TREX1 inhibits the innate immune response to human immunodeficiency virus type 1. Nat. Immunol. 11, 1005–1013 (2010).
Sharma, S. et al. Innate immune recognition of an AT-rich stem-loop DNA motif in the Plasmodium falciparum genome. Immunity 35, 194–207 (2011).
Gall, A. et al. Autoimmunity initiates in nonhematopoietic cells and progresses via lymphocytes in an interferon-dependent autoimmune disease. Immunity 36, 120–131 (2012). This paper identifies a critical role for STING in autoimmune responses to DNA.
Jin, L. et al. Identification and characterization of a loss-of-function human MPYS variant. Genes Immun. 12, 263–269 (2011).
Yin, Q. et al. Cyclic di-GMP sensing via the innate immune signaling protein STING. Mol. Cell 46, 735–745 (2012).
Ouyang, S. et al. Structural analysis of the STING adaptor protein reveals a hydrophobic dimer interface and mode of cyclic di-GMP binding. Immunity 36, 1073–1086 (2012).
Huang, Y.H., Liu, X.Y., Du, X.X., Jiang, Z.F. & Su, X.D. The structural basis for the sensing and binding of cyclic di-GMP by STING. Nat. Struct. Mol. Biol. 19, 728–730 (2012).
Shang, G. et al. Crystal structures of STING protein reveal basis for recognition of cyclic di-GMP. Nat. Struct. Mol. Biol. 19, 725–727 (2012).
Shu, C., Yi, G., Watts, T., Kao, C.C. & Li, P. Structure of STING bound to cyclic di-GMP reveals the mechanism of cyclic dinucleotide recognition by the immune system. Nat. Struct. Mol. Biol. 19, 722–724 (2012). References 30–34 describe crystal structures of STING not bound to ligand and bound to c-di-GMP.
Saitoh, T. et al. Atg9a controls dsDNA-driven dynamic translocation of STING and the innate immune response. Proc. Natl. Acad. Sci. USA 106, 20842–20846 (2009). This paper first connected STING to autophagic responses in cells.
Tamayo, R., Pratt, J.T. & Camilli, A. Roles of cyclic diguanylate in the regulation of bacterial pathogenesis. Annu. Rev. Microbiol. 61, 131–148 (2007).
Witte, G., Hartung, S., Buttner, K. & Hopfner, K.P. Structural biochemistry of a bacterial checkpoint protein reveals diadenylate cyclase activity regulated by DNA recombination intermediates. Mol. Cell 30, 167–178 (2008).
Davies, B.W., Bogard, R.W., Young, T.S. & Mekalanos, J.J. Coordinated regulation of accessory genetic elements produces cyclic di-nucleotides for V. cholerae virulence. Cell 149, 358–370 (2012).
Karaolis, D.K. et al. Bacterial c-di-GMP is an immunostimulatory molecule. J. Immunol. 178, 2171–2181 (2007).
McWhirter, S.M. et al. A host type I interferon response is induced by cytosolic sensing of the bacterial second messenger cyclic-di-GMP. J. Exp. Med. 206, 1899–1911 (2009).
Woodward, J.J., Iavarone, A.T. & Portnoy, D.A. c-di-AMP secreted by intracellular Listeria monocytogenes activates a host type I interferon response. Science 328, 1703–1705 (2010). This paper describes the first bacterial species known to secrete a cyclic dinucleotide ligand that activates STING-dependent signaling.
Burdette, D.L. et al. STING is a direct innate immune sensor of cyclic di-GMP. Nature 478, 515–518 (2011). This paper demonstrates that STING binds directly to cyclic-di-GMP.
Jin, L., Lenz, L.L. & Cambier, J.C. Cellular reactive oxygen species inhibit MPYS induction of IFNβ. PLoS ONE 5, e15142 (2010).
Tanaka, Y. & Chen, Z.J. STING specifies IRF3 phosphorylation by TBK1 in the cytosolic DNA signaling pathway. Sci. Signal. 5, ra20 (2012).
