Abstract
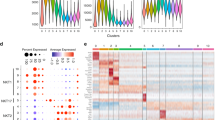
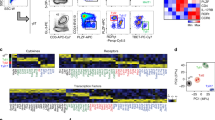
Invariant natural killer T cells (iNKT cells) are innate-like T lymphocytes that act as critical regulators of the immune response. To better characterize this population, we profiled gene expression in iNKT cells during ontogeny and in peripheral subsets as part of the Immunological Genome Project. High-resolution comparative transcriptional analyses defined developmental and subset-specific programs of gene expression by iNKT cells. In addition, we found that iNKT cells shared an extensive transcriptional program with NK cells, similar in magnitude to that shared with major histocompatibility complex (MHC)-restricted T cells. Notably, the program shared by NK cells and iNKT cells also operated constitutively in γδ T cells and in adaptive T cells after activation. Together our findings highlight a core effector program regulated distinctly in innate and adaptive lymphocytes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Heng, T.S.P. et al. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Cohen, N.R., Garg, S. & Brenner, M.B. Antigen presentation by CD1 lipids, T cells, and NKT cells in microbial immunity. Adv. Immunol. 102, 1–94 (2009).
Bendelac, A., Savage, P.B. & Teyton, L. The biology of NKT cells. Annu. Rev. Immunol. 25, 297–336 (2007).
Godfrey, D.I., Stankovic, S. & Baxter, A.G. Raising the NKT cell family. Nat. Immunol. 11, 197–206 (2010).
Coquet, J.M. et al. Diverse cytokine production by NKT cell subsets and identification of an IL-17-producing CD4–NK1.1- NKT cell population. Proc. Natl. Acad. Sci. USA 105, 11287–11292 (2008).
Cerundolo, V., Silk, J.D., Masri, S.H. & Salio, M. Harnessing invariant NKT cells in vaccination strategies. Nat. Rev. Immunol. 9, 28–38 (2009).
Makino, Y., Kanno, R., Ito, T., Higashino, K. & Taniguchi, M. Predominant expression of invariant Vα14+ TCR α chain in NK1.1+ T cell populations. Int. Immunol. 7, 1157–1161 (1995).
Bendelac, A. Mouse NK1+ T cells. Curr. Opin. Immunol. 7, 367–374 (1995).
Godfrey, D.I., MacDonald, H.R., Kronenberg, M., Smyth, M.J. & Van Kaer, L. NKT cells: what's in a name? Nat. Rev. Immunol. 4, 231–237 (2004).
Grégoire, C. et al. The trafficking of natural killer cells. Immunol. Rev. 220, 169–182 (2007).
Matsuda, J.L. et al. Tracking the response of natural killer T cells to a glycolipid antigen using CD1d tetramers. J. Exp. Med. 192, 741–754 (2000).
Cooper, M.A. et al. In vivo evidence for a dependence on interleukin 15 for survival of natural killer cells. Blood 100, 3633–3638 (2002).
Matsuda, J.L. et al. Homeostasis of V alpha 14i NKT cells. Nat. Immunol. 3, 966–974 (2002).
Brigl, M. & Brenner, M.B. How invariant natural killer T cells respond to infection by recognizing microbial or endogenous lipid antigens. Semin. Immunol. 22, 79–86 (2010).
Lanier, L.L. Evolutionary struggles between NK cells and viruses. Nat. Rev. Immunol. 8, 259–268 (2008).
Kuylenstierna, C. et al. NKG2D performs two functions in invariant NKT cells: direct TCR-independent activation of NK-like cytolysis and co-stimulation of activation by CD1d. Eur. J. Immunol. 41, 1913–1923 (2011).
Kawamura, T. et al. NKG2A inhibits invariant NKT cell activation in hepatic injury. J. Immunol. 182, 250–258 (2009).
Maeda, M., Lohwasser, S., Yamamura, T. & Takei, F. Regulation of NKT cells by Ly49: analysis of primary NKT cells and generation of NKT cell line. J. Immunol. 167, 4180–4186 (2001).
Ota, T. et al. IFN-γ-mediated negative feedback regulation of NKT-cell function by CD94/NKG2. Blood 106, 184–192 (2005).
Brennan, P.J. et al. Invariant natural killer T cells recognize lipid self antigen induced by microbial danger signals. Nat. Immunol. 12, 1202–1211 (2011).
