Abstract
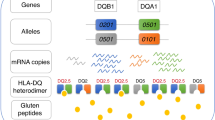
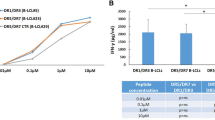
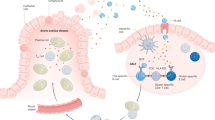
Celiac disease driven by an antigluten T cell response is strongly associated with the histocompatibility antigen HLA-DQ2.5 but is barely associated with HLA-DQ2.2. Yet these molecules have very similar peptide-binding motifs and both present gluten T cell epitopes. We found that DQ2.5+ antigen-presenting cells (APCs) had greater stability of bound peptides and protracted gluten presentation relative to that of DQ2.2+ cells. The improved ability of DQ2.5 to retain its peptide cargo can be ascribed to a polymorphism of DQα22 whereby DQ2.5 (tyrosine) can establish a hydrogen bond to the peptide main chain but DQ2.2 (phenylalanine) cannot. Our findings suggest that the kinetic stability of complexes of peptide and major histocompatibility complex (MHC) is of importance for the association of HLA with disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Change history
13 September 2009
NOTE: In the version of this article initially published online, the symbols for 0 h and 18 h were reversed in the legend for Figure 4 . The correct symbols are 0 h (▿) and 18 h (▵). The error has been corrected for the print, PDF and HTML versions of this article.
References
Sollid, L.M. Coeliac disease: dissecting a complex inflammatory disorder. Nat. Rev. Immunol. 2, 647–655 (2002).
Kagnoff, M.F. Celiac disease: pathogenesis of a model immunogenetic disease. J. Clin. Invest. 117, 41–49 (2007).
Lundin, K.E.A. et al. Gliadin-specific, HLA-DQ(α1*0501,β1*0201) restricted T cells isolated from the small intestinal mucosa of celiac disease patients. J. Exp. Med. 178, 187–196 (1993).
Kim, C.Y., Quarsten, H., Bergseng, E., Khosla, C. & Sollid, L.M. Structural basis for HLA-DQ2-mediated presentation of gluten epitopes in celiac disease. Proc. Natl. Acad. Sci. USA 101, 4175–4179 (2004).
Henderson, K.N. et al. A structural and immunological basis for the role of human leukocyte antigen DQ8 in celiac disease. Immunity 27, 23–34 (2007).
Bergseng, E., Sidney, J., Sette, A. & Sollid, L.M. Analysis of the binding of gluten T-cell epitopes to various human leukocyte antigen class II molecules. Hum. Immunol. 69, 94–100 (2008).
Sollid, L.M. et al. Evidence for a primary association of celiac disease to a particular HLA-DQ α/β heterodimer. J. Exp. Med. 169, 345–350 (1989).
Johansen, B.H. et al. Both α and β chain polymorphisms determine the specificity of the disease-associated HLA-DQ2 molecules, with β chain residues being most influential. Immunogenetics 45, 142–150 (1996).
van de Wal, Y. et al. Unique peptide binding characteristics of the disease-associated DQ(α1*0501, β1*0201) vs the non-disease-associated DQ(α1*0201, β1*0202) molecule. Immunogenetics 46, 484–492 (1997).
Vader, W. et al. The HLA-DQ2 gene dose effect in celiac disease is directly related to the magnitude and breadth of gluten-specific T cell responses. Proc. Natl. Acad. Sci. USA 100, 12390–12395 (2003).
Qiao, S.W. et al. Refining the rules of gliadin T cell epitope binding to the disease-associated DQ2 molecule in celiac disease: importance of proline spacing and glutamine deamidation. J. Immunol. 175, 254–261 (2005).
van de Wal, Y., Kooy, Y.M.C., Drijfhout, J.W., Amons, R. & Koning, F. Peptide binding characteristics of the coeliac disease-associated DQ(α1*0501, β1*0201) molecule. Immunogenetics 44, 246–253 (1996).
Vartdal, F. et al. The peptide binding motif of the disease associated HLA-DQ(α1*0501, β1*0201) molecule. Eur. J. Immunol. 26, 2764–2772 (1996).
Wiesner, M. et al. Dominance of an alternative CLIP sequence in the celiac disease associated HLA-DQ2 molecule. Immunogenetics 60, 551–555 (2008).
Fallang, L.E. et al. Complexes of two cohorts of CLIP peptides and HLA-DQ2 of the autoimmune DR3–DQ2 haplotype are poor substrates for HLA-DM. J. Immunol. 181, 5451–5461 (2008).
Pashine, A. et al. Interaction of HLA-DR with an acidic face of HLA-DM disrupts sequence-dependent interactions with peptides. Immunity 19, 183–192 (2003).
Hall, F.C. et al. Relationship between kinetic stability and immunogenicity of HLA-DR4/peptide complexes. Eur. J. Immunol. 32, 662–670 (2002).
Lazarski, C.A. et al. The kinetic stability of MHC class II:peptide complexes is a key parameter that dictates immunodominance. Immunity 23, 29–40 (2005).
Lanzavecchia, A., Reid, P.A. & Watts, C. Irreversible association of peptides with class II MHC molecules in living cells. Nature 357, 249–252 (1992).
Shan, L. et al. Structural basis for gluten intolerance in celiac sprue. Science 297, 2275–2279 (2002).