Tsuchida, T. et al. The ubiquitin ligase TRIM56 regulates innate immune responses to intracellular double-stranded DNA. Immunity 33, 765–776 (2010).
Zhong, B. et al. The ubiquitin ligase RNF5 regulates antiviral responses by mediating degradation of the adaptor protein MITA. Immunity 30, 397–407 (2009).
Zhang, J., Hu, M.M., Wang, Y.Y. & Shu, H.B. TRIM32 protein modulates type I interferon induction and cellular antiviral response by targeting MITA/STING protein for K63-linked ubiquitination. J. Biol. Chem. 287, 28646–28655 (2012).
Vance, J.E. Phospholipid synthesis in a membrane fraction associated with mitochondria. J. Biol. Chem. 265, 7248–7256 (1990).
Ishii, K.J. & Akira, S. Innate immune recognition of, and regulation by, DNA. Trends Immunol. 27, 525–532 (2006).
Stetson, D.B. & Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 24, 93–103 (2006).
Chiu, Y.H., Macmillan, J.B. & Chen, Z.J. RNA polymerase III detects cytosolic DNA and induces type I interferons through the RIG-I pathway. Cell 138, 576–591 (2009).
Ablasser, A. et al. RIG-I-dependent sensing of poly(dA:dT) through the induction of an RNA polymerase III–transcribed RNA intermediate. Nat. Immunol. 10, 1065–1072 (2009).
Unterholzner, L. et al. IFI16 is an innate immune sensor for intracellular DNA. Nat. Immunol. 11, 997–1004 (2010).
Babu, M.M., Luscombe, N.M., Aravind, L., Gerstein, M. & Teichmann, S.A. Structure and evolution of transcriptional regulatory networks. Curr. Opin. Struct. Biol. 14, 283–291 (2004).
Takaoka, A. et al. DAI (DLM-1/ZBP1) is a cytosolic DNA sensor and an activator of innate immune response. Nature 448, 501–505 (2007).
Wang, Z. et al. Regulation of innate immune responses by DAI (DLM-1/ZBP1) and other DNA-sensing molecules. Proc. Natl. Acad. Sci. USA 105, 5477–5482 (2008).
Ishii, K.J. et al. TANK-binding kinase-1 delineates innate and adaptive immune responses to DNA vaccines. Nature 451, 725–729 (2008).
Lippmann, J. et al. IFNβ responses induced by intracellular bacteria or cytosolic DNA in different human cells do not require ZBP1 (DLM-1/DAI). Cell. Microbiol. 10, 2579–2588 (2008).
Rebsamen, M. et al. DAI/ZBP1 recruits RIP1 and RIP3 through RIP homotypic interaction motifs to activate NF-κB. EMBO Rep. 10, 916–922 (2009).
Furr, S.R., Chauhan, V.S., Moerdyk-Schauwecker, M.J. & Marriott, I. A role for DNA-dependent activator of interferon regulatory factor in the recognition of herpes simplex virus type 1 by glial cells. J. Neuroinflammation 8, 99–111 (2011).
Chen, Q.Y. et al. DNA-dependent activator of interferon-regulatory factors inhibits hepatitis B virus replication. World J. Gastroenterol. 18, 2850–2858 (2012).
Upton, J.W., Kaiser, W.J. & Mocarski, E.S. DAI/ZBP1/DLM-1 complexes with RIP3 to mediate virus-induced programmed necrosis that is targeted by murine cytomegalovirus vIRA. Cell Host Microbe 11, 290–297 (2012).
Fuller-Pace, F.V. DExD/H box RNA helicases: multifunctional proteins with important roles in transcriptional regulation. Nucleic Acids Res. 34, 4206–4215 (2006).
Zhang, Z. et al. The helicase DDX41 senses intracellular DNA mediated by the adaptor STING in dendritic cells. Nat. Immunol. 12, 959–965 (2011).
