Abstract
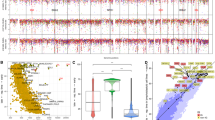
The cytidine deaminase AID hypermutates immunoglobulin genes but can also target oncogenes, leading to tumorigenesis. The extent of AID's promiscuity and its predilection for immunoglobulin genes are unknown. We report here that AID interacted broadly with promoter-proximal sequences associated with stalled polymerases and chromatin-activating marks. In contrast, genomic occupancy of replication protein A (RPA), an AID cofactor, was restricted to immunoglobulin genes. The recruitment of RPA to the immunoglobulin loci was facilitated by phosphorylation of AID at Ser38 and Thr140. We propose that stalled polymerases recruit AID, thereby resulting in low frequencies of hypermutation across the B cell genome. Efficient hypermutation and switch recombination required AID phosphorylation and correlated with recruitment of RPA. Our findings provide a rationale for the oncogenic role of AID in B cell malignancy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Stavnezer, J., Guikema, J.E. & Schrader, C.E. Mechanism and regulation of class switch recombination. Annu. Rev. Immunol. 26, 261–292 (2008).
Honjo, T., Kinoshita, K. & Muramatsu, M. Molecular mechanism of class switch recombination: linkage with somatic hypermutation. Annu. Rev. Immunol. 20, 165–196 (2002).
Revy, P. et al. Activation-induced cytidine deaminase (AID) deficiency causes the autosomal recessive form of the Hyper-IgM syndrome (HIGM2). Cell 102, 565–575 (2000).
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–563 (2000).
Di Noia, J.M. & Neuberger, M.S. Molecular mechanisms of antibody somatic hypermutation. Annu. Rev. Biochem. 76, 1–22 (2007).
Peled, J.U. et al. The biochemistry of somatic hypermutation. Annu. Rev. Immunol. 26, 481–511 (2008).
Delker, R.K., Fugmann, S.D. & Papavasiliou, F.N. A coming-of-age story: activation-induced cytidine deaminase turns 10. Nat. Immunol. 10, 1147–1153 (2009).
Peters, A. & Storb, U. Somatic hypermutation of immunoglobulin genes is linked to transcription initiation. Immunity 4, 57–65 (1996).
Robbiani, D.F. et al. AID produces DNA double-strand breaks in non-Ig genes and mature B cell lymphomas with reciprocal chromosome translocations. Mol. Cell 36, 631–641 (2009).
Shen, H.M., Peters, A., Baron, B., Zhu, X. & Storb, U. Mutation of BCL-6 gene in normal B cells by the process of somatic hypermutation of Ig genes. Science 280, 1750–1752 (1998).
Pasqualucci, L. et al. BCL-6 mutations in normal germinal center B cells: evidence of somatic hypermutation acting outside Ig loci. Proc. Natl. Acad. Sci. USA 95, 11816–11821 (1998).
Liu, M. et al. Two levels of protection for the B cell genome during somatic hypermutation. Nature 451, 841–845 (2008).
Gordon, M.S., Kanegai, C.M., Doerr, J.R. & Wall, R. Somatic hypermutation of the B cell receptor genes B29 (Igβ, CD79b) and mb1 (Igα, CD79a). Proc. Natl. Acad. Sci. USA 100, 4126–4131 (2003).
Nussenzweig, A. & Nussenzweig, M.C. Origin of chromosomal translocations in lymphoid cancer. Cell 141, 27–38 (2010).
Klemm, L. et al. The B cell mutator AID promotes B lymphoid blast crisis and drug resistance in chronic myeloid leukemia. Cancer Cell 16, 232–245 (2009).
Lin, C. et al. Nuclear receptor-induced chromosomal proximity and DNA breaks underlie specific translocations in cancer. Cell 139, 1069–1083 (2009).
Matsumoto, Y. et al. Helicobacter pylori infection triggers aberrant expression of activation-induced cytidine deaminase in gastric epithelium. Nat. Med. 13, 470–476 (2007).
Morgan, H.D., Dean, W., Coker, H.A., Reik, W. & Petersen-Mahrt, S.K. Activation-induced cytidine deaminase deaminates 5-methylcytosine in DNA and is expressed in pluripotent tissues: implications for epigenetic reprogramming. J. Biol. Chem. 279, 52353–52360 (2004).
Bhutani, N. et al. Reprogramming towards pluripotency requires AID-dependent DNA demethylation. Nature (2010).
Popp, C. et al. Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature 463, 1101–1105 (2010).
Rai, K. et al. DNA demethylation in zebrafish involves the coupling of a deaminase, a glycosylase, and gadd45. Cell 135, 1201–1212 (2008).
Chaudhuri, J., Khuong, C. & Alt, F.W. Replication protein A interacts with AID to promote deamination of somatic hypermutation targets. Nature 430, 992–998 (2004).
Basu, U. et al. The AID antibody diversification enzyme is regulated by protein kinase A phosphorylation. Nature 438, 508–511 (2005).
Vuong, B.Q. et al. Specific recruitment of protein kinase A to the immunoglobulin locus regulates class-switch recombination. Nat. Immunol. 10, 420–426 (2009).
McBride, K.M. et al. Regulation of hypermutation by activation-induced cytidine deaminase phosphorylation. Proc. Natl. Acad. Sci. USA 103, 8798–8803 (2006).
Pasqualucci, L., Kitaura, Y., Gu, H. & Dalla-Favera, R. PKA-mediated phosphorylation regulates the function of activation-induced deaminase (AID) in B cells. Proc. Natl. Acad. Sci. USA 103, 395–400 (2006).
Wold, M.S. Replication protein A: a heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu. Rev. Biochem. 66, 61–92 (1997).
