Abstract
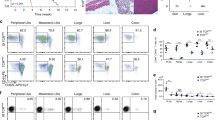
CD1 activates T cells, but the function and size of the possible human T cell repertoires that recognize each of the CD1 antigen-presenting molecules remain unknown. Using an experimental system that bypasses major histocompatibility complex (MHC) restriction and the requirement for defined antigens, we show that polyclonal T cells responded at higher rates to cells expressing CD1a than to those expressing CD1b, CD1c or CD1d. Unlike the repertoire of invariant natural killer T (NKT) cells, the CD1a-autoreactive repertoire contained diverse T cell antigen receptors (TCRs). Functionally, many CD1a-autoreactive T cells homed to skin, where they produced interleukin 22 (IL-22) in response to CD1a on Langerhans cells. The strong and frequent responses among genetically diverse donors define CD1a-autoreactive cells as a normal part of the human T cell repertoire and CD1a as a target of the TH22 subset of helper T cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Fowlkes, B.J. et al. A novel population of T-cell receptor α beta-bearing thymocytes which predominantly expresses a single Vβ gene family. Nature 329, 251–254 (1987).
Lantz, O. & Bendelac, A. An invariant T cell receptor alpha chain is used by a unique subset of major histocompatibility complex class I-specific CD4+ and CD4−8− T cells in mice and humans. J. Exp. Med. 180, 1097–1106 (1994).
Porcelli, S., Yockey, C.E., Brenner, M.B. & Balk, S.P. Analysis of T cell antigen receptor (TCR) expression by human peripheral blood CD4−8− α/β T cells demonstrates preferential use of several Vβ genes and an invariant TCR α chain. J. Exp. Med. 178, 1–16 (1993).
Benlagha, K., Weiss, A., Beavis, A., Teyton, L. & Bendelac, A. In vivo identification of glycolipid antigen-specific T cells using fluorescent CD1d tetramers. J. Exp. Med. 191, 1895–1903 (2000).
Gumperz, J.E., Miyake, S., Yamamura, T. & Brenner, M.B. Functionally distinct subsets of CD1d-restricted natural killer T cells revealed by CD1d tetramer staining. J. Exp. Med. 195, 625–636 (2002).
Matsuda, J.L. et al. Tracking the response of natural killer T cells to a glycolipid antigen using CD1d tetramers. J. Exp. Med. 192, 741–754 (2000).
Karadimitris, A. et al. Human CD1d-glycolipid tetramers generated by in vitro oxidative refolding chromatography. Proc. Natl. Acad. Sci. USA 98, 3294–3298 (2001).
Smiley, S.T., Kaplan, M.H. & Grusby, M.J. Immunoglobulin E production in the absence of interleukin-4-secreting CD1-dependent cells. Science 275, 977–979 (1997).
Cui, J. et al. Requirement for Vα14 NKT cells in IL-12-mediated rejection of tumors. Science 278, 1623–1626 (1997).
Godfrey, D.I. & Kronenberg, M. Going both ways: immune regulation via CD1d-dependent NKT cells. J. Clin. Invest. 114, 1379–1388 (2004).
Van Rhijn, I. et al. The bovine CD1 family contains group 1 CD1 proteins, but no functional CD1d. J. Immunol. 176, 4888–4893 (2006).
Looringh van Beeck, F.A. et al. Two canine CD1a proteins are differentially expressed in skin. Immunogenetics 60, 315–324 (2008).
Dascher, C.C. Evolutionary biology of CD1. Curr. Top. Microbiol. Immunol. 314, 3–26 (2007).
Kasmar, A., Van Rhijn, I. & Moody, D.B. The evolved functions of CD1 during infection. Curr. Opin. Immunol. 21, 397–403 (2009).
Roura-Mir, C. et al. Mycobacterium tuberculosis regulates CD1 antigen presentation pathways through TLR-2. J. Immunol. 175, 1758–1766 (2005).
Dougan, S.K., Kaser, A. & Blumberg, R.S. CD1 expression on antigen-presenting cells. Curr. Top. Microbiol. Immunol. 314, 113–141 (2007).
Sugita, M., Cernadas, M. & Brenner, M.B. New insights into pathways for CD1-mediated antigen presentation. Curr. Opin. Immunol. 16, 90–95 (2004).
Moody, D.B., Zajonc, D.M. & Wilson, I.A. Anatomy of CD1-lipid antigen complexes. Nat. Rev. Immunol. 5, 387–399 (2005).
Porcelli, S. et al. Recognition of cluster of differentiation 1 antigens by human CD4−CD8− cytolytic T lymphocytes. Nature 341, 447–450 (1989).