Paget, C. et al. Activation of invariant NKT cells by toll-like receptor 9-stimulated dendritic cells requires type I interferon and charged glycosphingolipids. Immunity 27, 597–609 (2007).
Salio, M. et al. Modulation of human natural killer T cell ligands on TLR-mediated antigen-presenting cell activation. Proc. Natl. Acad. Sci. USA 104, 20490–20495 (2007).
Reschner, A., Hubert, P., Delvenne, P., Boniver, J. & Jacobs, N. Innate lymphocyte and dendritic cell cross-talk: a key factor in the regulation of the immune response. Clin. Exp. Immunol. 152, 219–226 (2008).
Andrews, D.M., Scalzo, A.A., Yokoyama, W.M., Smyth, M.J. & Degli-Esposti, M.A. Functional interactions between dendritic cells and NK cells during viral infection. Nat. Immunol. 4, 175–181 (2003).
Brigl, M., Bry, L., Kent, S.C., Gumperz, J.E. & Brenner, M.B. Mechanism of CD1d-restricted natural killer T cell activation during microbial infection. Nat. Immunol. 4, 1230–1237 (2003).
Fernandez, N.C. et al. Dendritic cells directly trigger NK cell functions: cross-talk relevant in innate anti-tumor immune responses in vivo. Nat. Med. 5, 405–411 (1999).
Vincent, M.S. et al. CD1-dependent dendritic cell instruction. Nat. Immunol. 3, 1163–1168 (2002).
Walzer, T., Dalod, M., Vivier, E. & Zitvogel, L. Natural killer cell-dendritic cell crosstalk in the initiation of immune responses. Expert Opin. Biol. Ther. 5 Suppl 1, S49–S59 (2005).
Savage, A.K. et al. The transcription factor PLZF directs the effector program of the NKT cell lineage. Immunity 29, 391–403 (2008).
Kovalovsky, D. et al. The BTB-zinc finger transcriptional regulator PLZF controls the development of invariant natural killer T cell effector functions. Nat. Immunol. 9, 1055–1064 (2008).
Yu, S. & Cantorna, M.T. The vitamin D receptor is required for iNKT cell development. Proc. Natl. Acad. Sci. USA 105, 5207–5212 (2008).
Gumperz, J.E., Miyake, S., Yamamura, T. & Brenner, M.B. Functionally distinct subsets of CD1d-restricted natural killer T cells revealed by CD1d tetramer staining. J. Exp. Med. 195, 625–636 (2002).
Watarai, H. et al. Development and function of invariant natural killer T cells producing Th2- and Th17-cytokines. PLoS Biol. 10, e1001255 (2012).
Crowe, N.Y. et al. Differential antitumor immunity mediated by NKT cell subsets in vivo. J. Exp. Med. 202, 1279–1288 (2005).
Brigl, M. et al. Innate and cytokine-driven signals, rather than microbial antigens, dominate in natural killer T cell activation during microbial infection. J. Exp. Med. 208, 1163–1177 (2011).
Johnston, B., Kim, C.H., Soler, D., Emoto, M. & Butcher, E.C. Differential chemokine responses and homing patterns of murine TCRαβ NKT cell subsets. J. Immunol. 171, 2960–2969 (2003).
Townsend, M.J. et al. T-bet regulates the terminal maturation and homeostasis of NK and Vα14i NKT cells. Immunity 20, 477–494 (2004).
Thomas, P.D. et al. PANTHER: a browsable database of gene products organized by biological function, using curated protein family and subfamily classification. Nucleic Acids Res. 31, 334–341 (2003).
O'Brien, R.L. & Born, W.K. γδ T cell subsets: a link between TCR and function? Semin. Immunol. 22, 193–198 (2010).
Fahrer, A.M. et al. Attributes of gammadelta intraepithelial lymphocytes as suggested by their transcriptional profile. Proc. Natl. Acad. Sci. USA 98, 10261–10266 (2001).
Shires, J., Theodoridis, E. & Hayday, A.C. Biological insights into TCRγδ+ and TCRαβ+ intraepithelial lymphocytes provided by serial analysis of gene expression (SAGE). Immunity 15, 419–434 (2001).
Vivier, E. & Anfossi, N. Inhibitory NK-cell receptors on T cells: witness of the past, actors of the future. Nat. Rev. Immunol. 4, 190–198 (2004).
Gattinoni, L. et al. A human memory T cell subset with stem cell-like properties. Nat. Med. 17, 1290–1297 (2011).