McBride, K.M., Barreto, V., Ramiro, A.R., Stavropoulos, P. & Nussenzweig, M.C. Somatic hypermutation is limited by CRM1-dependent nuclear export of activation-induced deaminase. J. Exp. Med. 199, 1235–1244 (2004).
Nambu, Y. et al. Transcription-coupled events associating with immunoglobulin switch region chromatin. Science 302, 2137–2140 (2003).
Pavri, R. et al. Activation-induced cytidine deaminase targets DNA at sites of RNA polymerase II stalling by interaction with Spt5. Cell 143, 122–133 (2010).
Pasqualucci, L. et al. Hypermutation of multiple proto-oncogenes in B-cell diffuse large-cell lymphomas. Nature 412, 341–346 (2001).
Ji, Y. et al. The in vivo pattern of binding of RAG1 and RAG2 to antigen receptor loci. Cell 141, 419–431 (2010).
Lenz, G. et al. Aberrant immunoglobulin class switch recombination and switch translocations in activated B cell-like diffuse large B cell lymphoma. J. Exp. Med. 204, 633–643 (2007).
Kuchen, S. et al. Regulation of microRNA expression and abundance during lymphopoiesis. Immunity 32, 828–839 (2010).
Storb, U. et al. Targeting of AID to immunoglobulin genes. Adv. Exp. Med. Biol. 596, 83–91 (2007).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Mikkelsen, T.S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007).
Rahl, P.B. et al. c-Myc regulates transcriptional pause release. Cell 141, 432–445 (2010).
Fuda, N.J., Ardehali, M.B. & Lis, J.T. Defining mechanisms that regulate RNA polymerase II transcription in vivo. Nature 461, 186–192 (2009).
Buratowski, S. Transcription. Gene expression–where to start? Science 322, 1804–1805 (2008).
Wang, L., Wuerffel, R., Feldman, S., Khamlichi, A.A. & Kenter, A.L. S region sequence, RNA polymerase II, and histone modifications create chromatin accessibility during class switch recombination. J. Exp. Med. 206, 1817–1830 (2009).
Rajagopal, D. et al. Immunoglobulin switch mu sequence causes RNA polymerase II accumulation and reduces dA hypermutation. J. Exp. Med. 206, 1237–1244 (2009).
Xue, K., Rada, C. & Neuberger, M.S. The in vivo pattern of AID targeting to immunoglobulin switch regions deduced from mutation spectra in msh2−/− ung−/− mice. J. Exp. Med. 203, 2085–2094 (2006).
McBride, K.M. et al. Regulation of class switch recombination and somatic mutation by AID phosphorylation. J. Exp. Med. 205, 2585–2594 (2008).
Cheng, H.L. et al. Integrity of the AID serine-38 phosphorylation site is critical for class switch recombination and somatic hypermutation in mice. Proc. Natl. Acad. Sci. USA 106, 2717–2722 (2009).
Kovalchuk, A.L. et al. AID-deficient Bcl-xL transgenic mice develop delayed atypical plasma cell tumors with unusual Ig/Myc chromosomal rearrangements. J. Exp. Med. 204, 2989–3001 (2007).
Malynn, B.A. et al. N-myc can functionally replace c-myc in murine development, cellular growth, and differentiation. Genes Dev. 14, 1390–1399 (2000).
Takizawa, M. et al. AID expression levels determine the extent of cMyc oncogenic translocations and the incidence of B cell tumor development. J. Exp. Med. 205, 1949–1957 (2008).
Robbiani, D.F. et al. Activation induced deaminase is required for the chromosomal translocations in c-myc that lead to c-myc/IgH translocations. Cell 135, 1028–1038 (2008).
Chaudhuri, J. & Alt, F.W. Class-switch recombination: interplay of transcription, DNA deamination and DNA repair. Nat. Rev. Immunol. 4, 541–552 (2004).
Acknowledgements
We thank D. Schatz for comments on the manuscript; J. Chaudhuri (Memorial Sloan-Kettering Cancer Center) and F. Alt (Harvard University) for antibodies to AID; J. Simone for cell sorting; G. Gutierrez for technical assistance with the genome analyzer; and C. Ansarah-Sobrinho and S. Nelson for help with sequencing. Supported by the National Institutes of Health (Intramural Research Program of the National Institute of Arthritis and Musculoskeletal and Skin Diseases; and AI037526 to M.C.N.) and the Howard Hughes Medical Institute (M.C.N.).
Author information
Authors and Affiliations
Contributions
A.Y. did deep sequencing, cloning and conventional sequencing experiments; W.R. and H.-w.S. analyzed data; N.K. contributed data; Z.L. maintained the mouse colonies and cultured cells; D.F.R. contributed the Igk-AID mice; M.C.N. made suggestions for experiments and reviewed and wrote sections of the manuscript; R.C. designed the experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Text and Supplementary Tables 1–8 (PDF 3351 kb)
Rights and permissions
About this article
Cite this article
Yamane, A., Resch, W., Kuo, N. et al. Deep-sequencing identification of the genomic targets of the cytidine deaminase AID and its cofactor RPA in B lymphocytes. Nat Immunol 12, 62–69 (2011). https://doi.org/10.1038/ni.1964
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.1964
This article is cited by
-
Enhanced clinical assessment of hematologic malignancies through routine paired tumor and normal sequencing
Nature Communications (2023)
-
Transcribing malignancy: transcription-associated genomic instability in cancer
Oncogene (2018)
-
A licensing step links AID to transcription elongation for mutagenesis in B cells
Nature Communications (2018)
-
Phosphatidylinositol 3-kinase δ blockade increases genomic instability in B cells
Nature (2017)
-
Activation-induced cytidine deaminase targets SUV4-20-mediated histone H4K20 trimethylation to class-switch recombination sites
Scientific Reports (2017)