Rosat, J.P. et al. CD1-restricted microbial lipid antigen-specific recognition found in the CD8+ αβ T cell pool. J. Immunol. 162, 366–371 (1999).
Sieling, P.A. et al. Evidence for human CD4+ T cells in the CD1-restricted repertoire: derivation of mycobacteria-reactive T cells from leprosy lesions. J. Immunol. 164, 4790–4796 (2000).
Moody, D.B. The surprising diversity of lipid antigens for CD1-restricted T cells. Adv. Immunol. 89, 87–139 (2006).
Borg, N.A. et al. CD1d-lipid-antigen recognition by the semi-invariant NKT T-cell receptor. Nature 448, 44–49 (2007).
Shamshiev, A. et al. The αβ T cell response to self-glycolipids shows a novel mechanism of CD1b loading and a requirement for complex oligosaccharides. Immunity 13, 255–264 (2000).
Shamshiev, A. et al. Presentation of the same glycolipid by different CD1 molecules. J. Exp. Med. 195, 1013–1021 (2002).
Sieling, P.A. et al. Human double-negative T cells in systemic lupus erythematosus provide help for IgG and are restricted by CD1c. J. Immunol. 165, 5338–5344 (2000).
Vincent, M.S., Xiong, X., Grant, E.P., Peng, W. & Brenner, M.B. CD1a-, b-, and c-restricted TCRs recognize both self and foreign antigens. J. Immunol. 175, 6344–6351 (2005).
Zhou, D. et al. Lysosomal glycosphingolipid recognition by NKT cells. Science 306, 1786–1789 (2004).
Klein, E. et al. Properties of the K562 cell line, derived from a patient with chronic myeloid leukemia. Int. J. Cancer 18, 421–431 (1976).
Britten, C.M. et al. The use of HLA-A*0201-transfected K562 as standard antigen-presenting cells for CD8+ T lymphocytes in IFN-γ ELISPOT assays. J. Immunol. Methods 259, 95–110 (2002).
Shamshiev, A. et al. Self glycolipids as T-cell autoantigens. Eur. J. Immunol. 29, 1667–1675 (1999).
Roura-Mir, C. et al. CD1a and CD1c activate intrathyroidal T cells during Graves' disease and Hashimoto's thyroiditis. J. Immunol. 174, 3773–3780 (2005).
Moody, D.B. et al. Lipid length controls antigen entry into endosomal and nonendosomal pathways for CD1b presentation. Nat. Immunol. 3, 435–442 (2002).
Vincent, M.S. et al. CD1-dependent dendritic cell instruction. Nat. Immunol. 3, 1163–1168 (2002).
Meunier, L. et al. Quantification of CD1a, HLA-DR, and HLA class I expression on viable human Langerhans cells and keratinocytes. Cytometry 26, 260–264 (1996).
Yu, R.C., Abrams, D.C., Alaibac, M. & Chu, A.C. Morphological and quantitative analyses of normal epidermal Langerhans cells using confocal scanning laser microscopy. Br. J. Dermatol. 131, 843–848 (1994).
Chu, A. et al. Immunoelectron microscopic identification of Langerhans cells using a new antigenic marker. J. Invest. Dermatol. 78, 177–180 (1982).
Armerding, D. & Kupper, T.S. Functional cutaneous lymphocyte antigen can be induced in essentially all peripheral blood T lymphocytes. Int. Arch. Allergy Immunol. 119, 212–222 (1999).
Fuhlbrigge, R.C., Kieffer, J.D., Armerding, D. & Kupper, T.S. Cutaneous lymphocyte antigen is a specialized form of PSGL-1 expressed on skin-homing T cells. Nature 389, 978–981 (1997).
Agea, E. et al. Human CD1-restricted T cell recognition of lipids from pollens. J. Exp. Med. 202, 295–308 (2005).
Wang, X. et al. Natural killer T-cell autoreactivity leads to a specialized activation state. Blood 112, 4128–4138 (2008).
Wolk, K. et al. IL-22 increases the innate immunity of tissues. Immunity 21, 241–254 (2004).
Boniface, K. et al. IL-22 inhibits epidermal differentiation and induces proinflammatory gene expression and migration of human keratinocytes. J. Immunol. 174, 3695–3702 (2005).
Wolk, K. & Sabat, R. Interleukin-22: a novel T- and NK-cell derived cytokine that regulates the biology of tissue cells. Cytokine Growth Factor Rev. 17, 367–380 (2006).
Liang, S.C. et al. Interleukin (IL)-22 and IL-17 are coexpressed by Th17 cells and cooperatively enhance expression of antimicrobial peptides. J. Exp. Med. 203, 2271–2279 (2006).
