Abstract
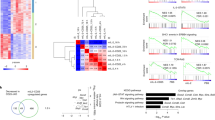
Interleukin 35 (IL-35) belongs to the IL-12 family of heterodimeric cytokines but has a distinct functional profile. IL-35 suppresses T cell proliferation and converts naive T cells into IL-35-producing induced regulatory T cells (iTr35 cells). Here we found that IL-35 signaled through a unique heterodimer of receptor chains IL-12Rβ2 and gp130 or homodimers of each chain. Conventional T cells were sensitive to IL-35-mediated suppression in the absence of one receptor chain but not both receptor chains, whereas signaling through both chains was required for IL-35 expression and conversion into iTr35 cells. Signaling through the IL-35 receptor required the transcription factors STAT1 and STAT4, which formed a unique heterodimer that bound to distinct sites in the promoters of the genes encoding the IL-12 subunits p35 and Ebi3. This unconventional mode of signaling, distinct from that of other members of the IL-12 family, may broaden the spectrum and specificity of IL-35-mediated suppression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Vignali, D.A., Collison, L.W. & Workman, C.J. How regulatory T cells work. Nat. Rev. Immunol. 8, 523–532 (2008).
Zou, W. Regulatory T cells, tumour immunity and immunotherapy. Nat. Rev. Immunol. 6, 295–307 (2006).
O'Shea, J.J. & Paul, W.E. Regulation of TH1 differentiation–controlling the controllers. Nat. Immunol. 3, 506–508 (2002).
Artis, D. et al. The IL-27 receptor (WSX-1) is an inhibitor of innate and adaptive elements of type 2 immunity. J. Immunol. 173, 5626–5634 (2004).
Awasthi, A. et al. A dominant function for interleukin 27 in generating interleukin 10-producing anti-inflammatory T cells. Nat. Immunol. 8, 1380–1389 (2007).
Fitzgerald, D.C. et al. Suppression of autoimmune inflammation of the central nervous system by interleukin 10 secreted by interleukin 27-stimulated T cells. Nat. Immunol. 8, 1372–1379 (2007).
Stumhofer, J.S. et al. Interleukins 27 and 6 induce STAT3-mediated T cell production of interleukin 10. Nat. Immunol. 8, 1363–1371 (2007).
Collison, L.W. et al. IL-35-mediated induction of a potent regulatory T cell population. Nat. Immunol. 11, 1093–1101 (2010).
Collison, L.W., Pillai, M.R., Chaturvedi, V. & Vignali, D.A. Regulatory T cell suppression is potentiated by target T cells in a cell contact, IL-35- and IL-10-dependent manner. J. Immunol. 182, 6121–6128 (2009).
Collison, L.W. et al. The inhibitory cytokine IL-35 contributes to regulatory T-cell function. Nature 450, 566–569 (2007).
Chua, A.O., Wilkinson, V.L., Presky, D.H. & Gubler, U. Cloning and characterization of a mouse IL-12 receptor-β component. J. Immunol. 155, 4286–4294 (1995).
Parham, C. et al. A receptor for the heterodimeric cytokine IL-23 is composed of IL-12Rβ1 and a novel cytokine receptor subunit, IL-23R. J. Immunol. 168, 5699–5708 (2002).
Pflanz, S. et al. WSX-1 and glycoprotein 130 constitute a signal-transducing receptor for IL-27. J. Immunol. 172, 2225–2231 (2004).
Silver, J.S. & Hunter, C.A. gp130 at the nexus of inflammation, autoimmunity, and cancer. J. Leukoc. Biol. 88, 1145–1156 (2010).
Bacon, C.M. et al. Interleukin 12 induces tyrosine phosphorylation and activation of STAT4 in human lymphocytes. Proc. Natl. Acad. Sci. USA 92, 7307–7311 (1995).
Delgoffe, G.M., Murray, P.J. & Vignali, D.A. Interpreting mixed signals: the cell's cytokine conundrum. Curr. Opin. Immunol. 23, 1–7 (2011).
Thierfelder, W.E. et al. Requirement for Stat4 in interleukin-12-mediated responses of natural killer and T cells. Nature 382, 171–174 (1996).
Huber, M. et al. IL-27 inhibits the development of regulatory T cells via STAT3. Int. Immunol. 20, 223–234 (2008).
Owaki, T. et al. STAT3 is indispensable to IL-27-mediated cell proliferation but not to IL-27-induced Th1 differentiation and suppression of proinflammatory cytokine production. J. Immunol. 180, 2903–2911 (2008).
Wu, C. et al. IL-12 receptor β 2 (IL-12Rβ2)-deficient mice are defective in IL-12-mediated signaling despite the presence of high affinity IL-12 binding sites. J. Immunol. 165, 6221–6228 (2000).
Presky, D.H. et al. A functional interleukin 12 receptor complex is composed of two β-type cytokine receptor subunits. Proc. Natl. Acad. Sci. USA 93, 14002–14007 (1996).
Villarino, A. et al. The IL-27R (WSX-1) is required to suppress T cell hyperactivity during infection. Immunity 19, 645–655 (2003).