Lee, K.H., Wucherpfennig, K.W. & Wiley, D.C. Structure of a human insulin peptide–HLA-DQ8 complex and susceptibility to type 1 diabetes. Nat. Immunol. 2, 501–507 (2001).
Siebold, C. et al. Crystal structure of HLA-DQ0602 that protects against type 1 diabetes and confers strong susceptibility to narcolepsy. Proc. Natl. Acad. Sci. USA 101, 1999–2004 (2004).
Shishova, E.Y., Di, C.L., Emig, F.A., Ash, D.E. & Christianson, D.W. Probing the specificity determinants of amino acid recognition by arginase. Biochemistry 48, 121–131 (2009).
Henrickson, S.E. et al. T cell sensing of antigen dose governs interactive behavior with dendritic cells and sets a threshold for T cell activation. Nat. Immunol. 9, 282–291 (2008).
Karell, K. et al. HLA types in celiac disease patients not carrying the DQA1*05–DQB1*02 (DQ2) heterodimer: results from the European Genetics Cluster on Celiac Disease. Hum. Immunol. 64, 469–477 (2003).
Ploski, R., Ek, J., Thorsby, E. & Sollid, L.M. On the HLA-DQ(α1*0501, β1*0201)-associated susceptibility in celiac disease: a possible gene dosage effect of DQB1*0201. Tissue Antigens 41, 173–177 (1993).
Stern, L.J. et al. Crystal structure of the human class II MHC protein HLA-DR1 complexed with an influenza virus peptide. Nature 368, 215–221 (1994).
Nelson, C.A. & Fremont, D.H. Structural principles of MHC class II antigen presentation. Rev. Immunogenet. 1, 47–59 (1999).
Fremont, D.H., Monnaie, D., Nelson, C.A., Hendrickson, W.A. & Unanue, E.R. Crystal structure of I-Ak in complex with a dominant epitope of lysozyme. Immunity 8, 305–317 (1998).
Price, P. et al. The genetic basis for the association of the 8.1 ancestral haplotype (A1, B8, DR3) with multiple immunopathological diseases. Immunol. Rev. 167, 257–274 (1999).
Erlich, H., Lee, J.S., Petersen, J.W., Bugawan, T. & DeMars, R. Molecular analysis of HLA class I and class II antigen loss mutants reveals a homozygous deletion of the DR, DQ, and part of the DP region: implications for class II gene order. Hum. Immunol. 16, 205–219 (1986).
Johansen, B.H. et al. Binding of peptides to HLA-DQ molecules: peptide binding properties of the disease-associated HLA-DQ(α1*0501, β1*0201) molecule. Int. Immunol. 6, 453–461 (1994).
Quarsten, H. et al. Staining of celiac disease-relevant T cells by peptide-DQ2 multimers. J. Immunol. 167, 4861–4868 (2001).
Viken, H.D. et al. Characterization of an HLA-DQ2-specific monoclonal antibody. Influence of amino acid substitutions in DQβ1*0202. Hum. Immunol. 42, 319–327 (1995).
Sloan, V.S. et al. Mediation by HLA-DM of dissociation of peptides from HLA-DR. Nature 375, 802–806 (1995).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Davis, I.W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Acknowledgements
We thank E. Mellins (Stanford University) for 8.1.6 B-LCLs and S2 cells expressing soluble HLA-DM; B. Roep (Leiden University Medical Center) for 721.82 B-LCLs; H. Soltani for flow-assisted cell sorting; T. Svingerud for assistance in making the sDQ2.2 and sDQ2.2 Fα22Y constructs; C. Khosla (Stanford University) and U. Jüse (University of Oslo) for fluorescence-labeled peptides; and B. Jabri and S. Buus for critical reading of the manuscript. Supported by the Research Council of Norway (L.M.S.), the Biomedical Research Council of Singapore (C.-Y.K.) and the Life Sciences Institute, National University of Singapore (C.-Y.K).
Author information
Authors and Affiliations
Contributions
L.-E.F. and E.B. designed and did experiments, analyzed data and contributed to the writing of the manuscript; K.H. contributed to data analysis and revised the manuscript; A.B.-L. did experiments, contributed to data analysis and revised the manuscript; C.-Y.K. contributed to data analysis and to the writing of the manuscript; and L.M.S. directed the research, designed experiments, analyzed data and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and TSupplementary able 1 (PDF 361 kb)
Rights and permissions
About this article
Cite this article
Fallang, LE., Bergseng, E., Hotta, K. et al. Differences in the risk of celiac disease associated with HLA-DQ2.5 or HLA-DQ2.2 are related to sustained gluten antigen presentation. Nat Immunol 10, 1096–1101 (2009). https://doi.org/10.1038/ni.1780
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.1780
This article is cited by
-
Influence of HLA on clinical and analytical features of pediatric celiac disease
BMC Gastroenterology (2019)
-
Opposing T cell responses in experimental autoimmune encephalomyelitis
Nature (2019)
-
HLA-DQ2 and -DQ8 genotype frequency in Syrian celiac disease children: HLA-DQ relative risks evaluation
BMC Gastroenterology (2018)
-
The roles of MHC class II genes and post-translational modification in celiac disease
Immunogenetics (2017)
-
Risk factors for celiac disease
Italian Journal of Pediatrics (2015)