Parvatiyar, K. et al. The helicase DDX41 recognizes the bacterial secondary messengers cyclic di-GMP and cyclic di-AMP to activate a type I interferon immune response. Nat. Immunol. 12, 1155–1161 (2012).
Kis-Toth, K., Szanto, A., Thai, T.H. & Tsokos, G.C. Cytosolic DNA-activated human dendritic cells are potent activators of the adaptive immune response. J. Immunol. 187, 1222–1234 (2011).
Conrady, C.D., Zheng, M., Fitzgerald, K.A., Liu, C. & Carr, D.J. Resistance to HSV-1 infection in the epithelium resides with the novel innate sensor, IFI-16. Mucosal Immunol. 5, 173–183 (2012).
Choubey, D. & Lengyel, P. Interferon action: nucleolar and nucleoplasmic localization of the interferon-inducible 72-kD protein that is encoded by the Ifi 204 gene from the gene 200 cluster. J. Cell Biol. 116, 1333–1341 (1992).
Li, T., Diner, B.A., Chen, J. & Cristea, I.M. Acetylation modulates cellular distribution and DNA sensing ability of interferon-inducible protein IFI16. Proc. Natl. Acad. Sci. USA 109, 10558–10563 (2012).
Kerur, N. et al. IFI16 acts as a nuclear pathogen sensor to induce the inflammasome in response to Kaposi sarcoma-associated herpesvirus infection. Cell Host Microbe 9, 363–375 (2011).
Gariano, G.R. et al. The intracellular DNA sensor IFI16 gene acts as restriction factor for human cytomegalovirus replication. PLoS Pathog. 8, e1002498 (2012).
Yang, P. et al. The cytosolic nucleic acid sensor LRRFIP1 mediates the production of type I interferon via a β-catenin-dependent pathway. Nat. Immunol. 11, 487–494 (2010).
Kim, T. et al. Aspartate-glutamate-alanine-histidine box motif (DEAH)/RNA helicase A helicases sense microbial DNA in human plasmacytoid dendritic cells. Proc. Natl. Acad. Sci. USA 107, 15181–15186 (2010).
Zhang, Z., Yuan, B., Lu, N., Facchinetti, V. & Liu, Y.J. DHX9 pairs with IPS-1 to sense double-stranded RNA in myeloid dendritic cells. J. Immunol. 187, 4501–4508 (2011).
Zhang, Z. et al. DDX1, DDX21, and DHX36 helicases form a complex with the adaptor molecule TRIF to sense dsRNA in dendritic cells. Immunity 34, 866–878 (2011).
Zhang, X. et al. Cutting edge: Ku70 is a novel cytosolic DNA sensor that induces type III rather than type I IFN. J. Immunol. 186, 4541–4545 (2011).
Acknowledgements
We thank our colleagues in the field of STING biology and members of the Vance and Barton laboratories for discussions. Supported by the US National Institutes of Health (AI091100 to D.L.B., and AI063302, AI075039 and AI082357 to R.E.V.) and the Burroughs Wellcome Fund (R.E.V.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Burdette, D., Vance, R. STING and the innate immune response to nucleic acids in the cytosol. Nat Immunol 14, 19–26 (2013). https://doi.org/10.1038/ni.2491
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2491
This article is cited by
-
Cytosolic DNA sensors in neurodegenerative diseases: from physiological defenders to pathological culprits
EMBO Molecular Medicine (2024)
-
STING mediates experimental osteoarthritis and mechanical allodynia in mouse
Arthritis Research & Therapy (2023)
-
PTK2B promotes TBK1 and STING oligomerization and enhances the STING-TBK1 signaling
Nature Communications (2023)
-
cGAMP-activated cGAS–STING signaling: its bacterial origins and evolutionary adaptation by metazoans
Nature Structural & Molecular Biology (2023)
-
The Mystery of Cancer Resistance: A Revelation Within Nature
Journal of Molecular Evolution (2023)