Schroers, A. et al. Dynamics of the gp130 cytokine complex: a model for assembly on the cellular membrane. Protein Sci. 14, 783–790 (2005).
Skiniotis, G., Lupardus, P.J., Martick, M., Walz, T. & Garcia, K.C. Structural organization of a full-length gp130/LIF-R cytokine receptor transmembrane complex. Mol. Cell 31, 737–748 (2008).
Betz, U.A. & Muller, W. Regulated expression of gp130 and IL-6 receptor α chain in T cell maturation and activation. Int. Immunol. 10, 1175–1184 (1998).
Charlot-Rabiega, P., Bardel, E., Dietrich, C., Kastelein, R. & Devergne, O. Signaling events involved in interleukin 27 (IL-27)-induced proliferation of human naive CD4+ T cells and B cells. J. Biol. Chem. 286, 27350–27362 (2011).
Hibbert, L., Pflanz, S., De Waal Malefyt, R. & Kastelein, R.A. IL-27 and IFN-α signal via Stat1 and Stat3 and induce T-bet and IL-12Rβ2 in naive T cells. J. Interferon Cytokine Res. 23, 513–522 (2003).
Takeda, A. et al. Cutting edge: role of IL-27/WSX-1 signaling for induction of T-bet through activation of STAT1 during initial Th1 commitment. J. Immunol. 170, 4886–4890 (2003).
Liao, W., Lin, J.X., Wang, L., Li, P. & Leonard, W.J. Modulation of cytokine receptors by IL-2 broadly regulates differentiation into helper T cell lineages. Nat. Immunol. 12, 551–559 (2011).
O'Shea, J.J. & Murray, P.J. Cytokine signaling modules in inflammatory responses. Immunity 28, 477–487 (2008).
Ho, H.H. & Ivashkiv, L.B. Role of STAT3 in type I interferon responses. Negative regulation of STAT1-dependent inflammatory gene activation. J. Biol. Chem. 281, 14111–14118 (2006).
Yamamoto, K., Miura, O., Hirosawa, S. & Miyasaka, N. Binding sequence of STAT4: STAT4 complex recognizes the IFN-γ activation site (GAS)-like sequence (T/A)TTCC(C/G)GGAA(T/A). Biochem. Biophys. Res. Commun. 233, 126–132 (1997).
Ehret, G.B. et al. DNA binding specificity of different STAT proteins. Comparison of in vitro specificity with natural target sites. J. Biol. Chem. 276, 6675–6688 (2001).
Yu, Q., Thieu, V.T. & Kaplan, M.H. Stat4 limits DNA methyltransferase recruitment and DNA methylation of the IL-18Rα gene during Th1 differentiation. EMBO J. 26, 2052–2060 (2007).
Ramsauer, K. et al. Distinct modes of action applied by transcription factors STAT1 and IRF1 to initiate transcription of the IFN-γ-inducible gbp2 gene. Proc. Natl. Acad. Sci. USA 104, 2849–2854 (2007).
Boulanger, M.J., Chow, D.C., Brevnova, E.E. & Garcia, K.C. Hexameric structure and assembly of the interleukin-6/IL-6 α-receptor/gp130 complex. Science 300, 2101–2104 (2003).
Stern, A.S., Gubler, U., Presky, D.H. & Magram, J. Structural and functional aspects of the IL-12 receptor complex. Chem. Immunol. 68, 23–37 (1997).
Huyton, T. et al. An unusual cytokine:Ig-domain interaction revealed in the crystal structure of leukemia inhibitory factor (LIF) in complex with the LIF receptor. Proc. Natl. Acad. Sci. USA 104, 12737–12742 (2007).
Lucas, S., Ghilardi, N., Li, J. & de Sauvage, F.J. IL-27 regulates IL-12 responsiveness of naive CD4+ T cells through Stat1-dependent and -independent mechanisms. Proc. Natl. Acad. Sci. USA 100, 15047–15052 (2003).
Canda-Sánchez, A. et al. Differential distribution of both IL-12Rβ chains in the plasma membrane of human T cells. J. Membr. Biol. 227, 1–12 (2009).
Maldonado, R.A., Irvine, D.J., Schreiber, R. & Glimcher, L.H. A role for the immunological synapse in lineage commitment of CD4 lymphocytes. Nature 431, 527–532 (2004).
Szabo, S.J., Dighe, A.S., Gubler, U. & Murphy, K.M. Regulation of the interleukin (IL)-12Rβ2 subunit expression in developing T helper 1 (Th1) and Th2 cells. J. Exp. Med. 185, 817–824 (1997).
Grohmann, U. et al. IL-12 acts directly on DC to promote nuclear localization of NF-κB and primes DC for IL-12 production. Immunity 9, 315–323 (1998).
Andersson, J. et al. CD4+FoxP3+ regulatory T cells confer infectious tolerance in a TGF-β-dependent manner. J. Exp. Med. 205, 1975–1981 (2008).
Pot, C. et al. Cutting edge: IL-27 induces the transcription factor c-Maf, cytokine IL-21, and the costimulatory receptor ICOS that coordinately act together to promote differentiation of IL-10-producing Tr1 cells. J. Immunol. 183, 797–801 (2009).