Matsuda, J.L. et al. T-bet concomitantly controls migration, survival, and effector functions during the development of Valpha14i NKT cells. Blood 107, 2797–2805 (2006).
Gordy, L.E. et al. IL-15 regulates homeostasis and terminal maturation of NKT cells. J. Immunol. 187, 6335–6345 (2011).
Kastner, P. et al. Bcl11b represses a mature T-cell gene expression program in immature CD4+CD8+ thymocytes. Eur. J. Immunol. 40, 2143–2154 (2010).
Yue, X., Izcue, A. & Borggrefe, T. Essential role of mediator subunit Med1 in invariant natural killer T-cell development. Proc. Natl. Acad. Sci. USA 108, 17105–17110 (2011).
Sullivan, B.M., Juedes, A., Szabo, S.J., von Herrath, M. & Glimcher, L.H. Antigen-driven effector CD8 T cell function regulated by T-bet. Proc. Natl. Acad. Sci. USA 100, 15818–15823 (2003).
Inagaki-Ohara, K., Nishimura, H., Mitani, A. & Yoshikai, Y. Interleukin-15 preferentially promotes the growth of intestinal intraepithelial lymphocytes bearing gamma delta T cell receptor in mice. Eur. J. Immunol. 27, 2885–2891 (1997).
Honma, S. et al. Dec1 and Dec2 are regulators of the mammalian molecular clock. Nature 419, 841–844 (2002).
Miyazaki, K. et al. The role of the basic helix-loop-helix transcription factor Dec1 in the regulatory T cells. J. Immunol. 185, 7330–7339 (2010).
Sun, H., Lu, B., Li, R.Q., Flavell, R.A. & Taneja, R. Defective T cell activation and autoimmune disorder in Stra13-deficient mice. Nat. Immunol. 2, 1040–1047 (2001).
Weber, B.N. et al. A critical role for TCF-1 in T-lineage specification and differentiation. Nature 476, 63–68 (2011).
Willinger, T. et al. Human naive CD8 T cells down-regulate expression of the WNT pathway transcription factors lymphoid enhancer binding factor 1 and transcription factor 7 (T cell factor-1) following antigen encounter in vitro and in vivo. J. Immunol. 176, 1439–1446 (2006).
Gattinoni, L. et al. Wnt signaling arrests effector T cell differentiation and generates CD8+ memory stem cells. Nat. Med. 15, 808–813 (2009).
Marshall, H.D. et al. Differential expression of Ly6C and T-bet distinguish effector and memory Th1 CD4+ cell properties during viral infection. Immunity 35, 633–646 (2011).
Huang da, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Acknowledgements
We thank the tetramer facility of the US National Institutes of Health for ongoing support; and S. Raychaudhuri, X. Hu and H. Li for advice, discussions and technical assistance. Supported by the US National Institutes of Health (R01AI063428 to M.B.B., T32AI007306 to P.J.B, and R24AI072073 to the ImmGen Project consortium).
Author information
Authors and Affiliations
Consortia
Contributions
N.R.C. primarily wrote the manuscript, conceived of and did experiments and analyzed the data; P.J.B. contributed substantially to the manuscript, conceived of and did experiments and analyzed the data; T.S. conceived of and did experiments and analyzed the data; G.F.W. did experiments; M.B. and J.K. assisted with experimental design and interpretation of the data; M.B.B. substantially contributed to the manuscript and supervised all experimental design, performance and data analysis; and The ImmGen Project Consortium contributed to experimental design and data collection.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Tables 1–16 (PDF 6998 kb)
Rights and permissions
About this article
Cite this article
Cohen, N., Brennan, P., Shay, T. et al. Shared and distinct transcriptional programs underlie the hybrid nature of iNKT cells. Nat Immunol 14, 90–99 (2013). https://doi.org/10.1038/ni.2490
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2490
This article is cited by
-
New cell sources for CAR-based immunotherapy
Biomarker Research (2023)
-
Invariant natural killer T cells in lung diseases
Experimental & Molecular Medicine (2023)
-
Differential location of NKT and MAIT cells within lymphoid tissue
Scientific Reports (2022)
-
PTEN directs developmental and metabolic signaling for innate-like T cell fate and tissue homeostasis
Nature Cell Biology (2022)
-
Gut microbiota-mediated immunomodulation in tumor
Journal of Experimental & Clinical Cancer Research (2021)