Nograles, K.E. et al. IL-22-producing “T22” T cells account for upregulated IL-22 in atopic dermatitis despite reduced IL-17-producing TH17 T cells. J. Allergy Clin. Immunol. 123, 1244–1252 (2009).
Duhen, T., Geiger, R., Jarrossay, D., Lanzavecchia, A. & Sallusto, F. Production of interleukin 22 but not interleukin 17 by a subset of human skin-homing memory T cells. Nat. Immunol. 10, 857–863 (2009).
Trifari, S., Kaplan, C.D., Tran, E.H., Crellin, N.K. & Spits, H. Identification of a human helper T cell population that has abundant production of interleukin 22 and is distinct from TH-17, TH1 and TH2 cells. Nat. Immunol. 10, 864–871 (2009).
Eyerich, S. et al. Th22 cells represent a distinct human T cell subset involved in epidermal immunity and remodeling. J. Clin. Invest. 119, 3573–3585 (2009).
Veldhoen, M. et al. The aryl hydrocarbon receptor links TH17-cell-mediated autoimmunity to environmental toxins. Nature 453, 106–109 (2008).
Clark, R.A. et al. A novel method for the isolation of skin resident T cells from normal and diseased human skin. J. Clin. Invest. 126, 1059–1070 (2006).
Strid, J., Tigelaar, R.E. & Hayday, A.C. Skin immune surveillance by T cells–a new order? Semin. Immunol. 21, 110–120 (2009).
Foster, C.A. et al. Human epidermal T cells predominantly belong to the lineage expressing α/β T cell receptor. J. Exp. Med. 171, 997–1013 (1990).
Clark, R.A. et al. The vast majority of CLA+ T cells are resident in normal skin. J. Immunol. 176, 4431–4439 (2006).
Alam, M.S. et al. Notch signaling drives IL-22 secretion in CD4+ T cells by stimulating the aryl hydrocarbon receptor. Proc. Natl. Acad. Sci. USA 107, 5943–5948 (2010).
Korn, T., Bettelli, E., Oukka, M. & Kuchroo, V.K. IL-17 and Th17 Cells. Annu. Rev. Immunol. 27, 485–517 (2009).
Boniface, K. et al. A role for T cell-derived interleukin 22 in psoriatic skin inflammation. Clin. Exp. Immunol. 150, 407–415 (2007).
Pena-Cruz, V. et al. Extraction of human Langerhans cells: a method for isolation of epidermis-resident dendritic cells. J. Immunol. Methods 255, 83–91 (2001).
Acknowledgements
We thank D.C. Barral, M. Brenner and M. Relloso (Harvard Medical School) and M. Sugita (Kyoto University) for CD1 plasmid constructs; J. Gumperz (University of Wisconsin) and M. Brigl (Harvard Medical School) for human NKT cell lines; G. Losyev for cell sorting; and K. Magalhães, S. Huang, I.C. Ho and R. Grenningloh for technical advice. Supported by the National Institute of Arthritis, Musculoskeletal and Skin Diseases (048632 to D.B.M. and AR056720 to R.A.C.), the National Institute of Allergy and Infectious Diseases (AI071155 to D.B.M., and AI054456 and AI056299), the Damon Runyon Cancer Research Foundation (R.A.C.), the Burroughs Wellcome Fund (D.B.M.) and the National Psoriasis Foundation (A.d.J.).
Author information
Authors and Affiliations
Contributions
A.d.J. designed and did the experiments; A.d.J. and D.B.M. prepared the manuscript; D.B.M. supervised the experiments; V.P.-C. isolated LCs from human epidermis and lymphocytes from the dermis; T.-Y.C. did T cell culture and immunoblot analysis; I.V.R. assisted in experiments; and R.A.C. provided T cells isolated from human skin biopsies.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Table 1 (PDF 2806 kb)
Rights and permissions
About this article
Cite this article
de Jong, A., Peña-Cruz, V., Cheng, TY. et al. CD1a-autoreactive T cells are a normal component of the human αβ T cell repertoire. Nat Immunol 11, 1102–1109 (2010). https://doi.org/10.1038/ni.1956
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.1956
This article is cited by
-
Staphylococcal phosphatidylglycerol antigens activate human T cells via CD1a
Nature Immunology (2023)
-
CD1a promotes systemic manifestations of skin inflammation
Nature Communications (2022)
-
CD4 and CD8 co-receptors modulate functional avidity of CD1b-restricted T cells
Nature Communications (2022)
-
Human T cells engineered with a leukemia lipid-specific TCR enables donor-unrestricted recognition of CD1c-expressing leukemia
Nature Communications (2021)
-
A T-cell receptor escape channel allows broad T-cell response to CD1b and membrane phospholipids
Nature Communications (2019)