Pistoia, V., Cocco, C. & Airoldi, I. Interleukin-12 receptor β2: from cytokine receptor to gatekeeper gene in human B-cell malignancies. J. Clin. Oncol. 27, 4809–4816 (2009).
Hirschfield, G.M. et al. Primary biliary cirrhosis associated with HLA, IL12A, and IL12RB2 variants. N. Engl. J. Med. 360, 2544–2555 (2009).
Matsui, E. et al. Mutations of the IL-12 receptor β2 chain gene in atopic subjects. Biochem. Biophys. Res. Commun. 266, 551–555 (1999).
Remmers, E.F. et al. Genome-wide association study identifies variants in the MHC class I, IL10, and IL23R–IL12RB2 regions associated with Behçet's disease. Nat. Genet. 42, 698–702 (2010).
Xu, Y. et al. Crystal structure of the entire ectodomain of gp130: insights into the molecular assembly of the tall cytokine receptor complexes. J. Biol. Chem. 285, 21214–21218 (2010).
Acknowledgements
We thank D. Fairweather and J.A. Frisancho (Johns Hopkins University) for spleens and lymph nodes from Il12rb1−/− mice; M. Karin and S. Grivennikov (University of California at San Diego) for Il6stΔT mice; C. Hunter and J. Stumhofer (University of Pennsylvania) for Il27ra−/− mice; A. Satoskar and P. Reville (Ohio State University) for Stat1−/− mice; C. Drake and H.R. Yen (Johns Hopkins University) for Stat3ΔT mice; R. McEver (University of Oklahoma Health Sciences Center) for mice used to establish the colony of CD4cre × Il6stfl/fl mice at St. Jude Children's Research Hospital; M.J. Turk (Dartmouth College) for B16-F10 melanoma; J.A. Frisancho, S. Grivennikov, M. Karin, J. Stumhofer, P. Reville and H.R. Yen for assistance with the collection and shipping of spleens and lymph nodes; J. Partridge and P. Brindle for assistance in designing ChIP experiments; K. Forbes, A. Castellaw and A. Krause for the maintenance, breeding and genotyping of mouse colonies; R. Cross, G. Lennon and S. Morgan for flow cytometry; the staff of the Shared Animal Resource Center at St. Jude Children's Research Hospital for the animal husbandry; and the Hartwell Center for Biotechnology and Bioinformatics at St. Jude Children's Research Hospital for the synthesis of real-time PCR primers and probes. Supported by the US National Institutes of Health (R01 AI091977 to D.A.A.V. and F32 AI072816 to L.W.C.), NovoNordisk (D.A.A.V.), the National Cancer Institute Comprehensive Cancer Center (CA21765 to D.A.A.V.) and the American Lebanese Syrian Associated Charities (D.A.A.V.).
Author information
Authors and Affiliations
Contributions
L.W.C. designed (with help from D.A.A.V.) and executed a substantial proportion of the experiments, analyzed data and wrote the manuscript; G.M.D. did STAT coimmunoprecipitation and ChIP–ChIP-reChIP experiments, cytokine-receptor coimmunoprecipitation experiments, some phosphorylated STAT analysis and functional assays and wrote the manuscript; C.S.G. did confocal microscopy–FRET experiments; K.M.V. generated all constructs; V.C. did TH1 and TH2 polarization for receptor analysis; D.F., A.R.S., C.A.H. and C.G.D. provided mice; K.C.G. did structural modeling of cytokine receptor complexes; P.J.M. analyzed phosphorylated STAT by immunoblot; C.A.H., P.M., K.C.G., C.G.D. and K.M.V. commented on the manuscript; and D.A.A.V. conceived of the research, directed the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.A.A.V., L.W.C. and K.M.V. have submitted (pending) patents and are entitled to a share of the income generated from licensing of those patent rights for commercial development; D.A.A.V. received support from a sponsored research agreement with NovoNordisk.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Table 1 (PDF 130 kb)
Rights and permissions
About this article
Cite this article
Collison, L., Delgoffe, G., Guy, C. et al. The composition and signaling of the IL-35 receptor are unconventional. Nat Immunol 13, 290–299 (2012). https://doi.org/10.1038/ni.2227
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2227
This article is cited by
-
Tumor-derived interleukin 35 mediates the dissemination of gemcitabine resistance in pancreatic adenocarcinoma
Oncogene (2024)
-
Interleukin-35 -producing B cells rescues inflammatory bowel disease in a mouse model via STAT3 phosphorylation and intestinal microbiota modification
Cell Death Discovery (2023)
-
A novel cytokine consisting of the p40 and EBI3 subunits suppresses experimental autoimmune arthritis via reciprocal regulation of Th17 and Treg cells
Cellular & Molecular Immunology (2022)
-
Interleukin-35 promotes Breg expansion and interleukin-10 production in CD19+ B cells in patients with ankylosing spondylitis
Clinical Rheumatology (2022)
-
Hairy cell leukemia: a specific 17-gene expression signature points to new targets for therapy
Journal of Cancer Research and Clinical Oncology (2022)